

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149050.9 - phase: 0 /pseudo
(157 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
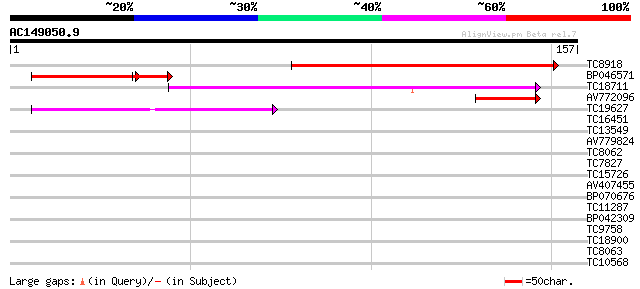
Score E
Sequences producing significant alignments: (bits) Value
TC8918 similar to UP|Q93Z42 (Q93Z42) AT5g19750/T29J13_170, parti... 133 2e-32
BP046571 49 1e-08
TC18711 41 1e-04
AV772096 39 5e-04
TC19627 similar to UP|Q9ZS51 (Q9ZS51) Peroxisomal membrane prote... 38 7e-04
TC16451 similar to UP|AAR20751 (AAR20751) At3g24570, partial (51%) 35 0.006
TC13549 similar to UP|Q9STG6 (Q9STG6) dUTP pyrophosphatase-like ... 28 0.72
AV779824 27 1.2
TC8062 UP|BAA31218 (BAA31218) Histone H3, complete 27 1.6
TC7827 PIR|A31886|A31886 H+-exporting ATPase 57K chain - Arabid... 26 2.7
TC15726 similar to UP|Q9FLB7 (Q9FLB7) Ubiquinol-cytochrome C red... 26 3.6
AV407455 25 4.7
BP070676 25 4.7
TC11287 similar to GB|AAO63429.1|28951011|BT005365 At4g25620 {Ar... 25 4.7
BP042309 25 6.1
TC9758 similar to UP|Q93VQ8 (Q93VQ8) AT4g08170/T12G13_10, partia... 25 6.1
TC18900 similar to UP|BAC83916 (BAC83916) Methylmalonate semi-al... 25 8.0
TC8063 UP|BAA31218 (BAA31218) Histone H3, complete 25 8.0
TC10568 similar to UP|Q84MM3 (Q84MM3) Lethal leaf spot 1-like pr... 25 8.0
>TC8918 similar to UP|Q93Z42 (Q93Z42) AT5g19750/T29J13_170, partial (33%)
Length = 614
Score = 133 bits (334), Expect = 2e-32
Identities = 62/74 (83%), Positives = 68/74 (91%)
Frame = +1
Query: 79 GTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFSAVLANWQLWIPF 138
G SGAILRLV+DQF+FSPIF+GVFL+SLVTLEG PS VPKLKQEWFS+VLANW+LWIPF
Sbjct: 1 GASGAILRLVIDQFIFSPIFIGVFLASLVTLEGSPSLVVPKLKQEWFSSVLANWKLWIPF 180
Query: 139 QFLNFRFVPQQFQV 152
FLNFRFVPQ FQV
Sbjct: 181 PFLNFRFVPQPFQV 222
>BP046571
Length = 490
Score = 49.3 bits (116), Expect(2) = 1e-08
Identities = 23/30 (76%), Positives = 27/30 (89%)
Frame = +3
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQL 36
Y+ALLAK+PV VKALTSA+L LIGDL+C L
Sbjct: 375 YLALLAKHPVLVKALTSALLTLIGDLVCSL 464
Score = 24.3 bits (51), Expect(2) = 1e-08
Identities = 10/11 (90%), Positives = 11/11 (99%)
Frame = +2
Query: 35 QLVIDKVQTPD 45
QLVIDKVQ+PD
Sbjct: 458 QLVIDKVQSPD 490
>TC18711
Length = 711
Score = 40.8 bits (94), Expect = 1e-04
Identities = 24/104 (23%), Positives = 46/104 (44%), Gaps = 1/104 (0%)
Frame = +1
Query: 45 DLKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLS 104
D R F +G L G H++Y + +L G +++V DQ +S ++ ++ +
Sbjct: 25 DRARMFRSGLVGFTLHGSLSHYYYQFCEELFPYKGWWVVPVKVVFDQTAWSAVWNSIYYT 204
Query: 105 SLVTLE-GRPSQAVPKLKQEWFSAVLANWQLWIPFQFLNFRFVP 147
+ L P +L+ +F + A W+LW + + VP
Sbjct: 205 VVGFLRFDSPINIFNELRATFFPMLTAGWKLWPFAHLITYGVVP 336
>AV772096
Length = 431
Score = 38.5 bits (88), Expect = 5e-04
Identities = 15/18 (83%), Positives = 16/18 (88%)
Frame = -2
Query: 130 ANWQLWIPFQFLNFRFVP 147
ANW+LWIPF FL FRFVP
Sbjct: 430 ANWKLWIPFPFLIFRFVP 377
>TC19627 similar to UP|Q9ZS51 (Q9ZS51) Peroxisomal membrane protein
(Peroxisomal membrane protein PMP22) (At4g04470),
partial (36%)
Length = 549
Score = 38.1 bits (87), Expect = 7e-04
Identities = 19/68 (27%), Positives = 37/68 (53%)
Frame = +3
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
Y+ L ++P+ K +T+ +L+ I D++ Q + +Q KR L G +GP HF
Sbjct: 189 YVKQLQQHPLRTKVITAGVLSGISDIVSQKLTG-IQRLQFKRLLLKVIFGAAYLGPFGHF 365
Query: 67 WYLYLSQL 74
+++ L ++
Sbjct: 366 FHIMLDKI 389
>TC16451 similar to UP|AAR20751 (AAR20751) At3g24570, partial (51%)
Length = 654
Score = 35.0 bits (79), Expect = 0.006
Identities = 17/64 (26%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Frame = +2
Query: 90 DQFVFSPIFLGVFLSSLVTLEGRPSQAVPK-LKQEWFSAVLANWQLWIPFQFLNFRFVPQ 148
D F+F P+ L VF + + G+ + + +K+++ A++ +W Q NFR++P
Sbjct: 86 DGFLFGPLDLLVFFTYMGFSTGKSVPQIKEDVKRDFLPALILEGGIWPVVQVANFRYIPV 265
Query: 149 QFQV 152
++Q+
Sbjct: 266 RYQL 277
>TC13549 similar to UP|Q9STG6 (Q9STG6) dUTP pyrophosphatase-like protein ,
partial (76%)
Length = 597
Score = 28.1 bits (61), Expect = 0.72
Identities = 11/18 (61%), Positives = 15/18 (83%)
Frame = +3
Query: 29 IGDLICQLVIDKVQTPDL 46
IGD I QL+I+K+ TPD+
Sbjct: 267 IGDRIAQLIIEKIVTPDV 320
>AV779824
Length = 506
Score = 27.3 bits (59), Expect = 1.2
Identities = 14/45 (31%), Positives = 22/45 (48%)
Frame = -3
Query: 28 LIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHFWYLYLS 72
L+ LIC ++ + P + S + LVL+GP FW + S
Sbjct: 351 LVKPLICYCIVQPLFCP-----CIISMVSLVLLGPRGEFWIIIFS 232
>TC8062 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 768
Score = 26.9 bits (58), Expect = 1.6
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 10/67 (14%)
Frame = -3
Query: 46 LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPG----------TSGAILRLVLDQFVFS 95
+ RT L+ G L+ ++ FWY +S+ T+PG GA LR L +
Sbjct: 274 ISRTSLWK--GSFLISSSVLFWYFLISRRATVPGR*R*GFFTPPVVGADLRAAL----VA 113
Query: 96 PIFLGVF 102
FLG F
Sbjct: 112 SCFLGAF 92
>TC7827 PIR|A31886|A31886 H+-exporting ATPase 57K chain - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial
(29%)
Length = 519
Score = 26.2 bits (56), Expect = 2.7
Identities = 22/87 (25%), Positives = 39/87 (44%), Gaps = 22/87 (25%)
Frame = +3
Query: 1 MTLYYRYMALLAKYPVPVKALTSAI------LNLIGDL-------------ICQLVIDKV 41
+ L+ RY+ L KYPV V L + I +N++ +L CQ++I ++
Sbjct: 246 LMLFVRYLLLEKKYPVDVGILVTCIQIWQQYMNVLEELRGEKALLHRFQF*PCQMMILRI 425
Query: 42 QTPDLKRTFL---FSFLGLVLVGPTLH 65
Q L+ L ++ G + G +H
Sbjct: 426 QLLILRDILLRGRYTLTGSCITGRYIH 506
>TC15726 similar to UP|Q9FLB7 (Q9FLB7) Ubiquinol-cytochrome C reductase
complex ubiquinone-binding protein, complete
Length = 564
Score = 25.8 bits (55), Expect = 3.6
Identities = 11/18 (61%), Positives = 14/18 (77%)
Frame = +3
Query: 55 LGLVLVGPTLHFWYLYLS 72
LGL+L+ LHF YLY+S
Sbjct: 456 LGLILLIVCLHFTYLYIS 509
>AV407455
Length = 399
Score = 25.4 bits (54), Expect = 4.7
Identities = 12/43 (27%), Positives = 20/43 (45%)
Frame = +2
Query: 94 FSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFSAVLANWQLWI 136
F P L LS L + P++ L+ ++ +WQLW+
Sbjct: 68 FHPFKLHSVLSPLPQIPSTPTERHLNLRLQFLPGGKTHWQLWL 196
>BP070676
Length = 420
Score = 25.4 bits (54), Expect = 4.7
Identities = 10/33 (30%), Positives = 19/33 (57%)
Frame = -3
Query: 105 SLVTLEGRPSQAVPKLKQEWFSAVLANWQLWIP 137
+ + + +Q + KLKQ WF +L++W +P
Sbjct: 238 AFICIHTTSTQQIQKLKQCWF*LLLSSWIHTLP 140
>TC11287 similar to GB|AAO63429.1|28951011|BT005365 At4g25620 {Arabidopsis
thaliana;}, partial (44%)
Length = 839
Score = 25.4 bits (54), Expect = 4.7
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = -1
Query: 23 SAILNLIGDLICQLVIDKVQTPDLKRTF 50
S ILNL+ +L ++ K+Q P L F
Sbjct: 797 SKILNLVLNLALHFLVSKIQIPSLNSIF 714
>BP042309
Length = 447
Score = 25.0 bits (53), Expect = 6.1
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = +1
Query: 78 PGTSGAILRLVLDQFVFSP 96
PGT G IL+L D+FVF P
Sbjct: 262 PGTPG-ILKLTQDKFVFKP 315
>TC9758 similar to UP|Q93VQ8 (Q93VQ8) AT4g08170/T12G13_10, partial (35%)
Length = 857
Score = 25.0 bits (53), Expect = 6.1
Identities = 24/70 (34%), Positives = 32/70 (45%), Gaps = 4/70 (5%)
Frame = -1
Query: 31 DLICQLVIDKVQTPDLKRTFLFSFLGLVL-VGPTLHFWYLYLSQLVTLPG---TSGAILR 86
D+ CQLV DKV LK F F L+L + W+ +S + T IL
Sbjct: 710 DIPCQLVADKV----LKDIMFFKFQDLLLSESIMMSIWFNKISFVFTFGCR**LQNIILF 543
Query: 87 LVLDQFVFSP 96
LV + +FSP
Sbjct: 542 LVRNIKMFSP 513
>TC18900 similar to UP|BAC83916 (BAC83916) Methylmalonate semi-aldehyde
dehydrogenase, partial (42%)
Length = 809
Score = 24.6 bits (52), Expect = 8.0
Identities = 18/40 (45%), Positives = 21/40 (52%)
Frame = +2
Query: 69 LYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVT 108
LYL LV L G+ ILRL +F FL +F SL T
Sbjct: 467 LYLLHLVLLQGSFRLILRLGRLASMFQFQFLCLFSHSLET 586
>TC8063 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 723
Score = 24.6 bits (52), Expect = 8.0
Identities = 9/24 (37%), Positives = 16/24 (66%)
Frame = -1
Query: 56 GLVLVGPTLHFWYLYLSQLVTLPG 79
G + + ++ FWYL +S+ T+PG
Sbjct: 270 GSLRIRSSVLFWYLRISRRATVPG 199
>TC10568 similar to UP|Q84MM3 (Q84MM3) Lethal leaf spot 1-like protein,
partial (28%)
Length = 742
Score = 24.6 bits (52), Expect = 8.0
Identities = 8/21 (38%), Positives = 15/21 (71%)
Frame = +2
Query: 52 FSFLGLVLVGPTLHFWYLYLS 72
F L ++++ TLH+W++ LS
Sbjct: 671 FCSLKIIILRKTLHYWFIKLS 733
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.332 0.146 0.456
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,932,730
Number of Sequences: 28460
Number of extensions: 41714
Number of successful extensions: 316
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 316
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 316
length of query: 157
length of database: 4,897,600
effective HSP length: 83
effective length of query: 74
effective length of database: 2,535,420
effective search space: 187621080
effective search space used: 187621080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC149050.9


