

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148970.12 + phase: 0
(560 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
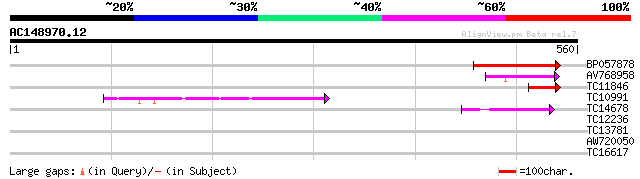
Score E
Sequences producing significant alignments: (bits) Value
BP057878 171 3e-43
AV768958 74 8e-14
TC11846 homologue to UP|Q9FEL1 (Q9FEL1) Lysyl-tRNA synthetase (F... 68 3e-12
TC10991 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-lik... 58 5e-09
TC14678 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-lik... 49 2e-06
TC12236 similar to UP|Q9LJE2 (Q9LJE2) Lysyl-tRNA synthetase, par... 36 0.019
TC13781 homologue to UP|SYN1_ARATH (Q9SW96) Asparaginyl-tRNA syn... 35 0.042
AW720050 28 5.1
TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protei... 28 5.1
>BP057878
Length = 546
Score = 171 bits (433), Expect = 3e-43
Identities = 81/86 (94%), Positives = 84/86 (97%)
Frame = -2
Query: 459 KHELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGI 518
KHE+ NAYTELNDPVVQRQRFA+QLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGI
Sbjct: 545 KHEVCNAYTELNDPVVQRQRFAEQLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGI 366
Query: 519 DRLTMLLTDSQNIKEVLLFPAMKPQD 544
DRLTM+ TDSQNIKEVLLFPAMKPQD
Sbjct: 365 DRLTMIFTDSQNIKEVLLFPAMKPQD 288
>AV768958
Length = 436
Score = 73.6 bits (179), Expect = 8e-14
Identities = 38/91 (41%), Positives = 52/91 (56%), Gaps = 18/91 (19%)
Frame = -3
Query: 471 DPVVQRQRFADQLKDRQS------------------GDDEAMALDETFCTALEYGLPPTA 512
DP+ QR R DQ++ + D + LD+ F TALEYG+PP +
Sbjct: 431 DPIDQRGRLEDQIRQHEKKKAAISTNGDPKEGIENEDDSYEVTLDDDFLTALEYGMPPAS 252
Query: 513 GWGLGIDRLTMLLTDSQNIKEVLLFPAMKPQ 543
G GLGIDRL MLLT+S +I++V+ FP +K Q
Sbjct: 251 GMGLGIDRLVMLLTNSPSIRDVIAFPVLKVQ 159
>TC11846 homologue to UP|Q9FEL1 (Q9FEL1) Lysyl-tRNA synthetase (Fragment),
partial (19%)
Length = 570
Score = 68.2 bits (165), Expect = 3e-12
Identities = 31/32 (96%), Positives = 32/32 (99%)
Frame = +2
Query: 513 GWGLGIDRLTMLLTDSQNIKEVLLFPAMKPQD 544
GWGLGIDRLTM+LTDSQNIKEVLLFPAMKPQD
Sbjct: 2 GWGLGIDRLTMILTDSQNIKEVLLFPAMKPQD 97
>TC10991 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-like protein,
partial (42%)
Length = 680
Score = 57.8 bits (138), Expect = 5e-09
Identities = 59/231 (25%), Positives = 101/231 (43%), Gaps = 7/231 (3%)
Frame = +2
Query: 93 LHDDGFKVQVMADASKSDLNEAGFDKVHSNVKRG---DIVGITGFPGKSRKG---ELSIF 146
+ ++GF VQ + A + D+ K + + R D+VG+ P KG ++ I
Sbjct: 5 VRENGFTVQCLVQA-QPDVVSPQMVKFAAALNRESIIDVVGVVSVPAAPIKGATQQVEIQ 181
Query: 147 PKTFVVLSHCLHMMPRQKSAAAAAADNAWLPGKTRNPEAYTLKDQETRYRLRHLDLMLNT 206
+ + + +P AA + + + + + +Q+TR R LDL
Sbjct: 182 VRKLYCVGRAVPTLPINLEDAARS--DVEVEKALQAGDQLVRVNQDTRLNFRVLDLRTPA 355
Query: 207 EVREIFRTRSKIISYVRSFLDKLDFLEVETPMMNMIAGGAAARPFVTHHNDLNMRLFMRI 266
+ IFR +S + + R FL K +F+E+ TP +IAG + V + +
Sbjct: 356 N-QGIFRIQSHVGNAFRQFLLKENFVEIHTP--KLIAGSSEGGAAVFRLDYKGQPACLAQ 526
Query: 267 APELYLKKLVVGGLNRVYEIGKQFRNE-GIDLTHNPEFTTCEFYMAYKDYY 316
+P+L+ + + G RV+EIG FR E H EFT + M K +Y
Sbjct: 527 SPQLHKQMSICGDFGRVFEIGPVFRAEDSFTHRHLCEFTGLDVEMEIKKHY 679
>TC14678 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-like protein,
partial (25%)
Length = 637
Score = 49.3 bits (116), Expect = 2e-06
Identities = 26/92 (28%), Positives = 45/92 (48%)
Frame = +3
Query: 447 PGLTERFELFINKHELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTALEY 506
P T F++FI E+ + QR D L++R + ++ T+ + Y
Sbjct: 36 PAYTNSFDVFIRGEEIISG--------AQRIHVPDFLEERAAACGIEVSTISTYIDSFRY 191
Query: 507 GLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFP 538
G PP G+G+G++R+ ML NI++ L+P
Sbjct: 192 GAPPHGGFGVGLERVVMLFCGLDNIRKTSLYP 287
>TC12236 similar to UP|Q9LJE2 (Q9LJE2) Lysyl-tRNA synthetase, partial (16%)
Length = 429
Score = 35.8 bits (81), Expect = 0.019
Identities = 22/72 (30%), Positives = 39/72 (53%), Gaps = 2/72 (2%)
Frame = +2
Query: 26 RLKYLSTEKEEGRNPYPHKFSATMTIKQYIKEYEGLSDGQHLDDVS--VSLTGKIMHKRS 83
RLK + + +G NPY + + T + Q Y+ L +G+ + + VS+ G+I+ +R+
Sbjct: 215 RLKKVEELRSKGLNPYAYGWDKTHSADQLQDIYKDLGNGEEANSENDHVSVAGRIVARRA 394
Query: 84 SGAKLVFYDLHD 95
G KL F + D
Sbjct: 395 FG-KLAFXTIRD 427
>TC13781 homologue to UP|SYN1_ARATH (Q9SW96) Asparaginyl-tRNA synthetase,
cytoplasmic 1 (Asparagine--tRNA ligase 1) (AsnRS 1) ,
partial (20%)
Length = 487
Score = 34.7 bits (78), Expect = 0.042
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +2
Query: 498 ETFCTALEYGLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFP 538
E + YG AG+GLG +R+ + T +NI++V+ FP
Sbjct: 194 EWYLDLRRYGTVKHAGFGLGFERMILFATGIENIRDVIPFP 316
>AW720050
Length = 436
Score = 27.7 bits (60), Expect = 5.1
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 1/67 (1%)
Frame = -2
Query: 460 HELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTA-LEYGLPPTAGWGLGI 518
HEL + + + PVV+RQ Q+ G + LD+ + E GL P GLG
Sbjct: 417 HELED-HQQQQHPVVERQSHEPQVL----GSEHHHELDQDQLRSHRELGLEPLDHHGLGQ 253
Query: 519 DRLTMLL 525
DRL LL
Sbjct: 252 DRLEALL 232
>TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (42%)
Length = 551
Score = 27.7 bits (60), Expect = 5.1
Identities = 13/45 (28%), Positives = 22/45 (48%)
Frame = -1
Query: 309 YMAYKDYYDLMEITENMLSGMVYELTKGSYKIKYHADGLDKDPIE 353
Y YK++ + E+ + L Y+ Y ++ HA LD+D E
Sbjct: 473 YHDYKEHPSVPELQRDFLPHFQYQQQSNPYFVQTHAAELDQDQTE 339
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,597,076
Number of Sequences: 28460
Number of extensions: 105150
Number of successful extensions: 447
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 445
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 446
length of query: 560
length of database: 4,897,600
effective HSP length: 95
effective length of query: 465
effective length of database: 2,193,900
effective search space: 1020163500
effective search space used: 1020163500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148970.12


