

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
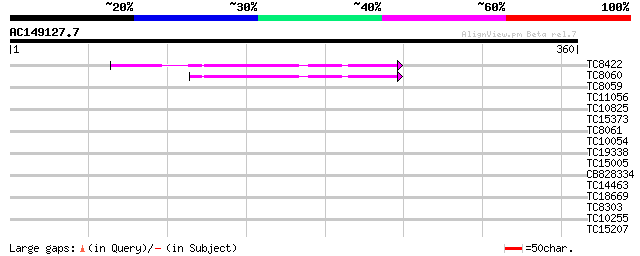
Query= AC149127.7 - phase: 0
(360 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete 42 2e-04
TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regu... 41 3e-04
TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus ann... 37 0.007
TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein ... 30 0.48
TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Ar... 30 0.63
TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 hom... 29 1.4
TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete 29 1.4
TC10054 similar to GB|AAO42896.1|28416731|BT004650 At4g13612 {Ar... 28 2.4
TC19338 similar to GB|AAC49358.1|1388088|PSU35831 thioredoxin m ... 27 4.1
TC15005 homologue to UP|Q9AR92 (Q9AR92) Protein kinase , partia... 27 4.1
CB828334 27 4.1
TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein (At2g... 27 5.3
TC18669 similar to UP|Q93Y39 (Q93Y39) ATP-dependent RNA helicase... 27 6.9
TC8303 homologue to UP|Q8L7F6 (Q8L7F6) Calcineurin B, partial (62%) 27 6.9
TC10255 26 9.1
TC15207 similar to UP|Q9FGP9 (Q9FGP9) Gb|AAF31706.1 (AT5g22790/K... 26 9.1
>TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete
Length = 832
Score = 41.6 bits (96), Expect = 2e-04
Identities = 41/185 (22%), Positives = 75/185 (40%)
Frame = +1
Query: 65 QQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKD 124
++ +N +N ++ T + T +I E E F L D+D D
Sbjct: 82 KERENPIQSNPILSHHTTMADQLTDDQISEFKE----------------AFSLFDKDG-D 210
Query: 125 GFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEKDIEKKNMAH 184
G + EL + + + Q + D +G+ + F E+L ++ K + +
Sbjct: 211 GSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEE 390
Query: 185 GEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKI 244
L E F V D D NG ++ ELR H + +++ V++ D + DG+I
Sbjct: 391 -----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIREADVDGDGQI 546
Query: 245 NFNQF 249
N+ +F
Sbjct: 547 NYEEF 561
>TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regulated
calmodulin) (Calmodulin TACAM1-1) (Calmodulin TACAM1-2)
(Calmodulin TACAM1-3) (Calmodulin TACAM3-1) (Calmodulin
TACAM3-2) (Calmodulin TACAM3-3) (Calmodulin TACAM4-1)
(CaM protein), complete
Length = 615
Score = 41.2 bits (95), Expect = 3e-04
Identities = 34/135 (25%), Positives = 60/135 (44%)
Frame = +3
Query: 115 FPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSE 174
F L D+D DG + EL + + + Q + D +G+ + F E+L ++
Sbjct: 117 FSLFDKDG-DGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 293
Query: 175 KDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
K + + L E F V D D NG ++ ELR H + +++ V++
Sbjct: 294 KMKDTDSEEE-----LKEAFRVFDKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIR 449
Query: 235 MDDYEHDGKINFNQF 249
D + DG+IN+ +F
Sbjct: 450 EADVDGDGQINYEEF 494
>TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus annuus;} ,
partial (83%)
Length = 836
Score = 36.6 bits (83), Expect = 0.007
Identities = 28/106 (26%), Positives = 50/106 (46%), Gaps = 4/106 (3%)
Frame = +2
Query: 148 TQVELESK----DKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNG 203
T+ EL+ D +G+ + F E+L ++ K + + L E F V D D NG
Sbjct: 221 TEAELQDMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEE-----LKEAFRVFDKDQNG 385
Query: 204 LLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQF 249
++ ELR H + +++ V++ D + DG+IN+ +F
Sbjct: 386 FISAAELR---HVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEF 514
>TC11056 similar to UP|O81390 (O81390) Calcium-dependent protein kinase ,
partial (22%)
Length = 675
Score = 30.4 bits (67), Expect = 0.48
Identities = 22/123 (17%), Positives = 58/123 (46%)
Frame = +3
Query: 127 VGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGE 186
+ + EL+S + + + + +E+ D +G+ ++ + E++ + + + E
Sbjct: 6 ITYEELKSGLARIGSRLSETEVKQLMEAADVDGNGSIDYLEFI----SATMHRHRLERDE 173
Query: 187 AGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINF 246
L + F D D++G + EL + ++ +K ++++ D ++DG+IN+
Sbjct: 174 H--LYKAFQYFDKDNSGHITIDELETAMTQHGMGDEASIKEIISEV----DTDNDGRINY 335
Query: 247 NQF 249
+F
Sbjct: 336 EEF 344
>TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Arabidopsis
thaliana;}, partial (32%)
Length = 1091
Score = 30.0 bits (66), Expect = 0.63
Identities = 30/113 (26%), Positives = 49/113 (42%), Gaps = 1/113 (0%)
Frame = +1
Query: 182 MAHGEAGWLMEKFDVADYDHNGLLNFTE-LRDFLHPEDSQNKEMLKWMVNDKFNMDDYEH 240
+A E LME VAD D NG+L++ E + +H + +N E + F D +
Sbjct: 112 LADQEIKMLME---VADVDGNGVLDYGEFVAVTIHLQKMENDEHF----HKAFKFFDKDG 270
Query: 241 DGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFAELDVNKDQFLSPEE 293
G I + ++ + + +G D D E+D +KD +S EE
Sbjct: 271 SGYIESGELQEAL---------ADESGVTDADVLNDIMREVDTDKDGRISYEE 402
>TC15373 weakly similar to UP|Q9STH9 (Q9STH9) Cytochrome P450 homolog,
partial (61%)
Length = 1059
Score = 28.9 bits (63), Expect = 1.4
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = -3
Query: 221 NKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETN 266
N + L W +N FN +HD K+ ++ V+ E Y F+T+
Sbjct: 991 NTDSLDWCMNITFNSIYKDHDWKVEGWGWQPEVFPFLEQYQIFQTH 854
>TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete
Length = 568
Score = 28.9 bits (63), Expect = 1.4
Identities = 15/52 (28%), Positives = 27/52 (51%)
Frame = +2
Query: 198 DYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQF 249
D D NG ++ ELR H + +++ V++ D + DG+IN+ +F
Sbjct: 326 DKDQNGFISAAELR---HVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEF 472
Score = 26.2 bits (56), Expect = 9.1
Identities = 16/62 (25%), Positives = 30/62 (47%)
Frame = +2
Query: 114 LFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLS 173
L LLD+D ++GF+ EL +T + D + D +GD +++ E++ +
Sbjct: 311 LSALLDKD-QNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEFVKVMM 487
Query: 174 EK 175
K
Sbjct: 488 AK 493
>TC10054 similar to GB|AAO42896.1|28416731|BT004650 At4g13612 {Arabidopsis
thaliana;}, partial (77%)
Length = 526
Score = 28.1 bits (61), Expect = 2.4
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = -1
Query: 40 YKILERAPTFDPLVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTT 87
+K+L P D T+++ ++ +N+QHK N ++ P T STT
Sbjct: 418 HKLLSSFP*NDTS-TRLQTQNPNQNKQHKFTISNTMHILPYTFPNSTT 278
>TC19338 similar to GB|AAC49358.1|1388088|PSU35831 thioredoxin m {Pisum
sativum;} , partial (65%)
Length = 476
Score = 27.3 bits (59), Expect = 4.1
Identities = 20/78 (25%), Positives = 37/78 (46%), Gaps = 5/78 (6%)
Frame = +2
Query: 51 PLVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTV-----SEIKETYEYLTSGGTL 105
PL+ ++ +E K +K + D++ N+A + G+ S TV E KE+ TL
Sbjct: 140 PLIDELAKEYAGKIACYKLNTDDSPNIATQYGIRSIPTVLFFKNGEKKESVIGAVPKSTL 319
Query: 106 NTTLRLIILFPLLDRDPK 123
+L I + + +P+
Sbjct: 320 TASLEKYIAI*VENHEPR 373
>TC15005 homologue to UP|Q9AR92 (Q9AR92) Protein kinase , partial (19%)
Length = 657
Score = 27.3 bits (59), Expect = 4.1
Identities = 17/60 (28%), Positives = 32/60 (53%), Gaps = 4/60 (6%)
Frame = +2
Query: 194 FDVADYDHNGLLNFTEL----RDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQF 249
F D D +G + EL +++ +D+ KE++ V+ + D +HDG+IN+ +F
Sbjct: 98 FPYFDKDSSGFITRDELEIAMKEYGMGDDATIKEIIS-EVDTIISEVDTDHDGRINYEEF 274
>CB828334
Length = 541
Score = 27.3 bits (59), Expect = 4.1
Identities = 16/54 (29%), Positives = 29/54 (53%)
Frame = +2
Query: 156 DKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTE 209
DKN D +S E + ++E +++ M++F+ D+D NG++NF E
Sbjct: 350 DKNKDGYVSKSEMVQAINETVSGERSSGR----IAMKRFEEMDWDKNGMVNFKE 499
>TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein
(At2g16660/T24I21.7), partial (40%)
Length = 1122
Score = 26.9 bits (58), Expect = 5.3
Identities = 16/56 (28%), Positives = 31/56 (54%), Gaps = 6/56 (10%)
Frame = -1
Query: 33 LNNRRFGYK------ILERAPTFDPLVTKIERESEQKNQQHKNDFDNNKNVAPRTG 82
LN +RFG+ + R P+F+ V + Q+NQQ+++ +++++ RTG
Sbjct: 972 LNQQRFGHPLPLNIPVRFRFPSFEVRVDRNPHRRSQQNQQNRH--EHSRHQGARTG 811
>TC18669 similar to UP|Q93Y39 (Q93Y39) ATP-dependent RNA helicase
(Fragment), partial (3%)
Length = 461
Score = 26.6 bits (57), Expect = 6.9
Identities = 13/41 (31%), Positives = 20/41 (48%), Gaps = 1/41 (2%)
Frame = -2
Query: 119 DRDPKDGFVGFNELE-SWVTQRALERLDYATQVELESKDKN 158
D DP F+G NELE +++ ++ DY + D N
Sbjct: 295 DNDPFSLFIGSNELEGGFLSLEEIDEADYGMDIHKPGGDDN 173
>TC8303 homologue to UP|Q8L7F6 (Q8L7F6) Calcineurin B, partial (62%)
Length = 784
Score = 26.6 bits (57), Expect = 6.9
Identities = 29/105 (27%), Positives = 44/105 (41%), Gaps = 5/105 (4%)
Frame = +1
Query: 194 FDVADYDHNGLLNFTELRDFL---HPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFE 250
FD+ D HNG+L F E L HP + + ++ F + D + G I + +
Sbjct: 10 FDLFDTKHNGILGFEEFARALSVFHPNAPIDDK-----IHFSFQLYDLKQQGFIERQEVK 174
Query: 251 DNVYVTY-ESYVDFETNGEGDIPTAKDK-FAELDVNKDQFLSPEE 293
V T ES ++ + I + DK F E D D + EE
Sbjct: 175 QMVVATLAESGMNL---SDDVIESIIDKTFEEADTKHDGKIDKEE 300
>TC10255
Length = 539
Score = 26.2 bits (56), Expect = 9.1
Identities = 25/89 (28%), Positives = 40/89 (44%), Gaps = 17/89 (19%)
Frame = -3
Query: 22 SPLNLEESKGRLNNRRFGYKIL--ERAPTFDPLVTKIERESEQKNQ----------QHK- 68
+PL + + KGR+ NR+ KI +R T D L T+ ++ + K + QH
Sbjct: 492 NPLKVGKMKGRVFNRKIE*KITKRKRQETKD*LATEQKKRKQNKRRTGMRQKPKQLQHT* 313
Query: 69 -NDFDN---NKNVAPRTGLGSTTTVSEIK 93
D DN KN+ R + T ++ K
Sbjct: 312 GEDLDNTPRTKNLGERGAKNNLETATKSK 226
>TC15207 similar to UP|Q9FGP9 (Q9FGP9) Gb|AAF31706.1 (AT5g22790/K8E10_2),
partial (55%)
Length = 1195
Score = 26.2 bits (56), Expect = 9.1
Identities = 14/29 (48%), Positives = 16/29 (54%)
Frame = -3
Query: 155 KDKNGDLALSFREYLPDLSEKDIEKKNMA 183
K+K G LA F LSEK + KKN A
Sbjct: 107 KEKKGQLAFDFSA*NNALSEKTVAKKNPA 21
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,886,228
Number of Sequences: 28460
Number of extensions: 54329
Number of successful extensions: 203
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 198
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 203
length of query: 360
length of database: 4,897,600
effective HSP length: 91
effective length of query: 269
effective length of database: 2,307,740
effective search space: 620782060
effective search space used: 620782060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149127.7


