

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
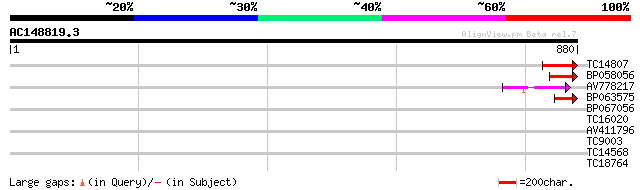
Score E
Sequences producing significant alignments: (bits) Value
TC14807 67 1e-11
BP058056 52 4e-07
AV778217 51 9e-07
BP063575 43 2e-04
BP067056 40 0.002
TC16020 similar to UP|Q9SLX0 (Q9SLX0) Importin alpha 1b, partial... 30 1.7
AV411796 30 1.7
TC9003 29 2.8
TC14568 29 2.8
TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding prote... 29 2.8
>TC14807
Length = 534
Score = 67.0 bits (162), Expect = 1e-11
Identities = 34/54 (62%), Positives = 40/54 (73%)
Frame = +1
Query: 827 KNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVSEDSSSHTTPPDS 880
K+A LMI EEIN L DEV RR+KPLL LET +D+SS S V +DSS +TPP S
Sbjct: 1 KSALLMIHEEINFLSDEVHRRLKPLLNLETAVDLSSPSVRVPKDSSFQSTPPHS 162
>BP058056
Length = 525
Score = 52.0 bits (123), Expect = 4e-07
Identities = 26/43 (60%), Positives = 31/43 (71%)
Frame = -1
Query: 838 NHLCDEVDRRIKPLLKLETVLDISSSSAPVSEDSSSHTTPPDS 880
N L DEV RR+KPLL LET +D+SS S V +DSS +TPP S
Sbjct: 525 NFLSDEVHRRLKPLLNLETAVDLSSPSVRVPKDSSFPSTPPHS 397
>AV778217
Length = 530
Score = 50.8 bits (120), Expect = 9e-07
Identities = 35/111 (31%), Positives = 49/111 (43%), Gaps = 6/111 (5%)
Frame = -3
Query: 766 APLAGCNPRVDDKHSKWLHLRIRPSALPFL------DPVKYNRHGKLKTKAFVDGRWILA 819
APL G P++D+ H LHLRIR P +P+ H DG+W L
Sbjct: 525 APLPGLRPKIDEDHPSCLHLRIREFGPPSYTITTRGNPLHMPDHSP-------DGKWTLG 367
Query: 820 FRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVSED 870
F D ++C+ A L I EI ++ + PLL+ + L S S D
Sbjct: 366 FTDAKACEEARLAILNEITKQRSALEYVLAPLLQNDPGLAES*GSLSCDHD 214
>BP063575
Length = 494
Score = 43.1 bits (100), Expect = 2e-04
Identities = 21/35 (60%), Positives = 26/35 (74%)
Frame = -1
Query: 846 RRIKPLLKLETVLDISSSSAPVSEDSSSHTTPPDS 880
RR+KPLL LET +D+SS S V +DSS +TPP S
Sbjct: 491 RRLKPLLHLETAVDLSSPSVRVPKDSSFPSTPPHS 387
>BP067056
Length = 572
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/28 (67%), Positives = 23/28 (81%)
Frame = -2
Query: 836 EINHLCDEVDRRIKPLLKLETVLDISSS 863
EIN L DEV RR+KPLL LET +D+SS+
Sbjct: 571 EINFLSDEVHRRLKPLLHLETPVDLSSA 488
>TC16020 similar to UP|Q9SLX0 (Q9SLX0) Importin alpha 1b, partial (55%)
Length = 951
Score = 30.0 bits (66), Expect = 1.7
Identities = 24/70 (34%), Positives = 39/70 (55%), Gaps = 8/70 (11%)
Frame = +3
Query: 221 LYYFSDIISAGIPDV-ERLITDSILMLLIFP---VLLPSLRTVAD----NDMQSGVVTSL 272
L Y SD + I V E + ++ LL+ P VL+P+LRTV + +DMQ+ V+ +
Sbjct: 174 LSYLSDGTNDKIQAVIEAGVCPRLVELLLHPSPSVLIPALRTVGNIVTGDDMQTQVIINH 353
Query: 273 YLLCCILRIV 282
L C+L ++
Sbjct: 354 QALPCLLNLL 383
>AV411796
Length = 365
Score = 30.0 bits (66), Expect = 1.7
Identities = 16/58 (27%), Positives = 28/58 (47%)
Frame = -1
Query: 304 KCSGGQVNGYIPDHGFTSKSDGIDNDNLAENSAKCLVVNVPCSSSSSDCHPQSITILN 361
K + GQ ++P H F+ + G L +N +C+ PC + DC + I ++N
Sbjct: 188 KSNVGQFFLHVP-HNFSLRGCGESVPTLGQNLHQCISEIAPCKVQTQDCMWKGIPLVN 18
>TC9003
Length = 613
Score = 29.3 bits (64), Expect = 2.8
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +2
Query: 729 GKERHFCFLAISVGTSGWLVLAEELP 754
GK+RHFC + V T + VL E+P
Sbjct: 389 GKKRHFCSFSCLVATRNYYVLVGEVP 466
>TC14568
Length = 1388
Score = 29.3 bits (64), Expect = 2.8
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = -1
Query: 630 EPKCMLFPPRMFFSEEDIPEGSSFTAGERMH 660
EP+ L R++F + +P+ SF+AGER H
Sbjct: 836 EPRFFLQFVRVYFFIKSVPDIESFSAGERAH 744
>TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding protein-like,
partial (35%)
Length = 848
Score = 29.3 bits (64), Expect = 2.8
Identities = 27/87 (31%), Positives = 40/87 (45%), Gaps = 1/87 (1%)
Frame = -1
Query: 308 GQVNGYIPDHGFTSKSDGIDNDNLAENSAKCLVVNVPCSSSSSDCHPQSITILNNGSSSN 367
G + + P + F+ S I N + A S PCSSSSS P S + + SSS+
Sbjct: 353 GVIFTHFPWNHFSQMSQQIQNSSAA*LSPHAPHKVSPCSSSSSSSKPSSSSSSPSPSSSS 174
Query: 368 VALREVLLEYVTG-GADVQVLGSLSVL 393
+ EY T G+D ++ L +L
Sbjct: 173 FDINP--FEYFTX*GSDPRLEFDLGLL 99
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,230,043
Number of Sequences: 28460
Number of extensions: 242256
Number of successful extensions: 1242
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1227
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1241
length of query: 880
length of database: 4,897,600
effective HSP length: 98
effective length of query: 782
effective length of database: 2,108,520
effective search space: 1648862640
effective search space used: 1648862640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148819.3


