

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149080.7 - phase: 0
(165 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
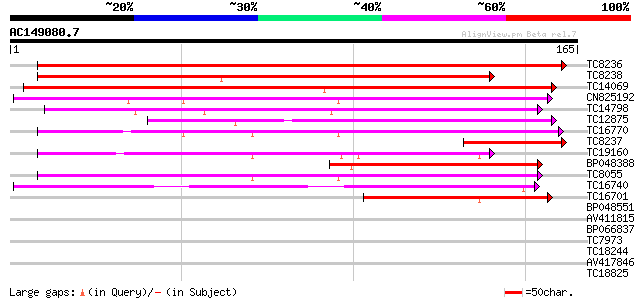
Score E
Sequences producing significant alignments: (bits) Value
TC8236 270 7e-74
TC8238 223 2e-59
TC14069 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fr... 125 3e-30
CN825192 104 7e-24
TC14798 similar to UP|Q84TF6 (Q84TF6) At1g09740, partial (85%) 104 9e-24
TC12875 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fr... 85 6e-18
TC16770 weakly similar to GB|AAL49942.1|17979251|AY070476 AT3g53... 70 2e-13
TC8237 62 4e-11
TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%) 62 5e-11
BP048388 60 2e-10
TC8055 similar to UP|Q9FES5 (Q9FES5) Early nodulin ENOD18, parti... 60 2e-10
TC16740 similar to UP|Q9SIJ8 (Q9SIJ8) Expressed protein (RD2 pro... 56 3e-09
TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome ... 53 3e-08
BP048551 32 0.055
AV411815 30 0.27
BP066837 27 1.4
TC7973 similar to AAQ56804 (AAQ56804) At1g63770, partial (19%) 27 1.8
TC18244 weakly similar to UP|Q9LSP5 (Q9LSP5) Gb|AAF26101.1 (AT3g... 26 3.0
AV417846 25 5.2
TC18825 similar to UP|BAD08451 (BAD08451) Glucose-6-phosphate is... 25 5.2
>TC8236
Length = 840
Score = 270 bits (691), Expect = 7e-74
Identities = 129/154 (83%), Positives = 143/154 (92%)
Frame = +3
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
RRIMV +DEG+ES+YAL+WCLKNL FQN KDHLILLYVKPPRVVYSAFDGTGYLFSSDIT
Sbjct: 141 RRIMVTVDEGDESMYALSWCLKNLAFQNDKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 320
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHGY 128
ATME+ SQQVA+ VLE+AK +CN+V+NVE + E+GDPRDVICQ VQK GVD+LVMGSHGY
Sbjct: 321 ATMERVSQQVAEGVLERAKGLCNNVENVEVKAESGDPRDVICQMVQKWGVDVLVMGSHGY 500
Query: 129 GVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTG 162
GVIKRAFLGSVSNHCAQNVKCPV+IVKKPKST G
Sbjct: 501 GVIKRAFLGSVSNHCAQNVKCPVVIVKKPKSTAG 602
>TC8238
Length = 506
Score = 223 bits (567), Expect = 2e-59
Identities = 110/139 (79%), Positives = 123/139 (88%), Gaps = 6/139 (4%)
Frame = +1
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTG------YL 62
RRIMV +DEG+ES+YAL+WCLKNL FQN KDHLILLYVKPPRVVYSAFDGTG YL
Sbjct: 88 RRIMVTVDEGDESMYALSWCLKNLAFQNDKDHLILLYVKPPRVVYSAFDGTGSFVVTRYL 267
Query: 63 FSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILV 122
FSSDITATME+ SQQVA+ VLE+AK +CN+V+NVE + E+GDPRDVICQ VQK GVD+LV
Sbjct: 268 FSSDITATMERVSQQVAEGVLERAKGLCNNVENVEVKAESGDPRDVICQMVQKWGVDVLV 447
Query: 123 MGSHGYGVIKRAFLGSVSN 141
MGSHGYGVIKRAFLGSVSN
Sbjct: 448 MGSHGYGVIKRAFLGSVSN 504
>TC14069 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fragment),
partial (58%)
Length = 824
Score = 125 bits (315), Expect = 3e-30
Identities = 60/157 (38%), Positives = 99/157 (62%), Gaps = 2/157 (1%)
Frame = +3
Query: 5 TENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFS 64
+E + +++ ID+ E S+YA+ W L + +N L+L++ +P F G Y +
Sbjct: 138 SEKKQVMVIGIDDSEHSVYAINWTLDHFFAKNPSFKLVLVHARPSATSAVGFAGPVYAGA 317
Query: 65 SDITATMEKYSQQVADCVLEKAKIVC--NDVQNVETRIENGDPRDVICQAVQKMGVDILV 122
+++ ++ +++A VLE AK +C N++ +V GDPR+V+C+AV+K +LV
Sbjct: 318 AEVLPIVDSDLKKIAARVLENAKQICIKNNITDVVVEAVEGDPRNVLCEAVEKYHASVLV 497
Query: 123 MGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
+GSHGYG +KRA LGSVS++CA + C V+IVKKPK+
Sbjct: 498 VGSHGYGALKRAVLGSVSDYCAHHAHCSVMIVKKPKT 608
>CN825192
Length = 697
Score = 104 bits (260), Expect = 7e-24
Identities = 62/172 (36%), Positives = 95/172 (55%), Gaps = 15/172 (8%)
Frame = +3
Query: 2 AGITENGRRIMVAIDEGEESIYALTWCLKNLV------------FQNSKDHLILLYVKPP 49
AG ++MVA+DE + S +AL W L N++ ++ + L++V+P
Sbjct: 54 AGERRMKMKVMVAVDESDCSFHALKWALDNVLNNMTTTATPDENIEDGGGMVFLVHVEPA 233
Query: 50 --RVVYSAFDGTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPR 106
VY Y S+ + M K ++ + L +A +C D Q E+ I GD R
Sbjct: 234 FHPAVYPIGTSALYPASASLEDLMRKAQREKSTSTLSRALQMCRDNQIKAESIILTGDAR 413
Query: 107 DVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
++ICQA +M VD+L+MGS G G++KR FLGSVS++CA + K P+LIVK P+
Sbjct: 414 EMICQAADQMHVDLLIMGSRGLGMLKRTFLGSVSDYCAHHAKTPILIVKPPE 569
>TC14798 similar to UP|Q84TF6 (Q84TF6) At1g09740, partial (85%)
Length = 1309
Score = 104 bits (259), Expect = 9e-24
Identities = 56/161 (34%), Positives = 91/161 (55%), Gaps = 16/161 (9%)
Frame = +3
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQ------NSKDHLILLYVKPPRVVYSA-------FD 57
+ VA+D +ES+ AL W + NL + + ++L+V+ P + + F
Sbjct: 282 VAVAVDGSDESMNALRWAMTNLKLRPQTPDSTTAGCFLILHVQSPPSIATGLNPGPIPFG 461
Query: 58 GTGYLFSSDITATMEKYSQQVADCVLEKAKIVCND---VQNVETRIENGDPRDVICQAVQ 114
G L A +E + +++ D +L+ A +C++ + V T + GDP++ IC+AVQ
Sbjct: 462 GPSNLEVPAFAAAIEAHQKRITDSILDHALGICSEFNFTEKVRTHVVIGDPKEKICEAVQ 641
Query: 115 KMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
D+LVMGS +G IKR FLGSVSN+CA + +CPV+I+K
Sbjct: 642 DQHADVLVMGSRAFGPIKRMFLGSVSNYCAHHAECPVIIIK 764
>TC12875 similar to UP|AAR07598 (AAR07598) Fiber protein Fb19 (Fragment),
partial (64%)
Length = 668
Score = 85.1 bits (209), Expect = 6e-18
Identities = 49/121 (40%), Positives = 70/121 (57%), Gaps = 2/121 (1%)
Frame = -3
Query: 41 LILLYVKPPRVVYSAFDGTGYLFS--SDITATMEKYSQQVADCVLEKAKIVCNDVQNVET 98
++L+Y KP S L S + A +++ + +V D + K V V +V
Sbjct: 582 IVLVYAKPSVATTSWLCWPWNLVSVIPVVDADLKRTAAKVTD--IAKELCVQKSVTDVTV 409
Query: 99 RIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
+ GD R+V+C V+K ILV+GSHGYG IKRA LGSVS++CA + C V+IVKKPK
Sbjct: 408 EVIEGDARNVLCDVVEKHHASILVVGSHGYGAIKRAVLGSVSDYCAHHAHCTVMIVKKPK 229
Query: 159 S 159
+
Sbjct: 228 T 226
>TC16770 weakly similar to GB|AAL49942.1|17979251|AY070476
AT3g53990/F5K20_290 {Arabidopsis thaliana;}, partial
(97%)
Length = 781
Score = 70.1 bits (170), Expect = 2e-13
Identities = 52/161 (32%), Positives = 77/161 (47%), Gaps = 8/161 (4%)
Frame = +2
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPP-----RVVYSAFDGTGYLF 63
R I VA+D + S AL W + NL+ + D L ++++ PP R + + G+ +
Sbjct: 71 RNIGVALDFSKGSKTALNWAVDNLL--RNGDTLYIIHINPPQDSESRNLLWSTTGSPLIP 244
Query: 64 SSDITA--TMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDI 120
S+ M Y VL+ Q + ++ GD R+ I AV+ + +D
Sbjct: 245 LSEFREREVMRHYEVDTDAEVLDLLDTASRQKQVTIVAKLYWGDAREKIVDAVEDLKLDA 424
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
LVMGS G G I+R LGSVS + N CPV IVK +T
Sbjct: 425 LVMGSRGLGAIQRVLLGSVSTYVTSNANCPVTIVKDSVPST 547
>TC8237
Length = 929
Score = 62.4 bits (150), Expect = 4e-11
Identities = 28/30 (93%), Positives = 29/30 (96%)
Frame = +3
Query: 133 RAFLGSVSNHCAQNVKCPVLIVKKPKSTTG 162
RAFLGSVSNHCAQNVKCPV+IVKKPKST G
Sbjct: 597 RAFLGSVSNHCAQNVKCPVVIVKKPKSTAG 686
>TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%)
Length = 557
Score = 62.0 bits (149), Expect = 5e-11
Identities = 45/147 (30%), Positives = 79/147 (53%), Gaps = 14/147 (9%)
Frame = +1
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R+I +A+D +ES YA+ W ++N + D +ILL+V+P V+Y A G G + D
Sbjct: 121 RKIAIAVDLSDESAYAVRWAVQN--YLRPGDEVILLHVRPTSVLYGADWGAGDHNAEDGD 294
Query: 69 A--TMEKYSQQVADCVLEKAKIVCNDVQN--VETRI-------ENGDPRDVICQAVQKMG 117
+ E+ +++ D +D+ VE +I ++ D ++ +C V+++G
Sbjct: 295 GGDSSEESRRKLEDDFDNFTSTKASDLAEPLVEEQIPFKIHIVKDHDMKERLCLEVERLG 474
Query: 118 VDILVMGSHGYGVIKRAF---LGSVSN 141
+ ++MGS G+G KRA LGSVS+
Sbjct: 475 LSAVIMGSRGFGASKRAAKGRLGSVSD 555
>BP048388
Length = 449
Score = 60.1 bits (144), Expect = 2e-10
Identities = 26/64 (40%), Positives = 43/64 (66%), Gaps = 2/64 (3%)
Frame = -3
Query: 94 QNVET--RIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPV 151
+N+E +I GD R+ + +A++ + +D ++MG+ G G ++RA +GSVSNH N CPV
Sbjct: 384 KNIEVLLKIYWGDAREKLLEAIEHIPLDSIIMGNRGLGTLRRAIMGSVSNHVVNNASCPV 205
Query: 152 LIVK 155
+VK
Sbjct: 204 TVVK 193
>TC8055 similar to UP|Q9FES5 (Q9FES5) Early nodulin ENOD18, partial (89%)
Length = 625
Score = 59.7 bits (143), Expect = 2e-10
Identities = 43/150 (28%), Positives = 70/150 (46%), Gaps = 3/150 (2%)
Frame = +1
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R++ VA D + S AL W ++N+ + ++I + R A G+ + S +
Sbjct: 34 RKVGVATDFSKSSNSALKWAIENMADKGDTFYIIHVMSDGSRTNIWAKSGSPLIPLSILR 213
Query: 69 A--TMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDILVMGS 125
M Y Q VL+ + N ++ G+ R + +++ + +D LVMGS
Sbjct: 214 QPEAMSNYGVQTDPEVLDMLDAAAGQKEVNFVAKLYWGEARQKLIDSIEDLKLDSLVMGS 393
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
G G IKR +GSVSN + CPV IV+
Sbjct: 394 RGRGSIKRILMGSVSNFLMIHATCPVAIVR 483
>TC16740 similar to UP|Q9SIJ8 (Q9SIJ8) Expressed protein (RD2 protein),
partial (90%)
Length = 765
Score = 56.2 bits (134), Expect = 3e-09
Identities = 44/154 (28%), Positives = 64/154 (40%), Gaps = 1/154 (0%)
Frame = +1
Query: 2 AGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGY 61
+G GR I++AID G S +A W L +L HLI ++ D
Sbjct: 157 SGERRRGRDIVLAIDHGPNSKHAFDWALIHLCRLADTIHLI----------HAVSDVKNQ 306
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDIL 121
L MEK + + + + K RI GD VIC +++ +
Sbjct: 307 LVYDTTQGLMEKLAVEAFEVAMVKTV----------ARIVEGDAGKVICNEAERIKPAAV 456
Query: 122 VMGSHGYGVIKRAFLGSVSNHCAQNVK-CPVLIV 154
VMG+ G +I+ GSV +C N K PV+IV
Sbjct: 457 VMGTRGRSLIQSVLQGSVGEYCVHNCKSAPVVIV 558
>TC16701 homologue to UP|Q9LSR2 (Q9LSR2) Genomic DNA, chromosome 5, BAC
clone:F24B18, partial (24%)
Length = 691
Score = 52.8 bits (125), Expect = 3e-08
Identities = 21/58 (36%), Positives = 40/58 (68%), Gaps = 3/58 (5%)
Frame = +1
Query: 104 DPRDVICQAVQKMGVDILVMGSHGYGVIKRAF---LGSVSNHCAQNVKCPVLIVKKPK 158
D ++ +C V+++G+ ++MGS G+G ++R LGSVS++C + CPV++V+ P+
Sbjct: 4 DMKERLCLEVERLGLSAVIMGSRGFGAVRRGSDGKLGSVSDYCVHHCVCPVVVVRYPE 177
>BP048551
Length = 540
Score = 32.0 bits (71), Expect = 0.055
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = -1
Query: 2 AGITENGRRIMVAIDEGEESIYALTWCLKNLV 33
AG ++MVAIDE E S +AL W L N++
Sbjct: 468 AGERRMKMKVMVAIDESELSFHALKWALDNVL 373
>AV411815
Length = 423
Score = 29.6 bits (65), Expect = 0.27
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +2
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPP 49
R + +A+D S AL W + NL+ N D +I++ V+PP
Sbjct: 152 RIVGIAMDFSPTSKLALRWAVDNLI--NRGDQIIIINVEPP 268
>BP066837
Length = 477
Score = 27.3 bits (59), Expect = 1.4
Identities = 16/62 (25%), Positives = 29/62 (45%)
Frame = +3
Query: 25 LTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDITATMEKYSQQVADCVLE 84
L W +K +F + + I ++ +PP V+ +F T SD + + S V C+
Sbjct: 216 LIWTIKLNMFISGCKYGICMHARPPTTVHESFHVT-----SDCMIMLHQESTDVMSCLSS 380
Query: 85 KA 86
+A
Sbjct: 381 QA 386
>TC7973 similar to AAQ56804 (AAQ56804) At1g63770, partial (19%)
Length = 858
Score = 26.9 bits (58), Expect = 1.8
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = +1
Query: 51 VVYSAFDGTGYLFSSDITATMEKYSQQVA 79
V + A DG+GY F +I ++K + QVA
Sbjct: 388 VNFHAKDGSGYTFLGEIVLQLDKINPQVA 474
>TC18244 weakly similar to UP|Q9LSP5 (Q9LSP5) Gb|AAF26101.1
(AT3g17020/K14A17_14), partial (26%)
Length = 451
Score = 26.2 bits (56), Expect = 3.0
Identities = 17/64 (26%), Positives = 31/64 (47%), Gaps = 2/64 (3%)
Frame = +3
Query: 94 QNVET--RIENGDPRDVICQAVQKMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPV 151
+N+E +I GD R+ + +A++ + +D ++MG + G NH N P
Sbjct: 183 KNIEVLLKIYWGDAREKLLEAIEHIPLDSIIMGEXRPWHSHKGHXGXCHNHVV-NTLLPR 359
Query: 152 LIVK 155
+VK
Sbjct: 360 PVVK 371
>AV417846
Length = 429
Score = 25.4 bits (54), Expect = 5.2
Identities = 19/75 (25%), Positives = 34/75 (45%)
Frame = +1
Query: 26 TWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDITATMEKYSQQVADCVLEK 85
+W L+ + + L L+ P+V+Y F + L + KYS +++D L +
Sbjct: 64 SWSLRLKIALGAAKGLAFLHSTEPKVIYRDFKTSNILLDT-------KYSAKLSDFGLAR 222
Query: 86 AKIVCNDVQNVETRI 100
V D +V TR+
Sbjct: 223 DGPV-GDKSHVSTRV 264
>TC18825 similar to UP|BAD08451 (BAD08451) Glucose-6-phosphate isomerase,
partial (39%)
Length = 773
Score = 25.4 bits (54), Expect = 5.2
Identities = 13/29 (44%), Positives = 18/29 (61%)
Frame = -3
Query: 42 ILLYVKPPRVVYSAFDGTGYLFSSDITAT 70
ILL+ +P ++ S F GYL S +IT T
Sbjct: 177 ILLHERPSVLLISHFWSQGYLHSLEITIT 91
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,561,132
Number of Sequences: 28460
Number of extensions: 28232
Number of successful extensions: 161
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 154
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 154
length of query: 165
length of database: 4,897,600
effective HSP length: 83
effective length of query: 82
effective length of database: 2,535,420
effective search space: 207904440
effective search space used: 207904440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC149080.7


