

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
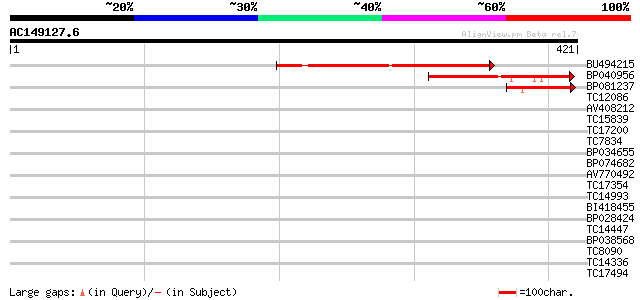
Query= AC149127.6 - phase: 0
(421 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BU494215 185 1e-47
BP040956 105 1e-23
BP081237 63 8e-11
TC12086 33 0.088
AV408212 30 0.75
TC15839 weakly similar to UP|Q8LL12 (Q8LL12) EDS1, partial (26%) 29 1.3
TC17200 similar to UP|Q84VX5 (Q84VX5) At4g02100, partial (27%) 29 1.7
TC7834 homologue to UP|AAS18241 (AAS18241) Cytosolic malate dehy... 29 1.7
BP034655 28 2.8
BP074682 28 3.7
AV770492 28 3.7
TC17354 weakly similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, part... 27 4.8
TC14993 similar to UP|O04237 (O04237) Transcription factor, part... 27 4.8
BI418455 27 4.8
BP028424 27 4.8
TC14447 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (86%) 27 6.3
BP038568 27 6.3
TC8090 homologue to UP|ACTB_ARATH (P53496) Actin 11, complete 27 6.3
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 27 8.3
TC17494 weakly similar to UP|Q940M0 (Q940M0) At1g63980/F22C12_9,... 27 8.3
>BU494215
Length = 491
Score = 185 bits (469), Expect = 1e-47
Identities = 99/162 (61%), Positives = 120/162 (73%)
Frame = +3
Query: 199 KKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTL 258
+KS E++ENCDLP P K + D ++ SSLNQK E SS+++G TS T
Sbjct: 15 QKSMEHIENCDLPNPQTKQ----RASYQKGFDHDKTTPSSLNQKAETGSSNADGYTSGTP 182
Query: 259 TSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQ 318
+SGCSFQDS+ F SS SKDS SS NK+ + NSE + AELL+ALC SQTRAREAEKAAQ
Sbjct: 183 SSGCSFQDSNEHFRSSSSKDSGSS-NKDCQTNSEKRSMAELLEALCHSQTRAREAEKAAQ 359
Query: 319 EACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSK 360
+A +EKEHILSLFF+QASQLFAYKQW H+LQLENLCLQ + +
Sbjct: 360 QAYNEKEHILSLFFRQASQLFAYKQWFHLLQLENLCLQLQKQ 485
>BP040956
Length = 560
Score = 105 bits (263), Expect = 1e-23
Identities = 61/123 (49%), Positives = 78/123 (62%), Gaps = 15/123 (12%)
Frame = -3
Query: 312 EAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFPY 371
EAE+AA++A EK+HI+ L FKQASQLFAYKQW +LQLE LC Q K+K+Q + LFP
Sbjct: 558 EAEEAAKQAFAEKDHIVKLIFKQASQLFAYKQWFQLLQLETLCSQMKNKDQQ-IPTLFPV 382
Query: 372 -------EGKKHRKNRRKVKNSRR---GIGKC-----IFAFAVGLALAGAGLLLGWTIGC 416
EG+ RK ++K N+++ KC A A+G L GAGLLLGWT+G
Sbjct: 381 GLPWLSNEGRNSRKRKQKFCNAKQEALAKPKCDVTTYAVAIALGFGLVGAGLLLGWTVGW 202
Query: 417 MFP 419
M P
Sbjct: 201 MLP 193
>BP081237
Length = 371
Score = 63.2 bits (152), Expect = 8e-11
Identities = 33/56 (58%), Positives = 38/56 (66%), Gaps = 5/56 (8%)
Frame = -3
Query: 370 PYEGKKHRKN-----RRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCMFPS 420
P G H+K+ +RK N R GI CI AFAVGL LAGAGLLLGWT+G MFP+
Sbjct: 366 PSRGTLHKKSPGKGGKRKNCNRRCGIQNCIVAFAVGLGLAGAGLLLGWTLGWMFPT 199
>TC12086
Length = 384
Score = 33.1 bits (74), Expect = 0.088
Identities = 20/59 (33%), Positives = 31/59 (51%)
Frame = -3
Query: 253 CTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAR 311
C S+ +TS SF S + SS E +++DSS + +S SE+ A+ +TR R
Sbjct: 304 CNSSNVTSASSFSISSSSSSSDEIEEADSSVSVSSCWGSETGILLLCF*AMSLRETRRR 128
>AV408212
Length = 362
Score = 30.0 bits (66), Expect = 0.75
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = +1
Query: 246 CSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSE 292
C + + C L+S + ++ SSS S S++S N NSKV E
Sbjct: 49 CRCNRSCCRGMQLSSNSNSNNNSSNSSSSSSNHSNNSHNHNSKVGIE 189
>TC15839 weakly similar to UP|Q8LL12 (Q8LL12) EDS1, partial (26%)
Length = 1161
Score = 29.3 bits (64), Expect = 1.3
Identities = 15/31 (48%), Positives = 17/31 (54%)
Frame = +3
Query: 253 CTSTTLTSGCSFQDSDRTFSSSESKDSDSSC 283
C S TL CS Q RTF S + K S +SC
Sbjct: 954 CRSATLVPHCSAQ*LHRTF*SCQQKHSATSC 1046
>TC17200 similar to UP|Q84VX5 (Q84VX5) At4g02100, partial (27%)
Length = 850
Score = 28.9 bits (63), Expect = 1.7
Identities = 39/159 (24%), Positives = 58/159 (35%), Gaps = 17/159 (10%)
Frame = +2
Query: 165 KSTSSDFEPHWLGAEKSQPWWRTTGK------DDLASLVA-KKSFEYVENCDLPEPNIK- 216
KST H L A + + WW T K D SL+A ++ E +L + +
Sbjct: 245 KSTKMAVTAHSLSATEKKHWWLTNRKIVEKYIKDARSLIATQEQSEIASALNLLDAALAI 424
Query: 217 --------PFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSD 268
R L R + + + L M + D S + +S CS Q+
Sbjct: 425 SPRLDQALELRARSLLYLRRFKEVADMLQDYIPSLRMANEDP---VSVSSSSDCSSQNLS 595
Query: 269 RTFSSSESKDSDSSCNKNS-KVNSESAAKAELLKALCRS 306
R S DS S K S S K +++ LC+S
Sbjct: 596 REGVKLLSSSPDSPVRYQSFKCFSVSDLKKKVMAGLCKS 712
>TC7834 homologue to UP|AAS18241 (AAS18241) Cytosolic malate dehydrogenase ,
complete
Length = 1572
Score = 28.9 bits (63), Expect = 1.7
Identities = 19/81 (23%), Positives = 36/81 (43%), Gaps = 11/81 (13%)
Frame = +1
Query: 15 FTQDFMMPPTSSKQLEPDFTTSDSTDSDMK---WWLHVKTNLAGDTDYTCQH-------- 63
F++D+++ + +FT D+ S ++ WWL+ ++ G+ C H
Sbjct: 1276 FSRDYIIVGHILNWIRTNFTCIDARSSGLELCLWWLYRNLDVCGNDAQACIHCD*VLCI* 1455
Query: 64 LNTLESKLDAFSTRLHGDNVS 84
+ + +AFS H NVS
Sbjct: 1456 IRSSLMLFNAFSQHFHIMNVS 1518
>BP034655
Length = 517
Score = 28.1 bits (61), Expect = 2.8
Identities = 16/46 (34%), Positives = 26/46 (55%)
Frame = -3
Query: 250 SNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
S+ +S++ +S S S + SSS S S SS + +S + +SAA
Sbjct: 251 SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSESPKSAA 114
Score = 27.7 bits (60), Expect = 3.7
Identities = 16/44 (36%), Positives = 24/44 (54%)
Frame = -3
Query: 254 TSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKA 297
+S++ +S S S + SSS S S SS + +S +SES A
Sbjct: 248 SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSESPKSA 117
>BP074682
Length = 497
Score = 27.7 bits (60), Expect = 3.7
Identities = 27/79 (34%), Positives = 37/79 (46%)
Frame = +1
Query: 214 NIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSS 273
++ PF ++ RET EE V D+ TS++ +SG SD SS
Sbjct: 43 SLAPFVEVQ----RETRAEERNVPPSLPVHGEIQVDNGSKTSSSSSSG-----SDSGSSS 195
Query: 274 SESKDSDSSCNKNSKVNSE 292
S+S DSDSS S V S+
Sbjct: 196 SDS-DSDSSSASGSDVGSQ 249
>AV770492
Length = 479
Score = 27.7 bits (60), Expect = 3.7
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = -2
Query: 382 KVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCMFPS 420
K + RRG+G F+F + + GLLLG+ + +F S
Sbjct: 319 KRQRHRRGVGDPGFSFTFAVFVGLLGLLLGFLLKLLFSS 203
>TC17354 weakly similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, partial (24%)
Length = 638
Score = 27.3 bits (59), Expect = 4.8
Identities = 17/64 (26%), Positives = 27/64 (41%)
Frame = +3
Query: 228 ETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNS 287
E D+ N +L ++ D TS++ + S + + S S S SDS +
Sbjct: 144 EMDRGTNAAEALRAQVNAEGRDEQTSTSSSSSGSGSSESGSGSGSRSSSSSSDSEGSDED 323
Query: 288 KVNS 291
VNS
Sbjct: 324 TVNS 335
>TC14993 similar to UP|O04237 (O04237) Transcription factor, partial (31%)
Length = 846
Score = 27.3 bits (59), Expect = 4.8
Identities = 26/66 (39%), Positives = 32/66 (48%), Gaps = 4/66 (6%)
Frame = -2
Query: 221 IHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCS----FQDSDRTFSSSES 276
I +L+ +D ++SSL+ E DSN ST LTS S F S TFS S S
Sbjct: 287 IASLRLSLSDPNLAKISSLSFLWESGCMDSNSLASTFLTSVLSACRAFLFSSSTFSYSSS 108
Query: 277 KDSDSS 282
S SS
Sbjct: 107 VISLSS 90
>BI418455
Length = 477
Score = 27.3 bits (59), Expect = 4.8
Identities = 13/47 (27%), Positives = 27/47 (56%), Gaps = 4/47 (8%)
Frame = -3
Query: 345 LHVLQLENLCLQYKSKNQPLLNNLFPY----EGKKHRKNRRKVKNSR 387
L + Q++ LC ++S+ +L + FP+ EG HR+ ++++ R
Sbjct: 184 LQLSQVQGLCRIFRSQIHHILGH*FPHRNLVEGSGHRQRQKRMPKPR 44
>BP028424
Length = 206
Score = 27.3 bits (59), Expect = 4.8
Identities = 11/22 (50%), Positives = 17/22 (77%)
Frame = +2
Query: 333 KQASQLFAYKQWLHVLQLENLC 354
+++S LF+Y+ LHV+QL LC
Sbjct: 95 EKSSPLFSYEPSLHVIQLYRLC 160
>TC14447 homologue to UP|Q9M531 (Q9M531) Histone H2A, partial (86%)
Length = 755
Score = 26.9 bits (58), Expect = 6.3
Identities = 16/50 (32%), Positives = 24/50 (48%)
Frame = -2
Query: 211 PEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTS 260
P+PN+ R+ HTL R N +SSL ++ + N S + TS
Sbjct: 538 PKPNLSFLRRFHTLLSRRLRLLGNLLSSLLRQKHRVNIRKNTAMSNSHTS 389
>BP038568
Length = 553
Score = 26.9 bits (58), Expect = 6.3
Identities = 26/66 (39%), Positives = 32/66 (48%), Gaps = 4/66 (6%)
Frame = -3
Query: 221 IHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCS----FQDSDRTFSSSES 276
I +L+ +D ++SSL+ E DSN ST LTS S F S TFS S S
Sbjct: 317 IASLRLSLSDPNLAKISSLSFLWESGCMDSNSFASTFLTSVLSACRAFLFSSSTFSYSSS 138
Query: 277 KDSDSS 282
S SS
Sbjct: 137 VISLSS 120
>TC8090 homologue to UP|ACTB_ARATH (P53496) Actin 11, complete
Length = 1624
Score = 26.9 bits (58), Expect = 6.3
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = +1
Query: 160 ISEHCKSTSSDFEPHWLGAEKSQPW 184
+S+HC+S+SS + W G E+ W
Sbjct: 193 VSQHCRSSSSHWCDGWHGPERCICW 267
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 26.6 bits (57), Expect = 8.3
Identities = 18/44 (40%), Positives = 22/44 (49%)
Frame = -1
Query: 205 VENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSS 248
VEN P+P + P + TLQ EEN SSL+ M SS
Sbjct: 822 VENATNPDPRLMPLGSLITLQEVMVP*EEN--SSLSFSSAMLSS 697
>TC17494 weakly similar to UP|Q940M0 (Q940M0) At1g63980/F22C12_9, partial
(18%)
Length = 634
Score = 26.6 bits (57), Expect = 8.3
Identities = 13/51 (25%), Positives = 28/51 (54%), Gaps = 5/51 (9%)
Frame = +2
Query: 260 SGCSFQDSDRTFSSSESKDSDSSC----NKNSKVNSESAAKAEL-LKALCR 305
+GC F+ +F +S+ +DSD+ ++ + E ++++ LK LC+
Sbjct: 11 AGCYFEGKKTSFDNSDEEDSDTDSMEKQTRDDLIKVEKVVESKVKLKKLCK 163
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.127 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,129,072
Number of Sequences: 28460
Number of extensions: 113845
Number of successful extensions: 685
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 671
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 681
length of query: 421
length of database: 4,897,600
effective HSP length: 93
effective length of query: 328
effective length of database: 2,250,820
effective search space: 738268960
effective search space used: 738268960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149127.6


