

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
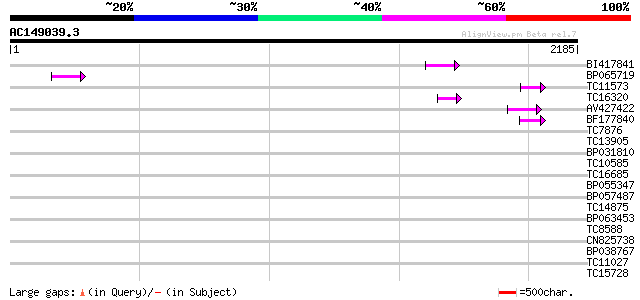
Query= AC149039.3 + phase: 0 /pseudo
(2185 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI417841 72 1e-12
BP065719 64 3e-10
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 61 2e-09
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 59 1e-08
AV427422 55 1e-07
BF177840 48 1e-05
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 32 1.1
TC13905 similar to UP|Q8H1U4 (Q8H1U4) APC5, partial (12%) 32 1.1
BP031810 31 1.8
TC10585 similar to UP|Q9ZP40 (Q9ZP40) Plastoglobule associated p... 30 3.1
TC16685 homologue to UP|O64614 (O64614) At2g18840 protein (Expre... 30 4.0
BP055347 30 4.0
BP057487 30 4.0
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 30 4.0
BP063453 30 5.3
TC8588 similar to UP|CAE54085 (CAE54085) Superoxide dismutase (... 30 5.3
CN825738 30 5.3
BP038767 29 6.9
TC11027 homologue to UP|Q39828 (Q39828) SDL5A, partial (44%) 29 6.9
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 29 6.9
>BI417841
Length = 617
Score = 71.6 bits (174), Expect = 1e-12
Identities = 41/136 (30%), Positives = 66/136 (48%), Gaps = 2/136 (1%)
Frame = +1
Query: 1601 NSKWGLVFDGAV--NAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEA 1658
N L FDG+ N G GAV+ + G + + + TNN AEY I G++ A
Sbjct: 127 NRSCTLEFDGSSKGNPGSAGAGAVLRAEDGSKV-YLREGVGNQTNNQAEYRGLILGLKHA 303
Query: 1659 IDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQ 1718
+ +H+++ GDS LV Q++G W+ + + + A+ L + F +++H+PR N
Sbjct: 304 HEQGYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNS 483
Query: 1719 MADALATLSSMFRVNH 1734
AD A L H
Sbjct: 484 EADVQANLGVNLPAGH 531
>BP065719
Length = 567
Score = 63.5 bits (153), Expect = 3e-10
Identities = 38/132 (28%), Positives = 70/132 (52%), Gaps = 2/132 (1%)
Frame = +3
Query: 162 VSKVDVPKKFKVPEFDKYNGLTCPQN--HIVKYVRKMSNYKDNDSLMIHCFQDSLMEDAA 219
V + ++P+ +KVP+F K++G + HI +Y + + N++L + F SL ++A
Sbjct: 147 VLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAF 326
Query: 220 EWYTSLSKDDVHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAA 279
W+T+L+ VHT+ +L F + F K + + L S+ +K ES +Y R+R
Sbjct: 327 TWFTTLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLK 503
Query: 280 ARITPALDEEEM 291
+R + E E+
Sbjct: 504 SRCFTHVSEHEL 539
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 61.2 bits (147), Expect = 2e-09
Identities = 35/96 (36%), Positives = 48/96 (49%), Gaps = 1/96 (1%)
Frame = +2
Query: 1968 DNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNI-KRIVQKMVTTYKDWHE 2026
+NG + Q C+ I+ SS PQ NG EA NK I K I +++ W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 2027 MLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2062
LP L Y T +SS TPF L YG + +LP+E+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 58.5 bits (140), Expect = 1e-08
Identities = 29/91 (31%), Positives = 44/91 (47%)
Frame = +2
Query: 1648 YEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKV 1707
Y I G++ AI KH+ + GDS LV NQ++G W+ + + A+ L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1708 ELHHIPRDENQMADALATLSSMFRVNHWNDV 1738
+++HIPR+ N AD A R +V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>AV427422
Length = 417
Score = 55.1 bits (131), Expect = 1e-07
Identities = 33/133 (24%), Positives = 63/133 (46%), Gaps = 2/133 (1%)
Frame = +1
Query: 1917 SNGHHFILVAIDYFTKWVEAASYTN-VTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNN 1975
S G+ +LV +D +K+ + T +V+A ++ +GVP I++D +
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 1976 NVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMVTTY-KDWHEMLPYALHG 2034
N + L + + S+ Y P+ +G E N+ ++ ++ + K W +P+A +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 2035 YRTTVRSSTGATP 2047
Y T+ STG TP
Sbjct: 379 YNTSYHVSTGQTP 417
>BF177840
Length = 410
Score = 48.1 bits (113), Expect = 1e-05
Identities = 26/101 (25%), Positives = 49/101 (47%), Gaps = 1/101 (0%)
Frame = +2
Query: 1963 SKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEATNKNIKRIVQKMVT-TY 2021
+ I++D T ++ + L + + S+ PQ +G E NK + +++ ++
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 2022 KDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2062
K W LP+ Y V S+T +PF +VYG + PL++
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 32.0 bits (71), Expect = 1.1
Identities = 24/86 (27%), Positives = 40/86 (45%), Gaps = 11/86 (12%)
Frame = +1
Query: 25 LLAAQEQPA--PVVSAAANVIPT-----ATTDPRFTMPPGYPY----GLPPYYTANTGAG 73
L+A+ PA V SA+ ++P ++ PR + P P PP TA + G
Sbjct: 289 LMASSPSPAHGTVSSASGTLLPAPPPAASSVTPRTSSPSLSPLTTVRSCPPPATAPSSCG 468
Query: 74 TSGTTSNGPISGANLTPVSTALTQAA 99
T +++ P LTP+ +A+ +A
Sbjct: 469 TPSVSASTPFRTTTLTPIGSAV*DSA 546
>TC13905 similar to UP|Q8H1U4 (Q8H1U4) APC5, partial (12%)
Length = 733
Score = 32.0 bits (71), Expect = 1.1
Identities = 18/40 (45%), Positives = 26/40 (65%), Gaps = 1/40 (2%)
Frame = +3
Query: 479 F*FHFTFVKITF-FSSILFLNHNLVQNFLIFIFCPFLIAV 517
F F F+F+ F FS I+F +H+L +FL +IF FL+ V
Sbjct: 474 FSFIFSFIIFPFTFSFIIFPHHSLHLSFLSYIFISFLVWV 593
>BP031810
Length = 393
Score = 31.2 bits (69), Expect = 1.8
Identities = 16/52 (30%), Positives = 26/52 (49%), Gaps = 2/52 (3%)
Frame = -3
Query: 1296 IGMPLTFPQSCMFLSNIPCCLSILFFQNNN--PLALPKARVIFSVRACLFRH 1345
+G+PLT +C+ SN+PC +L +N P+ + A + R L H
Sbjct: 217 LGVPLTLSNTCLASSNLPCSTRLLGVSGHNKPPINITVAGTVAKPRDNLHPH 62
>TC10585 similar to UP|Q9ZP40 (Q9ZP40) Plastoglobule associated protein PG1
precursor, partial (52%)
Length = 807
Score = 30.4 bits (67), Expect = 3.1
Identities = 22/76 (28%), Positives = 37/76 (47%), Gaps = 4/76 (5%)
Frame = -3
Query: 32 PAPVVSAAANVIPTATTDPRFTMPP--GYPYGLPPYYTANTGAGTSGT--TSNGPISGAN 87
P+ SA A+V P T PP P PP+ +++ G TSG+ + SG+
Sbjct: 412 PSTPSSAGASVGVAGVPLPAST*PPPVSAPAATPPHSSSSLGGSTSGSLGADDASGSGSE 233
Query: 88 LTPVSTALTQAATTVT 103
L+P S+ ++ A + +
Sbjct: 232 LSPHSSEMSADALSAS 185
>TC16685 homologue to UP|O64614 (O64614) At2g18840 protein (Expressed
protein), partial (30%)
Length = 465
Score = 30.0 bits (66), Expect = 4.0
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = +2
Query: 485 FVKITFFSSILFLNHNLVQNFLIFIFCPFLIAVQSFFIIF 524
FV IT ++ +L N L+ NFL F F P + SFF +F
Sbjct: 281 FVPITSWNLLLNRNFCLMVNFL-FFFSPVFVPTFSFFFLF 397
>BP055347
Length = 538
Score = 30.0 bits (66), Expect = 4.0
Identities = 14/42 (33%), Positives = 23/42 (54%)
Frame = +3
Query: 1032 LLLVTTSGLVKKSCRKWERVTIWLCTSR*IANQT*YQMCWWI 1073
L L+ T L+K+ C WE IW S+*+ + ++ WW+
Sbjct: 411 LYLMGTKVLLKRICEVWEA*LIWRQCSK*VKHM--IRLIWWL 530
>BP057487
Length = 512
Score = 30.0 bits (66), Expect = 4.0
Identities = 16/47 (34%), Positives = 23/47 (48%)
Frame = -3
Query: 1296 IGMPLTFPQSCMFLSNIPCCLSILFFQNNNPLALPKARVIFSVRACL 1342
I MPL P + CC + LF + N P+ + AR+ F + A L
Sbjct: 309 IEMPLAAP--------LWCCSTALFCKRNKPVVIVTARMFFRIEAAL 193
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 30.0 bits (66), Expect = 4.0
Identities = 20/82 (24%), Positives = 36/82 (43%), Gaps = 1/82 (1%)
Frame = +1
Query: 32 PAPVVSAAANVIPTATTDPRFTMPPGYPYGL-PPYYTANTGAGTSGTTSNGPISGANLTP 90
PAP + +N T+T+ P PP L PP + ++ T+ +++ P++ T
Sbjct: 118 PAPTI*PPSNP-KTSTSTPTPASPPASTSPLTPPTTSPSSSTSTAAPSASAPLTTTPTTA 294
Query: 91 VSTALTQAATTVTEPIVNTVPQ 112
ST + T + P P+
Sbjct: 295 TSTTSPRRQTPSSSPSTTASPR 360
>BP063453
Length = 522
Score = 29.6 bits (65), Expect = 5.3
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +2
Query: 341 FVNSLLFFSESLLSF*NIFYCLCLSFLALQV 371
F+ + FS +L + ++F CLC+S L +Q+
Sbjct: 398 FIKEICMFSNNLNKWSSVFMCLCVSKLLIQI 490
>TC8588 similar to UP|CAE54085 (CAE54085) Superoxide dismutase (Fragment)
, complete
Length = 837
Score = 29.6 bits (65), Expect = 5.3
Identities = 20/71 (28%), Positives = 33/71 (46%), Gaps = 7/71 (9%)
Frame = -1
Query: 42 VIPTATTDPRFTMPPGYPYGL----PPYYTANTGAGTSGTTSN---GPISGANLTPVSTA 94
V P P+ T+PP +P L PP+ ++ A T+ GP+ G L+ +
Sbjct: 531 VYPCKPIIPQATLPPAFPVVLLSSWPPFPRSSGSAWTTTALPMMEFGPVRGI*LSVIEKL 352
Query: 95 LTQAATTVTEP 105
+ ++ TVT P
Sbjct: 351 VVPSSPTVTFP 319
>CN825738
Length = 335
Score = 29.6 bits (65), Expect = 5.3
Identities = 28/108 (25%), Positives = 55/108 (50%)
Frame = -1
Query: 25 LLAAQEQPAPVVSAAANVIPTATTDPRFTMPPGYPYGLPPYYTANTGAGTSGTTSNGPIS 84
L+ + +P P+ S++A+VIP + T + T+P LPP T++ GA + S
Sbjct: 314 LIPSSTKPLPLTSSSASVIPFSPTSTKSTIPFS---PLPP--TSSKGAESIS------FS 168
Query: 85 GANLTPVSTALTQAATTVTEPIVNTVPQFVHANAHRGSIATTGTMEER 132
G + + ++LT+ + T ++ +P F + S+A + + EE+
Sbjct: 167 GNSSKSLFSSLTREESVCT--*MSALPSFNSST----SVAISASAEEK 42
>BP038767
Length = 561
Score = 29.3 bits (64), Expect = 6.9
Identities = 25/70 (35%), Positives = 30/70 (42%), Gaps = 5/70 (7%)
Frame = +1
Query: 1390 HKKPSF*NHPTILRRLNYNEPVGHSNPAFPP---NFEFPVHD-AENEEDDDIPYEITRLL 1445
H P+ +HP I R N + +S P PP P H EED PY+ RL
Sbjct: 247 HHAPNSNSHPKIRTRRNQQQ---NSLPVAPPEEATISVPFHPLLRQEEDSRNPYQP*RLP 417
Query: 1446 EQEG-RAIQP 1454
Q G R I P
Sbjct: 418 RQPGSRPIPP 447
>TC11027 homologue to UP|Q39828 (Q39828) SDL5A, partial (44%)
Length = 826
Score = 29.3 bits (64), Expect = 6.9
Identities = 22/95 (23%), Positives = 41/95 (43%), Gaps = 4/95 (4%)
Frame = -2
Query: 1298 MPLTFPQSCMFLSNIPCCLSILFFQNNNPLALPKARVIFSVRA----CLFRHCR*LIKNC 1353
+P++F ++ + + + FQ++ L + + R+ +F++ LIK
Sbjct: 600 IPISFSNKFPYIFH*ELLIKLQPFQSSRELIVKDIINFITSRSHAIKMIFKY---LIK*A 430
Query: 1354 RFFHDCFFLVPFFSWK*W*CQKKPKYLYFLINFKN 1388
+FHDC L P FS C + + F F N
Sbjct: 429 TYFHDCIQLAPIFS-----CNRLSQTTQFSFKFSN 340
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 29.3 bits (64), Expect = 6.9
Identities = 25/91 (27%), Positives = 35/91 (37%)
Frame = +2
Query: 15 LAKAHDAMTALLAAQEQPAPVVSAAANVIPTATTDPRFTMPPGYPYGLPPYYTANTGAGT 74
LA+AH A T P P +A P+ PP PP ++G
Sbjct: 191 LARAHSAATTATTPAHAPTPPPLETTASCSSACASPKAPPPP------PP----SSGRAL 340
Query: 75 SGTTSNGPISGANLTPVSTALTQAATTVTEP 105
+ TTS I+ N TP++ T T+ P
Sbjct: 341 A*TTSPS-ITNPNPTPLTQLATPPTTSFIPP 430
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.148 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,463,525
Number of Sequences: 28460
Number of extensions: 608357
Number of successful extensions: 4926
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 4802
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4925
length of query: 2185
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2080
effective length of database: 1,909,300
effective search space: 3971344000
effective search space used: 3971344000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149039.3


