

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
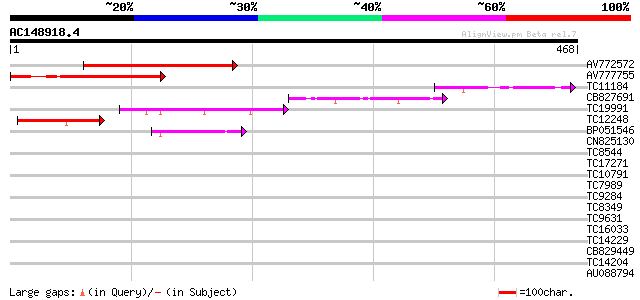
Query= AC148918.4 - phase: 0
(468 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV772572 250 4e-67
AV777755 179 7e-46
TC11184 similar to GB|AAM10292.1|20147151|AY091693 AT3g13810/MCP... 97 6e-21
CB827691 92 2e-19
TC19991 72 1e-13
TC12248 67 5e-12
BP051546 67 8e-12
CN825130 37 0.007
TC8544 32 0.17
TC17271 UP|Q7PHQ1 (Q7PHQ1) ENSANGP00000024044 (Fragment), partia... 31 0.38
TC10791 similar to UP|Q8LSK7 (Q8LSK7) Auxin-regulated protein, p... 30 0.84
TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , par... 30 0.84
TC9284 weakly similar to UP|Q9LPY0 (Q9LPY0) T23J18.22, partial (... 30 0.84
TC8349 30 0.84
TC9631 similar to GB|AAC19270.1|3193286|T14P8 coded for by A. th... 30 1.1
TC16033 UP|Q7X9C1 (Q7X9C1) NIN-like protein 1, partial (88%) 29 1.9
TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%) 28 2.4
CB829449 28 3.2
TC14204 homologue to UP|Q9AVG8 (Q9AVG8) Isopentenyl diphosphate ... 28 3.2
AU088794 28 4.2
>AV772572
Length = 491
Score = 250 bits (638), Expect = 4e-67
Identities = 109/127 (85%), Positives = 120/127 (93%)
Frame = -3
Query: 62 IALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEP 121
IALSPKTLMATNRF+C ICN GFQRDQNLQLH+RGHNLPWKL+QR++ E+RKKVY+CPE
Sbjct: 489 IALSPKTLMATNRFVCVICN*GFQRDQNLQLHRRGHNLPWKLRQRSNKEVRKKVYICPEK 310
Query: 122 TCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCD 181
TCVHHD SRALGDLTGIKKHF RKHGEKKWKC+KCSKKYAV SDWKAH++TCGTREY+CD
Sbjct: 309 TCVHHDASRALGDLTGIKKHFSRKHGEKKWKCEKCSKKYAVPSDWKAHTQTCGTREYKCD 130
Query: 182 CGTLFSR 188
CGTLFSR
Sbjct: 129 CGTLFSR 109
>AV777755
Length = 373
Score = 179 bits (455), Expect = 7e-46
Identities = 87/128 (67%), Positives = 99/128 (76%)
Frame = +2
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAE 60
+ N+ +ASGEAS +SSGN Q + T T AKK+RNLPG PDP AE
Sbjct: 29 LDNVCTASGEASVSSSGN-----------QIVPPPKAAT--TTTAKKKRNLPGMPDPDAE 169
Query: 61 VIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPE 120
VIALSPKTLMATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKL+QR+S E+RK+VYVCPE
Sbjct: 170 VIALSPKTLMATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLRQRSSKEVRKRVYVCPE 349
Query: 121 PTCVHHDP 128
PTC HHDP
Sbjct: 350 PTCAHHDP 373
>TC11184 similar to GB|AAM10292.1|20147151|AY091693 AT3g13810/MCP4_2
{Arabidopsis thaliana;}, partial (4%)
Length = 481
Score = 97.1 bits (240), Expect = 6e-21
Identities = 66/141 (46%), Positives = 79/141 (55%), Gaps = 24/141 (17%)
Frame = +2
Query: 351 SNGNF-SLNLSSCEDQMINNSFSS---------------------SGFHGTSFEDTFA-G 387
S+GNF +LNLS+CEDQM+ + ++ SGF GTSFEDTF G
Sbjct: 11 SSGNFGNLNLSTCEDQMVTMTTTTAAAATTGVSTSFLHHVMDSFPSGFEGTSFEDTFGPG 190
Query: 388 NILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGNN-AFTRDFLGLKPLSDSDILTIAGMG 446
IL+SN DI TT+ V GGNN TRDFLGL+PLS SDILT+A MG
Sbjct: 191 GILNSNHQ---------DIDSITTS---VVKGGNNEGLTRDFLGLRPLSHSDILTLASMG 334
Query: 447 SCMNPSNSNHQENHSQKPWEG 467
+CMN + Q N S KPW G
Sbjct: 335 NCMN----HDQPNPSPKPWPG 385
>CB827691
Length = 419
Score = 91.7 bits (226), Expect = 2e-19
Identities = 67/142 (47%), Positives = 81/142 (56%), Gaps = 11/142 (7%)
Frame = +3
Query: 231 IQGFTLKKEHQSFNMLRPEQEVQIPSWLCQSSIDLSS------NYSSLDQDLHLYENPNP 284
+ GF LKKE S N QE+ +P WL S +++ N SS ENPNP
Sbjct: 6 LHGFPLKKEQLSXNF---GQEI-MPPWLGPQSSTINNVDHHHHNLSSASMFSSQEENPNP 173
Query: 285 RNGPTSTLPSYQPSSAASPHMSATALLQKAAQMGA-----TSSCSSQSMMSGTHQQGHVS 339
GPT LP YQ + SPHMSATALLQKAAQMGA T++ S+ +M+S TH Q HV
Sbjct: 174 SMGPT--LPPYQ-TVPPSPHMSATALLQKAAQMGATMTTTTTAASAPAMISRTHHQPHVP 344
Query: 340 IVDSATNNMINSNGNFSLNLSS 361
+A NN +S GNF NLSS
Sbjct: 345 PHSAANNN--SSIGNFGFNLSS 404
>TC19991
Length = 540
Score = 72.4 bits (176), Expect = 1e-13
Identities = 44/161 (27%), Positives = 69/161 (42%), Gaps = 21/161 (13%)
Frame = +1
Query: 91 QLHKRGHNLPWKLKQRTSNEI-------RKKVYVCPEPTC---VHHDPSRALGDLTGIKK 140
++H R H +K + + + R++ + CP C H RAL + ++
Sbjct: 1 RMHMRAHGNQFKTAEALAKPLVEDAGGQRERRFSCPFKGCNRNKRHRRFRALKSVVCVRN 180
Query: 141 HFCRKHGEKKWKCDKCSKK--YAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRA- 197
HF R H K + CD+C K+ ++V SD K+H + C +RC CGT FSR+D H A
Sbjct: 181 HFKRSHCPKMYACDRCHKRKSFSVLSDLKSHMRHCSESRWRCSCGTTFSRKDKLFGHIAR 360
Query: 198 --------FCDALAEESSRTVIPQPTQPNSHHNMNNLQTQD 230
D + E+ V P ++N D
Sbjct: 361 FEGHVPAIALDEIGEKGKHVVEENEEDPIKEDELSNCFLND 483
>TC12248
Length = 591
Score = 67.4 bits (163), Expect = 5e-12
Identities = 37/77 (48%), Positives = 47/77 (60%), Gaps = 5/77 (6%)
Frame = +3
Query: 7 ASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPA-----KKRRNLPGNPDPQAEV 61
A+ +SA+ G R + QQ+ + + TP+ KK+RN PG P+P AEV
Sbjct: 360 AASSSSASLFGFREEDNQMNQQQQHSLVAAAPSSSATPSAPPPQKKKRNQPGTPNPDAEV 539
Query: 62 IALSPKTLMATNRFICE 78
IALSPKTLMATNRFICE
Sbjct: 540 IALSPKTLMATNRFICE 590
>BP051546
Length = 574
Score = 66.6 bits (161), Expect = 8e-12
Identities = 31/81 (38%), Positives = 43/81 (52%), Gaps = 3/81 (3%)
Frame = -2
Query: 118 CPEPTC---VHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCG 174
C P C + H ++ L D ++ H+ RKHG K + C KC K +AV+ DW+ H K CG
Sbjct: 510 CCAPGCKNNIDHPRAKPLKDFRTLQTHYKRKHGIKPFMCRKCCKAFAVRGDWRTHEKNCG 331
Query: 175 TREYRCDCGTLFSRRDSFITH 195
Y C CG+ F + S H
Sbjct: 330 KLWY-CSCGSDFKHKRSLKDH 271
>CN825130
Length = 742
Score = 37.0 bits (84), Expect = 0.007
Identities = 30/80 (37%), Positives = 39/80 (48%), Gaps = 13/80 (16%)
Frame = +1
Query: 256 SWLCQSSIDLSSNYSSLDQ--DLHLYENPNPRNGPTSTL---PSYQPS--------SAAS 302
+W+ + + N+ L L L N N N L PS S ++AS
Sbjct: 226 NWVFGNKFSSNGNHQELTSTTSLPLVNNNNIVNDGAHNLISVPSLYSSQQQHQSHQTSAS 405
Query: 303 PHMSATALLQKAAQMGATSS 322
+MSATALLQKAAQ+G TSS
Sbjct: 406 ANMSATALLQKAAQIGTTSS 465
>TC8544
Length = 1278
Score = 32.3 bits (72), Expect = 0.17
Identities = 28/95 (29%), Positives = 41/95 (42%), Gaps = 5/95 (5%)
Frame = +2
Query: 333 HQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQMINNSFSSSGFHGTSFEDTFAGNILHS 392
H +V ATN + +FS++ S+ NS SSS +F+D+F N L
Sbjct: 170 HNDDDEPVVSDATNLALIRPDHFSID-STPNSNSSCNSNSSSDEVDRNFDDSFVANQLFP 346
Query: 393 NQD-HNINHDGDNDI----PKTTTNDDDVAAGGNN 422
N + HD D+D P T D AA ++
Sbjct: 347 NPNPQRWTHDSDSDSITTDPVKETEDSASAAAASS 451
>TC17271 UP|Q7PHQ1 (Q7PHQ1) ENSANGP00000024044 (Fragment), partial (11%)
Length = 539
Score = 31.2 bits (69), Expect = 0.38
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 1/55 (1%)
Frame = +2
Query: 241 QSFNMLRPEQEVQIPSWLCQSS-IDLSSNYSSLDQDLHLYENPNPRNGPTSTLPS 294
QS N+ P Q P Q++ + LS ++ SL+ +HL NP+P+ T LP+
Sbjct: 11 QSQNLKSPSQTTTPPCRKLQTT*MVLSKHFKSLN*LIHLRPNPHPQTWSTPMLPT 175
>TC10791 similar to UP|Q8LSK7 (Q8LSK7) Auxin-regulated protein, partial
(68%)
Length = 1004
Score = 30.0 bits (66), Expect = 0.84
Identities = 31/125 (24%), Positives = 51/125 (40%), Gaps = 5/125 (4%)
Frame = +1
Query: 304 HMSATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVD-----SATNNMINSNGNFSLN 358
H T L G + +S ++G VD S+TNN++ S G+ S +
Sbjct: 55 HFEETELRLGLPLSGKEGDSTLKSTLNGKRVFSDTKTVDLKLNLSSTNNVLAS-GSSSSS 231
Query: 359 LSSCEDQMINNSFSSSGFHGTSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAA 418
+S + + + G +F NI++ + +NIN + K T A+
Sbjct: 232 VSKESRNPAKSPPAKAQVVGWPPVRSFRKNIVNVQKSNNIN-----EAEKATITSTATAS 396
Query: 419 GGNNA 423
GGNNA
Sbjct: 397 GGNNA 411
>TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (85%)
Length = 1352
Score = 30.0 bits (66), Expect = 0.84
Identities = 16/31 (51%), Positives = 21/31 (67%), Gaps = 2/31 (6%)
Frame = +1
Query: 282 PNPRNGP--TSTLPSYQPSSAASPHMSATAL 310
P+ + P TST P++ PS AA+P SATAL
Sbjct: 580 PSSTHSPPKTSTRPTWYPSLAATPSASATAL 672
>TC9284 weakly similar to UP|Q9LPY0 (Q9LPY0) T23J18.22, partial (33%)
Length = 898
Score = 30.0 bits (66), Expect = 0.84
Identities = 28/82 (34%), Positives = 37/82 (44%), Gaps = 12/82 (14%)
Frame = +1
Query: 281 NPNPRNGPTSTLPSYQ-----------PSSAASPHMSATALLQKAAQMGATSSCSSQSMM 329
+PNP + ++T PS PSS++S +S+T A+ ATSS SS
Sbjct: 193 DPNPASSASTTKPSAATSPPSKPHAPTPSSSSSTPLSSTTNPNSASPSFATSSPSSPPPS 372
Query: 330 S-GTHQQGHVSIVDSATNNMIN 350
S T H I SAT N N
Sbjct: 373 SPTTPTPPHSRISSSATLNSRN 438
>TC8349
Length = 749
Score = 30.0 bits (66), Expect = 0.84
Identities = 32/125 (25%), Positives = 49/125 (38%), Gaps = 3/125 (2%)
Frame = +2
Query: 305 MSATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNF---SLNLSS 361
+SA ++LQ+ G+ ++ + Q M + M +SNG SL S
Sbjct: 11 LSANSILQQGHSQGSQANQALQQQM-----------IQQLLQEMSHSNGGMQPQSLGGSH 157
Query: 362 CEDQMINNSFSSSGFHGTSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGN 421
M N GF G + AG+ + + + N K +N D AAGGN
Sbjct: 158 GNGNMAKNG--GVGFGGQTLR--VAGDTSNVSGSNGPVPSRSNSF-KAASNSDSSAAGGN 322
Query: 422 NAFTR 426
N F +
Sbjct: 323 NGFNQ 337
>TC9631 similar to GB|AAC19270.1|3193286|T14P8 coded for by A. thaliana
cDNA H76583 {Arabidopsis thaliana;}, partial (30%)
Length = 624
Score = 29.6 bits (65), Expect = 1.1
Identities = 20/70 (28%), Positives = 28/70 (39%), Gaps = 6/70 (8%)
Frame = +2
Query: 115 VYVCPEPTCVHHDPSRALGDLTGIKKHFCRK------HGEKKWKCDKCSKKYAVQSDWKA 168
+YVC P CVH + R G+ K+F R + + K D A + W
Sbjct: 14 LYVCDWPGCVHSEEKRKYMLFRGVFKNFKRTRVWRTINDGNRRKLDLPCAFCACKHTWDL 193
Query: 169 HSKTCGTREY 178
HS C R +
Sbjct: 194 HSAFCLRRGF 223
>TC16033 UP|Q7X9C1 (Q7X9C1) NIN-like protein 1, partial (88%)
Length = 2554
Score = 28.9 bits (63), Expect = 1.9
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +3
Query: 127 DPSRALGDLTGIKKHFCRKHGEKKWKCDKCSK 158
D ++ +G T K CR+HG K+W K K
Sbjct: 1518 DAAKNIGVCTTTLKRVCRQHGIKRWPSRKIKK 1613
>TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%)
Length = 1175
Score = 28.5 bits (62), Expect = 2.4
Identities = 20/71 (28%), Positives = 29/71 (40%), Gaps = 4/71 (5%)
Frame = +1
Query: 45 AKKRRNLPGNPDPQAEVI----ALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLP 100
A R++ N P A + ++S T C IC+K F Q L HKR H
Sbjct: 436 ASHRKSSSENQTPAAAAVNDNASVSTSTGNGGKMHECSICHKSFPTGQALGGHKRCHYEG 615
Query: 101 WKLKQRTSNEI 111
K T++ +
Sbjct: 616 GKSSNSTNSNV 648
>CB829449
Length = 561
Score = 28.1 bits (61), Expect = 3.2
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 445 MGSCMNPSNSNHQENHSQKPWEG 467
+G C P+ S +NHS KP +G
Sbjct: 467 IGHCTEPAQSQQPQNHSMKP*DG 399
>TC14204 homologue to UP|Q9AVG8 (Q9AVG8) Isopentenyl diphosphate isomerase
1, partial (79%)
Length = 1217
Score = 28.1 bits (61), Expect = 3.2
Identities = 26/95 (27%), Positives = 43/95 (44%)
Frame = -1
Query: 282 PNPRNGPTSTLPSYQPSSAASPHMSATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIV 341
PN N T T + PSS+ S +A ++ + + + S CS H +G+V+ V
Sbjct: 569 PNGVN*STGTSSAGMPSSSKSFL*AAFLTPREFSSISSDSRCSGWLQQVFVHTRGNVTFV 390
Query: 342 DSATNNMINSNGNFSLNLSSCEDQMINNSFSSSGF 376
+ SN F LN ++ + + + FS S F
Sbjct: 389 ---ADRC*RSNSYFELNRNTL--KALCSKFSDSIF 300
>AU088794
Length = 433
Score = 27.7 bits (60), Expect = 4.2
Identities = 20/61 (32%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Frame = +2
Query: 298 SSAASPHMSATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMIN-SNGNFS 356
+S A PH+S A L KAA + + ++M+ + VD+ + M++ SNGN S
Sbjct: 170 TSMACPHVSGLAALLKAAHPXWSPAAIKSALMTTAY------TVDNKGDAMLDESNGNVS 331
Query: 357 L 357
L
Sbjct: 332 L 334
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.126 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,551,959
Number of Sequences: 28460
Number of extensions: 156848
Number of successful extensions: 845
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 829
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 839
length of query: 468
length of database: 4,897,600
effective HSP length: 94
effective length of query: 374
effective length of database: 2,222,360
effective search space: 831162640
effective search space used: 831162640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148918.4


