

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149129.7 + phase: 0
(726 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
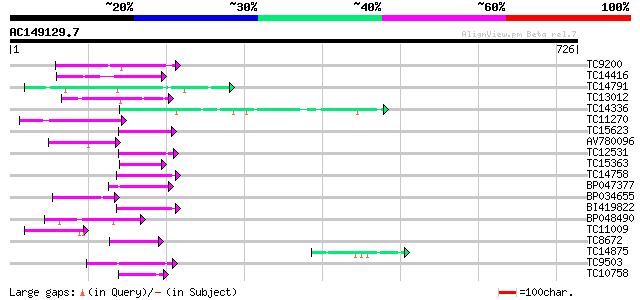
Score E
Sequences producing significant alignments: (bits) Value
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 67 8e-12
TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, parti... 60 2e-09
TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), par... 57 1e-08
TC13012 similar to PIR|T50004|T50004 RNA binding protein-like - ... 54 7e-08
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 54 9e-08
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 50 1e-06
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 48 6e-06
AV780096 46 2e-05
TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, parti... 46 2e-05
TC15363 similar to UP|Q84LL8 (Q84LL8) Salt tolerance protein 4, ... 45 3e-05
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 45 4e-05
BP047377 45 5e-05
BP034655 45 5e-05
BI419822 44 7e-05
BP048490 44 7e-05
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 43 2e-04
TC8672 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partia... 43 2e-04
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 42 3e-04
TC9503 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprot... 42 3e-04
TC10758 42 3e-04
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 67.4 bits (163), Expect = 8e-12
Identities = 51/166 (30%), Positives = 81/166 (48%), Gaps = 6/166 (3%)
Frame = +1
Query: 59 QEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDD-VAGDDEHASEENVHEH 117
+E +EVEEE EE +EEEG+ +EE EE E V E E +DD+ + E +E +
Sbjct: 211 EEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDDEPIQKLLEPMGKEQIISL 390
Query: 118 LADVKEEEQVEVIKETQKQKELEV-----FVGGLDKEATEHDLRKVFSKVGEITEVRMTV 172
LAD + +V +K + +V FV GL + T L F + G I + +
Sbjct: 391 LADAASNHR-DVADRIRKVADEDVSHRKIFVHGLGWDTTATTLVYAFQQYGAIEDCKAVT 567
Query: 173 NPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQ 218
+ T ++KG+ + F++ + AL E + K+ G +T CQ
Sbjct: 568 DKVTGKSKGYGFILFKSRRGARNALKEPQ------KKIGNRMTACQ 687
>TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (28%)
Length = 695
Score = 59.7 bits (143), Expect = 2e-09
Identities = 36/141 (25%), Positives = 70/141 (49%)
Frame = +1
Query: 60 EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLA 119
+ D ++E + + E+ + G E V E E +E G+D ++ G +E EE +E
Sbjct: 316 QTSDWAQQEEDNTLTLEDQEEGAEGVSETEAGWEP-NGEDAEIEGAEEGEGEEGDYE--- 483
Query: 120 DVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRN 179
+ +E ++FVG L +A H L ++F++VG + + N +T ++
Sbjct: 484 --------------EPPEEAKLFVGNLPYDADGHKLAELFNQVGTVDLAEVIYNRETDQS 621
Query: 180 KGFALLRFETVEHVKRALAEL 200
+GF + +TVE ++A+ +L
Sbjct: 622 RGFGFVTMKTVEDAEKAVDKL 684
>TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), partial (37%)
Length = 1779
Score = 57.0 bits (136), Expect = 1e-08
Identities = 68/312 (21%), Positives = 116/312 (36%), Gaps = 43/312 (13%)
Frame = +2
Query: 19 IDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEP-------EE 71
+D+ +D E + + E+ DD +E+++ EA E EP E+
Sbjct: 191 LDQSNNTMAVDEEPEEQDQETEDQPQIDD----AVEEEQDPEAPATTESEPQQQQETLED 358
Query: 72 NVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIK 131
N D + E V ++++ +D EE V L E+ ++K
Sbjct: 359 NADAPNESSAAAAAAASESVANGSNNNEEEEEEEDLELEEEPVERLLEPFTREQLHSLVK 538
Query: 132 ETQKQ----------------KELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ 175
+ ++ ++FV GL + T L VFSK GEI + + +
Sbjct: 539 QAAEKYPDFVENVRQLADVDPAHRKIFVHGLGWDTTAETLTSVFSKYGEIEDCKAVTDKV 718
Query: 176 TKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDT------------- 222
T ++KG+A + F+ + +RA LKNP K+ G T+ Q +
Sbjct: 719 TGKSKGYAFILFKHRDGCRRA---LKNP---QKKIGNRTTSSQLASAGPVPATPPNAPPV 880
Query: 223 -------LYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSH 275
+++ N+ + L E K +G L L D N G G A + S
Sbjct: 881 SEYTQRKIFVSNVSSDIEPLKLLEFFKQFGEVEDGPLGL---DKNTGRPKGFALFVYRSV 1051
Query: 276 SDSKDAYKRLQK 287
+K A + K
Sbjct: 1052ESAKKALEEPNK 1087
Score = 28.1 bits (61), Expect = 5.2
Identities = 19/93 (20%), Positives = 37/93 (39%), Gaps = 2/93 (2%)
Frame = +2
Query: 329 WDE--DYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESITSAGLGE 386
WD + + ++ KYG +E + + + K Y F+ F C ++ +
Sbjct: 635 WDTTAETLTSVFSKYGEIEDCKAVTDKVTGKSKGYAFILFKHRDG---CRRALKNPQKKI 805
Query: 387 GNKKAKVRARLSRPLPKVRGKHVPHRDISGRKI 419
GN+ + + P+P P + + RKI
Sbjct: 806 GNRTTSSQLASAGPVPATPPNAPPVSEYTQRKI 904
>TC13012 similar to PIR|T50004|T50004 RNA binding protein-like - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (84%)
Length = 625
Score = 54.3 bits (129), Expect = 7e-08
Identities = 46/168 (27%), Positives = 78/168 (46%), Gaps = 25/168 (14%)
Frame = +3
Query: 67 EEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKE-EE 125
E+ ++N+DEEE + E+ +V E+E D D A DD+ A+ + + E +KE EE
Sbjct: 36 EQNQKNMDEEEHEVYGGEIPDV----GEMEADVDMSAADDDAAAVKELDEMKRRLKEMEE 203
Query: 126 QVEVIKETQKQKELE------------------------VFVGGLDKEATEHDLRKVFSK 161
+ ++E Q + E E VFVG +D T ++++ F
Sbjct: 204 EAAALREMQAKVEKEIGTVQDPAAAAASQANKEETDGRSVFVGNVDYACTPEEVQQHFQS 383
Query: 162 VGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQ 209
G + V + + + + KGFA + F E V+ AL L ++G+Q
Sbjct: 384 CGTVNRVTI-LTDKFGQPKGFAYVEFVEAEAVQEAL-RLNESELHGRQ 521
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 53.9 bits (128), Expect = 9e-08
Identities = 81/370 (21%), Positives = 144/370 (38%), Gaps = 26/370 (7%)
Frame = +2
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
+F+ LDK L FS G I ++ + + ++KG+ ++FET E + A+ +L
Sbjct: 98 IFIKNLDKAIDHKALHDTFSTFGNILSCKVATD-SSGQSKGYGFVQFETEEAAQNAIEKL 274
Query: 201 KNPVINGKQCGI----------TITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFK 250
++N KQ + + T + +++ N+ +S + LK +G +
Sbjct: 275 NGMLLNDKQVYVGPFLRKQERESSTDKAKFNNVFVKNLSESTTDDELKNVFGEFG--TIT 448
Query: 251 DLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRL---QKTDVVFGVDKPAEVS------ 301
++ D + + G F+ F + D+ A + L + D + V K + S
Sbjct: 449 SAVVMRDGDGKSKCFG--FVNFENADDAARAVESLNGKKVDDKEWYVGKAQKKSEREQEL 622
Query: 302 ---FANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARR 358
F S + D Q +++ L S ++ ++ L YG + ++ ++ G R
Sbjct: 623 KNKFEQSMKEAADKY--QGANLYVKNLDDSIADEKLKELFSSYGTITSCKVMRDPNGISR 796
Query: 359 KNYGFVTFGTHAAAVECAESITSAGLGEGNKKAKVRARLSRPLPKVRGKHVPHRDISGRK 418
+ GFV F T A S L E N K V S+PL + R +
Sbjct: 797 GS-GFVAFSTPEEA--------SRALLEMNGKMVV----SKPLYVTLAQRKEDRRARLQA 937
Query: 419 IGRLERPSRSRPAPRSRPAPQSRPA---HVFRRLG-SRIPPARPSSARNRRPVTSIPVRA 474
RP PAPR P P +F G I P++P ++ V +
Sbjct: 938 QFAQMRPVAMPPAPRVPMYPPGGPGIGQQIFYGQGPPAIIPSQPGFGYQQQLVPGM---- 1105
Query: 475 RPASPPARSY 484
RP P ++
Sbjct: 1106RPGGAPVPNF 1135
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 50.4 bits (119), Expect = 1e-06
Identities = 38/139 (27%), Positives = 69/139 (49%), Gaps = 2/139 (1%)
Frame = +3
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
E++ + ++E E+DE + E H V E E+DE +E G+E EEE E
Sbjct: 318 EKLNQGVEEEEEDEEDEEEQTPHPTRGRSRV------EHAQEEDEEEENGEEEEEEEEVV 479
Query: 73 VDEEEGDTGDEEVEEVE-YVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVE-VI 130
V++E+G+ ++E EE E V EV+G D G + + ++ + ++ +E ++
Sbjct: 480 VEDEQGNEDEDEDEEEEGVVVVEVKGRKVDSKGLHSGSGTPSKVAYVIPLPDKRTLELIL 659
Query: 131 KETQKQKELEVFVGGLDKE 149
+ QK+ V+ +D E
Sbjct: 660 DKLQKKDTYGVYAEPVDPE 716
Score = 33.1 bits (74), Expect = 0.16
Identities = 28/94 (29%), Positives = 46/94 (48%), Gaps = 14/94 (14%)
Frame = +3
Query: 58 GQEAGDEVEEEPEENVDEEEGDTGDEEVE---------EVEYVYEEVEGDDDDVAGDDEH 108
G++ ++ E+ + V+EEE D DEE + VE+ EE D+++ G++E
Sbjct: 291 GRKKKLKLVEKLNQGVEEEEEDEEDEEEQTPHPTRGRSRVEHAQEE---DEEEENGEEEE 461
Query: 109 ASEENVHE-----HLADVKEEEQVEVIKETQKQK 137
EE V E D EEE+ V+ E + +K
Sbjct: 462 EEEEVVVEDEQGNEDEDEDEEEEGVVVVEVKGRK 563
Score = 27.3 bits (59), Expect = 8.8
Identities = 15/43 (34%), Positives = 26/43 (59%)
Frame = +2
Query: 24 KDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVE 66
+ ++ + +Y + + E+ D+D DERG E++E Q AG E E
Sbjct: 203 RPRNVRYNIDYLDEEDEDEDEDEDDDERGREKEEAQ-AG*EAE 328
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 47.8 bits (112), Expect = 6e-06
Identities = 21/74 (28%), Positives = 39/74 (52%)
Frame = +1
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++F+GGL + L FS G + E R+ V+ T R++GF + F++ E AL+
Sbjct: 187 KLFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDTGRSRGFGFVSFDSEESASSALSS 366
Query: 200 LKNPVINGKQCGIT 213
+ +NG+ ++
Sbjct: 367 MDGQDLNGRNIRVS 408
>AV780096
Length = 564
Score = 46.2 bits (108), Expect = 2e-05
Identities = 31/102 (30%), Positives = 54/102 (52%), Gaps = 10/102 (9%)
Frame = +1
Query: 50 ERGIE-QDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDD--------D 100
ER ++ +D + G + EEE EE +EE D +E+ EE+E V DD D
Sbjct: 4 ERNLDLRDMARRNGSDSEEEEEEQQEEEVHDQIEEDDEELEAVARSASSDDEPDESGGED 183
Query: 101 DVAGDDEHASEE-NVHEHLADVKEEEQVEVIKETQKQKELEV 141
VA DD++ +E N + +E+ +++ +++ +KQK E+
Sbjct: 184 AVAADDDNQDDEGNAADPEISKREKARLKELQQLKKQKIQEI 309
>TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, partial (39%)
Length = 688
Score = 45.8 bits (107), Expect = 2e-05
Identities = 24/77 (31%), Positives = 41/77 (53%)
Frame = -1
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+VFVGGL E + ++K F + G+I E + + T R+KG+ + F E RA +
Sbjct: 358 KVFVGGLAWETQKETMKKYFEQFGDILEAVVITDKATGRSKGYGFVTFREPEAAMRACVD 179
Query: 200 LKNPVINGKQCGITITT 216
PVI+G++ + +
Sbjct: 178 AA-PVIDGRRANCNLAS 131
>TC15363 similar to UP|Q84LL8 (Q84LL8) Salt tolerance protein 4, partial
(79%)
Length = 1163
Score = 45.4 bits (106), Expect = 3e-05
Identities = 21/60 (35%), Positives = 35/60 (58%)
Frame = +1
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
+ V G+ +EA E DL+ F + GEI + + ++ +T KG+AL+ +E E K A+ L
Sbjct: 388 ILVTGVHEEAQEDDLQNAFGEYGEIKNLHLNLDRRTGFVKGYALIEYERAEEAKNAIENL 567
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 45.1 bits (105), Expect = 4e-05
Identities = 24/81 (29%), Positives = 42/81 (51%)
Frame = +1
Query: 138 ELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRAL 197
E FVGGL H L + FS GEI + ++ + +T R++GF + F + E ++ A+
Sbjct: 79 EYRCFVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRSRGFGFVTFTSEEAMRSAI 258
Query: 198 AELKNPVINGKQCGITITTCQ 218
+ ++G+ IT+ Q
Sbjct: 259 EGMNGNELDGR--NITVNEAQ 315
Score = 30.8 bits (68), Expect = 0.80
Identities = 26/75 (34%), Positives = 28/75 (36%), Gaps = 5/75 (6%)
Frame = +1
Query: 656 GAYSSRQDPSYGRGS---LGGSDGGIYSSSYGGDYLSRGSDVGGSSYSSMYSS--RGMNG 710
G R YG G GG GG S YGG G GG Y G G
Sbjct: 331 GGGGGRGGGGYGGGGGRREGGYGGGGGSRGYGGGGGGYGGG-GGGGYGGRREGGYGGDGG 507
Query: 711 SSSYMSGSGGGSRSY 725
S Y SG GGG ++
Sbjct: 508 GSRYGSGGGGGGGNW 552
Score = 30.4 bits (67), Expect = 1.0
Identities = 30/103 (29%), Positives = 35/103 (33%), Gaps = 16/103 (15%)
Frame = -1
Query: 425 PSRSRPAPRSRPAPQSRPAHVFRRLGSRIPPARPSSARNRRPVTSIPVRARPASPPARSY 484
PSR P P P P P R PP P S R P P R P PP R++
Sbjct: 485 PSRRPP*PPPPPPPYPPPPPP*PR--DPPPPPYPPSRRPPPPPYPPPPRPPPPPPPPRAW 312
Query: 485 R----------------RLAAAYPKSSMKRDYGRPVDLPPPRS 511
+A S +K P+DLP RS
Sbjct: 311 ASFTVMLRPSSSLPFIPSIALLIASSEVKVTNPNPLDLPVSRS 183
Score = 30.0 bits (66), Expect = 1.4
Identities = 23/81 (28%), Positives = 25/81 (30%)
Frame = +1
Query: 621 GGSASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQDPSYGRGSLGGSDGGIYS 680
GG R YG R GY G S G Y YG GG Y
Sbjct: 331 GGGGGRGGGGYGGGGGRREGGYGGGGGSRGYGGGGGGYGGGGGGGYGGRREGG-----YG 495
Query: 681 SSYGGDYLSRGSDVGGSSYSS 701
GG G GG ++ S
Sbjct: 496 GDGGGSRYGSGGGGGGGNWRS 558
Score = 29.6 bits (65), Expect = 1.8
Identities = 21/90 (23%), Positives = 35/90 (38%), Gaps = 1/90 (1%)
Frame = +1
Query: 300 VSFANSFIDLGDDIMAQVK-TVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARR 358
+S F+ D MA V+ F+ L + D + YG + ++ + R
Sbjct: 25 LSLFPQFLGFSDSAMADVEYRCFVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRS 204
Query: 359 KNYGFVTFGTHAAAVECAESITSAGLGEGN 388
+ +GFVTF + A E + L N
Sbjct: 205 RGFGFVTFTSEEAMRSAIEGMNGNELDGRN 294
Score = 29.3 bits (64), Expect = 2.3
Identities = 18/50 (36%), Positives = 18/50 (36%)
Frame = +1
Query: 672 GGSDGGIYSSSYGGDYLSRGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGG 721
GG GG YGG R GG SRG G G GGG
Sbjct: 328 GGGGGGRGGGGYGGGGGRREGGYGGGG-----GSRGYGGGGGGYGGGGGG 462
>BP047377
Length = 564
Score = 44.7 bits (104), Expect = 5e-05
Identities = 24/83 (28%), Positives = 43/83 (50%)
Frame = +3
Query: 127 VEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLR 186
V+ + +Q Q VFVG + +ATE L ++ +VG + R+ ++ +T + KG+
Sbjct: 96 VDFMASSQSQHRC-VFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDRETGKPKGYGFCE 272
Query: 187 FETVEHVKRALAELKNPVINGKQ 209
++ E A L+ ING+Q
Sbjct: 273 YKDEETALSARRNLQGYEINGRQ 341
>BP034655
Length = 517
Score = 44.7 bits (104), Expect = 5e-05
Identities = 27/85 (31%), Positives = 44/85 (51%)
Frame = +1
Query: 56 DEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVH 115
D G DE EEE EE +EEE + +EE EE + +E + D+D + H NVH
Sbjct: 121 DFGLSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRVSPNVH 300
Query: 116 EHLADVKEEEQVEVIKETQKQKELE 140
++ + +V +I ++++Q E
Sbjct: 301 QN--PTSDSVEVILIPDSEQQNYSE 369
>BI419822
Length = 380
Score = 44.3 bits (103), Expect = 7e-05
Identities = 25/81 (30%), Positives = 41/81 (49%)
Frame = +2
Query: 138 ELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRAL 197
E FVGGL L K FS GEI E ++ + +T R++GF + F + + +K A+
Sbjct: 74 EFRCFVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVTFASEQAMKDAI 253
Query: 198 AELKNPVINGKQCGITITTCQ 218
+ ++G+ IT+ Q
Sbjct: 254 EAMNGQNLDGR--NITVNEAQ 310
Score = 29.6 bits (65), Expect = 1.8
Identities = 15/68 (22%), Positives = 30/68 (44%)
Frame = +2
Query: 321 FIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESIT 380
F+ L + D + + YG + + ++ + R + +GFVTF + A + E++
Sbjct: 86 FVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVTFASEQAMKDAIEAMN 265
Query: 381 SAGLGEGN 388
L N
Sbjct: 266 GQNLDGRN 289
>BP048490
Length = 452
Score = 44.3 bits (103), Expect = 7e-05
Identities = 30/145 (20%), Positives = 62/145 (42%), Gaps = 16/145 (11%)
Frame = +3
Query: 45 DDDYDERGIEQDEGQEAG------DEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGD 98
DD+ E E+++ + A V++ + D+E+ D EV++ ++++ D
Sbjct: 27 DDESSESSDEENDSKPATTTVAKPSAVKKSASSDSDDEDSSDSDSEVKK-----DKMDVD 191
Query: 99 DDDVAGDDEHASEENVHEHLADVKEEEQVEVIK----------ETQKQKELEVFVGGLDK 148
+ D + D + E++ VK+ E V+ +T+ + +FVG L
Sbjct: 192 ESDSSEDSDEEDEKSTKTPQKKVKDVEMVDASSGKNAPKTPAAQTENEGSKTLFVGNLSF 371
Query: 149 EATEHDLRKVFSKVGEITEVRMTVN 173
D+ F GE+ +VR+ +
Sbjct: 372 SVQRSDVENFFKDCGEVVDVRLATD 446
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 43.1 bits (100), Expect = 2e-04
Identities = 31/103 (30%), Positives = 48/103 (46%), Gaps = 21/103 (20%)
Frame = +2
Query: 20 DEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGD 79
DE + D+ D + + + D GDDD DE E++ G E G +P+++ D+++ D
Sbjct: 26 DEDDDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEEGGVEGGRGGGGDPDDDDDDDDDD 205
Query: 80 TGDEEVEE---VEYVY------------------EEVEGDDDD 101
+EE EE EY+ EE EGDD+D
Sbjct: 206 DEEEEEEEDLGTEYLVRRTVAAAEDEEASSDFEPEEDEGDDND 334
Score = 37.7 bits (86), Expect = 0.007
Identities = 25/83 (30%), Positives = 40/83 (48%)
Frame = +2
Query: 25 DEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEE 84
D+ D +D+ E + EE +G + R + E +EA + E E E+EGD D +
Sbjct: 179 DDDDDDDDDDEEEEEEEDLGTEYLVRRTVAAAEDEEASSDFEPE------EDEGDDNDND 340
Query: 85 VEEVEYVYEEVEGDDDDVAGDDE 107
E V + + D D +GDD+
Sbjct: 341 DGEKAGVPSKRKRSDKDGSGDDD 409
Score = 35.8 bits (81), Expect = 0.025
Identities = 28/76 (36%), Positives = 38/76 (49%), Gaps = 7/76 (9%)
Frame = +2
Query: 44 GDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYE-------EVE 96
GD+D D+ G E D+G E D+ ++ P D++ DEE EE E E + +
Sbjct: 23 GDEDDDDDG-EDDDGDE--DDDDDAPGGGDDDD-----DEEDEEEEGGVEGGRGGGGDPD 178
Query: 97 GDDDDVAGDDEHASEE 112
DDDD DDE EE
Sbjct: 179 DDDDDDDDDDEEEEEE 226
Score = 35.8 bits (81), Expect = 0.025
Identities = 24/76 (31%), Positives = 34/76 (44%), Gaps = 6/76 (7%)
Frame = +2
Query: 45 DDDYDERGIEQDE------GQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGD 98
DD D+ G E D+ G + DE +EE E V+ G GD + + D
Sbjct: 41 DDGEDDDGDEDDDDDAPGGGDDDDDEEDEEEEGGVEGGRGGGGDPD-----------DDD 187
Query: 99 DDDVAGDDEHASEENV 114
DDD D+E EE++
Sbjct: 188 DDDDDDDEEEEEEEDL 235
>TC8672 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (26%)
Length = 665
Score = 42.7 bits (99), Expect = 2e-04
Identities = 18/70 (25%), Positives = 37/70 (52%)
Frame = +1
Query: 128 EVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRF 187
EV+ + ++L++FVG L + +L +F + G + + N T +++GF +
Sbjct: 445 EVVASAEPSEDLKIFVGNLPWDVESENLAMLFEEAGSVEFAEVIYNKATNQSRGFGFVIM 624
Query: 188 ETVEHVKRAL 197
T E +++AL
Sbjct: 625 STAEDLEKAL 654
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 42.4 bits (98), Expect = 3e-04
Identities = 44/154 (28%), Positives = 62/154 (39%), Gaps = 28/154 (18%)
Frame = +2
Query: 387 GNKKAKVRARLSRPLPKVRGKHVPHRDISG-RKIGRLERPSRSRPAPRSRPAPQSR---- 441
G+K + R+R+ PLP ++ P R ++G R RPS RP R + P+ R
Sbjct: 17 GSKPQRRRSRVPSPLPTLQ--RWPRRTLNGHRHHPSRHRPSNRRPIQRRQHQPRHRRRRP 190
Query: 442 ------------PAHVFRR--------LGSRIPPA---RPSSARNRRPVTSIPVRARPAS 478
P H+ R L R+PP R + RR + +P RP +
Sbjct: 191 PLPPP*RRQQPPPPHLHPRRRLLHLHPLQQRLPPPPQQRRREGKRRRRLRPLP--PRPGA 364
Query: 479 PPARSYRRLAAAYPKSSMKRDYGRPVDLPPPRSR 512
PP RR ++P RPV P RSR
Sbjct: 365 PPPHRLRRHLGSHPVGCRP---PRPVAQGPRRSR 457
>TC9503 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprotein-like
{Arabidopsis thaliana;}, partial (26%)
Length = 577
Score = 42.4 bits (98), Expect = 3e-04
Identities = 30/117 (25%), Positives = 51/117 (42%)
Frame = +1
Query: 99 DDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKV 158
+D V G E + H D + + + ++F+GGL +E T K
Sbjct: 124 NDHVVGGGESNDVARTYSHREDHDQHHNSQPLTGDGASPG-KIFIGGLARETTIAQFIKH 300
Query: 159 FSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITIT 215
F K GEIT+ + + +T + +GF + + V + + + +INGKQ I T
Sbjct: 301 FGKYGEITDSVIMKDRKTAQPRGFGFITYADPSVVDKVIED--THIINGKQVEIKRT 465
>TC10758
Length = 532
Score = 42.0 bits (97), Expect = 3e-04
Identities = 23/64 (35%), Positives = 36/64 (55%)
Frame = +1
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++FV GL E T LR VFS G++ E + + T R+KG+ + F +H+ A+
Sbjct: 310 KLFVRGLACETTTETLRSVFSAYGDLDEAIVIFDKSTGRSKGYGFVVF---KHIDGAILA 480
Query: 200 LKNP 203
LK+P
Sbjct: 481 LKDP 492
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,128,555
Number of Sequences: 28460
Number of extensions: 175351
Number of successful extensions: 2120
Number of sequences better than 10.0: 272
Number of HSP's better than 10.0 without gapping: 1541
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1834
length of query: 726
length of database: 4,897,600
effective HSP length: 97
effective length of query: 629
effective length of database: 2,136,980
effective search space: 1344160420
effective search space used: 1344160420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149129.7


