

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
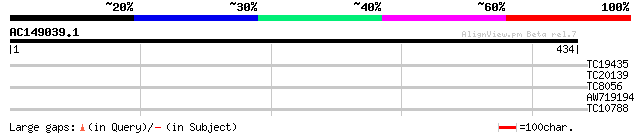
Query= AC149039.1 + phase: 0 /pseudo
(434 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein red... 25 0.77
TC20139 similar to GB|AAG31805.1|11139702|AF315590 Pumilio 2 {Mu... 28 3.9
TC8056 27 6.6
AW719194 27 6.6
TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type b... 27 8.6
>TC19435 similar to UP|Q39036 (Q39036) NADPH-ferrihemoprotein reductase ,
partial (12%)
Length = 459
Score = 25.4 bits (54), Expect(2) = 0.77
Identities = 9/30 (30%), Positives = 17/30 (56%), Gaps = 2/30 (6%)
Frame = -2
Query: 304 WKQFFLWK--WRSHLSV*SWKQSYPRLNGA 331
W + W+ W+ + +*SW+ SYP + +
Sbjct: 182 WSRRRRWRRRWKGRVCL*SWRTSYPAVKSS 93
Score = 23.1 bits (48), Expect(2) = 0.77
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -1
Query: 240 ITLLPIDLR*MVQLRPPTRISRELSR 265
+ LLP+DLR M P R + E+ R
Sbjct: 288 LALLPVDLRQMRTTTQPMRTAMEVVR 211
>TC20139 similar to GB|AAG31805.1|11139702|AF315590 Pumilio 2 {Mus
musculus;} , partial (7%)
Length = 425
Score = 27.7 bits (60), Expect = 3.9
Identities = 17/56 (30%), Positives = 27/56 (47%), Gaps = 3/56 (5%)
Frame = -3
Query: 97 GILCQGSCYGQVTTGWP---WSMIATSTPESATNVKSMLIRFMCLRTLSMLFHPHG 149
G C +C+ TT +P W+ + T+ S+ +L C RTL + HP+G
Sbjct: 216 GFPCSSTCW---TT*FPY*SWANMLTAPKISSITFCCVLGSLQCSRTLCITRHPYG 58
>TC8056
Length = 730
Score = 26.9 bits (58), Expect = 6.6
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = -1
Query: 85 MYMTVPSGPMLQGILCQGSCYGQVTTGWPWSMIATS 120
++ +VP P+LQ +L QG C G+ P + +TS
Sbjct: 589 IWSSVPEIPLLQTLLHQGQCGGRFYRRCPHKVDSTS 482
>AW719194
Length = 526
Score = 26.9 bits (58), Expect = 6.6
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = -1
Query: 85 MYMTVPSGPMLQGILCQGSCYGQVTTGWPWSMIATS 120
++ +VP P+LQ +L QG C G+ P + +TS
Sbjct: 115 IWSSVPEIPLLQTLLHQGQCGGRFYRRCPHKVDSTS 8
>TC10788 weakly similar to UP|Q8YG29 (Q8YG29) Cytochrome C-type biogenesis
protein, partial (25%)
Length = 656
Score = 26.6 bits (57), Expect = 8.6
Identities = 19/54 (35%), Positives = 27/54 (49%), Gaps = 10/54 (18%)
Frame = -2
Query: 86 YMTVPSGPM-------LQGIL-CQGSCYGQVT--TGWPWSMIATSTPESATNVK 129
Y T+PS L+G+L C S VT T W W +++T+T AT+ K
Sbjct: 553 YTTLPSRTRPPNRNFALEGLLRCFSSMSVGVT*NTNWSWKLLSTTTMNPATHPK 392
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.354 0.153 0.564
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,437,478
Number of Sequences: 28460
Number of extensions: 127794
Number of successful extensions: 1508
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1497
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1508
length of query: 434
length of database: 4,897,600
effective HSP length: 93
effective length of query: 341
effective length of database: 2,250,820
effective search space: 767529620
effective search space used: 767529620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC149039.1


