

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148995.8 + phase: 0
(325 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
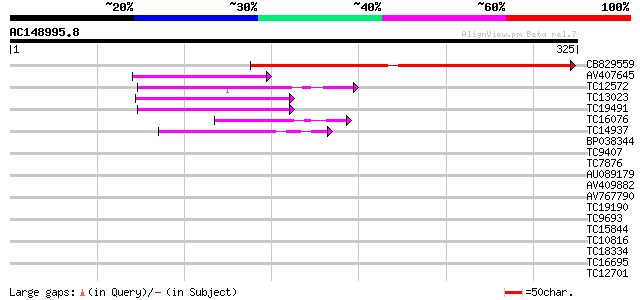
Score E
Sequences producing significant alignments: (bits) Value
CB829559 340 1e-94
AV407645 47 4e-06
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 45 1e-05
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 45 2e-05
TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 42 1e-04
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 40 4e-04
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 40 5e-04
BP038344 39 0.001
TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription in... 38 0.003
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 37 0.003
AU089179 36 0.008
AV409882 35 0.013
AV767790 35 0.017
TC19190 weakly similar to PIR|T46032|T46032 WD-40 repeat regulat... 34 0.038
TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit ... 33 0.086
TC15844 homologue to UP|Q9FV61 (Q9FV61) Heterotrimeric GTP-bindi... 32 0.11
TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subun... 31 0.25
TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1 (AT3... 30 0.43
TC16695 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-intera... 30 0.43
TC12701 similar to UP|O63266 (O63266) Cytochrome oxidase subunit... 30 0.56
>CB829559
Length = 552
Score = 340 bits (873), Expect = 1e-94
Identities = 166/186 (89%), Positives = 170/186 (91%)
Frame = +1
Query: 139 VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELE 198
VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI YRFGSVGQDTQLLLW+LE
Sbjct: 1 VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETIMYRFGSVGQDTQLLLWDLE 180
Query: 199 MDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAH 258
MDEIVVPLRR PPGGSPT+ S GSQSSHWDN VPLGTLQPAPSMRDVPKISPLVAH
Sbjct: 181 MDEIVVPLRRCPPGGSPTY-----STGSQSSHWDNVVPLGTLQPAPSMRDVPKISPLVAH 345
Query: 259 RVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLATSLKEKPSLSSKI 318
RVHTEPLS LIFTQESVLTACREGHIK+W RPG ESQPSNSET LATSLKEKP LSSK+
Sbjct: 346 RVHTEPLSGLIFTQESVLTACREGHIKVWMRPGAPESQPSNSETSLATSLKEKPLLSSKL 525
Query: 319 SNSIYK 324
SNS YK
Sbjct: 526 SNSSYK 543
>AV407645
Length = 413
Score = 47.0 bits (110), Expect = 4e-06
Identities = 24/80 (30%), Positives = 43/80 (53%)
Frame = +2
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
Q + +SIS+S + L T G + +R +D + + + GL+ C W GKYIL+
Sbjct: 140 QKAFSSISWSPNDQELLTCGVEEAIRRWDVSTGKCLQIYEKAGAGLISCTWFPSGKYILS 319
Query: 131 GGEDDLVQVWSMEDRKVVAW 150
G D + +W ++ ++V +W
Sbjct: 320 GLSDKSICMWELDGKEVESW 379
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 45.4 bits (106), Expect = 1e-05
Identities = 29/130 (22%), Positives = 58/130 (44%), Gaps = 3/130 (2%)
Frame = +3
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSM---DGKYILT 130
++ + FS + ++ D LR+++Y+ + + C + + +GKY++
Sbjct: 177 VSFVKFSPNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISSTFSTTNGKYVVG 356
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
G ED + +W ++ RK+V EGH+ V V+ P N G++G D
Sbjct: 357 GSEDHGIYLWELQTRKIVQKLEGHSDTVVSVS-----CHPTEN------MIASGALGNDK 503
Query: 191 QLLLWELEMD 200
+ +W + D
Sbjct: 504 TVKIWTQQKD 533
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 45.1 bits (105), Expect = 2e-05
Identities = 26/91 (28%), Positives = 43/91 (46%)
Frame = +1
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S+ +++ S D L + G +RV+D + V K + G ++C + G + TGG
Sbjct: 289 SVTALALSPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGPVMCMSCHPSGGLLATGG 468
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
D V VW ++ + +GH VS V F
Sbjct: 469 ADRKVLVWDVDGGYCTHFFKGHGGVVSCVMF 561
>TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (39%)
Length = 527
Score = 42.4 bits (98), Expect = 1e-04
Identities = 29/91 (31%), Positives = 41/91 (44%), Gaps = 1/91 (1%)
Frame = +1
Query: 74 INSISFSA-DGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
+ S+ +S D L T DG + ++D + LV + +LC S DG I TG
Sbjct: 25 VRSLVYSPYDPRLLFTASDDGNVHMYDAEGKALVGTMSGHASWVLCVDVSPDGGAIATGS 204
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
D V++W + R V H V GVAF
Sbjct: 205 SDRTVRLWDLAMRASVQTMSNHTDQVWGVAF 297
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 40.4 bits (93), Expect = 4e-04
Identities = 23/79 (29%), Positives = 38/79 (47%)
Frame = +3
Query: 118 CCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGE 177
CC +S DGK + + G+D V +W+M+ + + E H S +S V F PNS+
Sbjct: 9 CCHFSSDGKMLASAGDDKKVVLWNMDTLQTESTPEQHKSVISDVRF-----RPNSS---- 161
Query: 178 TITYRFGSVGQDTQLLLWE 196
+ + D + LW+
Sbjct: 162 ----QLATASIDKSVRLWD 206
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 40.0 bits (92), Expect = 5e-04
Identities = 25/100 (25%), Positives = 47/100 (47%)
Frame = +1
Query: 86 LATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR 145
LA+ D ++++D L+ + G+ A+S +G+Y+ +G D + +WS+++
Sbjct: 55 LASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIWSLKEG 234
Query: 146 KVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGS 185
K+V G SG F+ W N G+ I F +
Sbjct: 235 KIVKTYTG-----SGGIFEVCW-----NKEGDKIAACFAN 324
>BP038344
Length = 525
Score = 38.9 bits (89), Expect = 0.001
Identities = 27/101 (26%), Positives = 42/101 (40%)
Frame = +2
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
++ + FS DG LA+ D L ++ L+ + G+ AWS D YI +
Sbjct: 104 AVACVKFSNDGTLLASASLDKTLIIWSSATLSLLHRLTGHSEGISDLAWSSDSHYICSAS 283
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSN 173
+D +++W V GH V V F +P SN
Sbjct: 284 DDRTLRIWDATGGDCVKTLRGHTHAVFCVNF-----NPQSN 391
>TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 530
Score = 37.7 bits (86), Expect = 0.003
Identities = 21/83 (25%), Positives = 41/83 (49%), Gaps = 3/83 (3%)
Frame = +2
Query: 76 SISFSADGAYLATVGRDGYLRVFD---YTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S F YLAT D +++++ +T + + G + + C +S+DG Y++T
Sbjct: 29 SPEFCEPHRYLATASADHTVKIWNVDGFTLDKTLIGHQRWVWD---CVFSVDGAYLITAS 199
Query: 133 EDDLVQVWSMEDRKVVAWGEGHN 155
D ++WSM + + +GH+
Sbjct: 200 SDTTARLWSMSTGEDIKVYQGHH 268
Score = 26.9 bits (58), Expect = 4.7
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +2
Query: 79 FSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCA 120
FS DGAYL T D R++ + + + ++ CCA
Sbjct: 164 FSVDGAYLITASSDTTARLWSMSTGEDIKVYQGHHKATTCCA 289
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 37.4 bits (85), Expect = 0.003
Identities = 19/81 (23%), Positives = 40/81 (48%), Gaps = 4/81 (4%)
Frame = +3
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYY----GGLLCCAWSMDGKY 127
G +N+++ S DG+ A+ G+DG + ++D + GK Y G ++ +Y
Sbjct: 651 GYVNTVAVSPDGSLCASGGKDGVILLWDLAE------GKRLYSLDAGSIIHALCFSPNRY 812
Query: 128 ILTGGEDDLVQVWSMEDRKVV 148
L + +++W +E + +V
Sbjct: 813 WLCAATETSIKIWDLESKSIV 875
Score = 32.0 bits (71), Expect = 0.15
Identities = 29/126 (23%), Positives = 59/126 (46%), Gaps = 4/126 (3%)
Frame = +3
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKE--HLVCGGKSYYGGLLCCAWSMDGKY--ILTG 131
S++FS D + + RD +++++ E + + ++ + C +S I++
Sbjct: 399 SVAFSIDNRQIVSASRDRTIKLWNTLGECKYTIQDNDAHSDWVSCVRFSPSTLQPTIVSA 578
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D V+VW++ + K+ GH+ +V+ VA SP+ + S G+D
Sbjct: 579 SWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAV-----SPDGS--------LCASGGKDGV 719
Query: 192 LLLWEL 197
+LLW+L
Sbjct: 720 ILLWDL 737
Score = 28.5 bits (62), Expect = 1.6
Identities = 29/117 (24%), Positives = 47/117 (39%), Gaps = 7/117 (5%)
Frame = +3
Query: 86 LATVGRDGYLRVFDYTKEHLVCG-------GKSYYGGLLCCAWSMDGKYILTGGEDDLVQ 138
+ T RD + ++ TKE G G S++ + S DG++ L+G D ++
Sbjct: 162 IVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHF--VQDVVLSSDGQFALSGSWDGELR 335
Query: 139 VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLW 195
+W + GH V VAF S DN + + S +D + LW
Sbjct: 336 LWDLAAGTSARRFVGHTKDVLSVAF--------SIDNRQIV-----SASRDRTIKLW 467
Score = 27.7 bits (60), Expect = 2.8
Identities = 14/51 (27%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Frame = +3
Query: 95 LRVFDYTKEHLVCGGKSYYGGLLCCA---WSMDGKYILTGGEDDLVQVWSM 142
L+V T+ GG + ++ C WS DG + +G D +++VW++
Sbjct: 882 LKVDLKTEADATTGGNTNKKKVIYCTSLNWSADGSTLFSGYTDGVIRVWAI 1034
>AU089179
Length = 340
Score = 36.2 bits (82), Expect = 0.008
Identities = 21/73 (28%), Positives = 39/73 (52%), Gaps = 1/73 (1%)
Frame = +3
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVC-GGKSYYGGLLCCAWSMDGKYIL 129
+G +++F+ DG LAT G DG LRVF + ++ ++ + +S +GK+I+
Sbjct: 6 EGQQLALAFNNDGTALATGGEDGSLRVFKWPXMKVILEHSTAHSSSVKDLHFSSNGKWIV 185
Query: 130 TGGEDDLVQVWSM 142
+ G +VW +
Sbjct: 186 SLGSGGPCRVWDL 224
>AV409882
Length = 422
Score = 35.4 bits (80), Expect = 0.013
Identities = 23/108 (21%), Positives = 47/108 (43%), Gaps = 1/108 (0%)
Frame = +3
Query: 58 YSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLL 117
Y + + + + S F A ++ D ++RV++Y V +++ +
Sbjct: 57 YQSQTMAKSFEVTELPVRSAKFVARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIR 236
Query: 118 CCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAW-GEGHNSWVSGVAFD 164
C A Y+L+ +D L+++W E + EGH+ +V V F+
Sbjct: 237 CVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQIFEGHSHYVMQVTFN 380
>AV767790
Length = 617
Score = 35.0 bits (79), Expect = 0.017
Identities = 13/40 (32%), Positives = 24/40 (59%)
Frame = +3
Query: 116 LLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHN 155
++CC +S DGK + + G D V +W+ME + + + H+
Sbjct: 483 VVCCHFSSDGKLLASAGHDQKVVLWNMETLQTESTPDAHS 602
>TC19190 weakly similar to PIR|T46032|T46032 WD-40 repeat regulatory protein
tup1 homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (18%)
Length = 511
Score = 33.9 bits (76), Expect = 0.038
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = +3
Query: 124 DGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGV 161
+G+YI++G ED V +W ++ + ++ EGH V V
Sbjct: 12 NGRYIVSGSEDRCVYLWDLQQKNMIQKLEGHTDTVISV 125
>TC9693 similar to UP|COPP_MOUSE (O55029) Coatomer beta' subunit
(Beta'-coat protein) (Beta'-COP) (p102), partial (20%)
Length = 743
Score = 32.7 bits (73), Expect = 0.086
Identities = 22/108 (20%), Positives = 46/108 (42%), Gaps = 1/108 (0%)
Frame = +1
Query: 58 YSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLL 117
Y + + + + S F A ++ D ++RV++Y V +++ +
Sbjct: 337 YQTQTMAKSFEVTELPVRSAKFIARKQWVVAGADDMFIRVYNYNTMDKVKVFEAHTDYIR 516
Query: 118 CCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAW-GEGHNSWVSGVAFD 164
C A Y+L+ +D L+++W E + E H+ +V V F+
Sbjct: 517 CVAVHPTLPYVLSSSDDMLIKLWDWEKGWICTQICEAHSHYVMQVTFN 660
>TC15844 homologue to UP|Q9FV61 (Q9FV61) Heterotrimeric GTP-binding protein
subunit beta 1, partial (28%)
Length = 668
Score = 32.3 bits (72), Expect = 0.11
Identities = 21/72 (29%), Positives = 33/72 (45%), Gaps = 4/72 (5%)
Frame = +3
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCG----GKSYYGGLLCCAWSMDGKYIL 129
+ SI+FS G L +G V+D +V S+ G + C S DG +
Sbjct: 93 VTSIAFSISGRLLFAGYANGDCYVWDTLLAKVVLNIGSLQDSHEGRISCLGLSADGSALC 272
Query: 130 TGGEDDLVQVWS 141
TGG D +++W+
Sbjct: 273 TGGWDTNLKIWA 308
>TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (20%)
Length = 979
Score = 31.2 bits (69), Expect = 0.25
Identities = 15/92 (16%), Positives = 39/92 (42%)
Frame = +1
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTG 131
G + + F + G D ++V++Y + + + + + +I++
Sbjct: 394 GPVRGVHFHHSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSA 573
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
+D +++W+ + R ++ GHN +V F
Sbjct: 574 SDDQTIRIWNWQSRTCISVLTGHNHYVMCALF 669
>TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1
(AT3g18860/MCB22_3), partial (21%)
Length = 595
Score = 30.4 bits (67), Expect = 0.43
Identities = 23/89 (25%), Positives = 39/89 (42%), Gaps = 10/89 (11%)
Frame = +2
Query: 86 LATVGRDGYLRVFD-----YTKEHLVCGGKSYYGGLLCCAW-----SMDGKYILTGGEDD 135
+AT RD +R + + ++ G S+ G L+ W + + +GG D
Sbjct: 191 IATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVGPLV---WIPPNPELPQGGVASGGMDT 361
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
LV VW + + +GH V+ +AFD
Sbjct: 362 LVLVWDLSTGEKAHTLKGHQLQVTSIAFD 448
>TC16695 homologue to UP|Q94KS2 (Q94KS2) TGF-beta receptor-interacting
protein 1, partial (46%)
Length = 531
Score = 30.4 bits (67), Expect = 0.43
Identities = 14/67 (20%), Positives = 31/67 (45%)
Frame = +3
Query: 77 ISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDL 136
+ ++ DG L + +D V+ + + + G + CC S D ++TG D
Sbjct: 123 LKYNRDGDLLFSCAKDHNPTVWFADNGERLGTYRGHNGAVWCCDVSRDSSRLITGSADQT 302
Query: 137 VQVWSME 143
++W+++
Sbjct: 303 AKLWNVQ 323
>TC12701 similar to UP|O63266 (O63266) Cytochrome oxidase subunit I
(Cytochrome c oxidase polypeptide I) (Fragment) ,
partial (5%)
Length = 698
Score = 30.0 bits (66), Expect = 0.56
Identities = 22/75 (29%), Positives = 31/75 (41%)
Frame = +3
Query: 213 GSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQ 272
GS F S G Q PL T A ++ HR L+S+ TQ
Sbjct: 162 GSDDFYETKSSFGVQVVQETKRRPLTTKVTAKAILAAAATDSAGCHRDSIVSLASVKLTQ 341
Query: 273 ESVLTACREGHIKIW 287
+L++ R+G IK+W
Sbjct: 342 RLLLSSGRDGAIKVW 386
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,827,878
Number of Sequences: 28460
Number of extensions: 107300
Number of successful extensions: 621
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 593
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 612
length of query: 325
length of database: 4,897,600
effective HSP length: 90
effective length of query: 235
effective length of database: 2,336,200
effective search space: 549007000
effective search space used: 549007000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148995.8


