

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148817.2 - phase: 0
(608 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
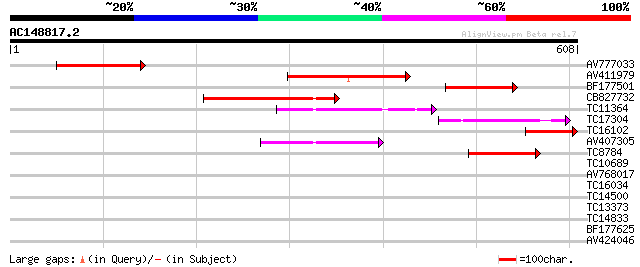
Score E
Sequences producing significant alignments: (bits) Value
AV777033 180 7e-46
AV411979 167 5e-42
BF177501 142 2e-34
CB827732 122 1e-28
TC11364 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent ... 107 4e-24
TC17304 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent ... 92 2e-19
TC16102 homologue to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzy... 82 2e-16
AV407305 81 4e-16
TC8784 similar to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic protei... 72 2e-13
TC10689 UP|Q9ATF6 (Q9ATF6) Ribosomal protein L17, complete 30 1.5
AV768017 30 1.5
TC16034 homologue to UP|FKB2_VICFA (Q41649) FK506-binding protei... 29 1.9
TC14500 UP|Q9S7B1 (Q9S7B1) Nodule INCEPTION protein, complete 29 1.9
TC13373 similar to UP|Q9ASP8 (Q9ASP8) AT3g55400/T22E16_60, parti... 27 7.3
TC14833 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (... 27 9.5
BF177625 27 9.5
AV424046 27 9.5
>AV777033
Length = 592
Score = 180 bits (456), Expect = 7e-46
Identities = 84/95 (88%), Positives = 91/95 (95%)
Frame = +2
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GFPLTERDRLGLRGLLPPR+ISF++QYDRFM+SYRSLEKNT Q +K+VSL+KWRILNRL
Sbjct: 305 GFPLTERDRLGLRGLLPPRVISFEQQYDRFMNSYRSLEKNTQCQPEKVVSLAKWRILNRL 484
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNY 145
HDRNETLYYR LIDNIKEFAPIIYTPTVGLVCQNY
Sbjct: 485 HDRNETLYYRVLIDNIKEFAPIIYTPTVGLVCQNY 589
>AV411979
Length = 408
Score = 167 bits (423), Expect = 5e-42
Identities = 86/136 (63%), Positives = 110/136 (80%), Gaps = 5/136 (3%)
Frame = +1
Query: 299 DDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCS 358
DD+QGTAGVA+AGLLG VRAQGRP+ DF KQKIV+ GAGSAG+GVLN A + ++RM G +
Sbjct: 1 DDVQGTAGVAIAGLLGAVRAQGRPMIDFPKQKIVVAGAGSAGIGVLNAARKTMARMLGNN 180
Query: 359 ETA---AKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPR--DLEGLAEGASIIEVVKKVK 413
E A AKSQF+++D GL+T R N+DP A+PFA+N + D +GL EGAS++EVVK+VK
Sbjct: 181 EAAFESAKSQFWVVDAKGLITEGRENIDPDALPFARNLKEMDRQGLREGASLVEVVKQVK 360
Query: 414 PHVLLGLSGVGGVFNE 429
P VLLGLS VGG+F++
Sbjct: 361 PDVLLGLSAVGGLFSK 408
>BF177501
Length = 247
Score = 142 bits (358), Expect = 2e-34
Identities = 69/77 (89%), Positives = 72/77 (92%)
Frame = +1
Query: 468 ENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECL 527
ENIVFASGSPFENVDLGNG+ GHVNQANNMYLFPGIGLG+LLSGA ITDGMLQAASECL
Sbjct: 16 ENIVFASGSPFENVDLGNGKVGHVNQANNMYLFPGIGLGTLLSGARLITDGMLQAASECL 195
Query: 528 ASYMTEEDIQKGILYPS 544
ASYM E+DI KGILYPS
Sbjct: 196 ASYMVEDDILKGILYPS 246
>CB827732
Length = 438
Score = 122 bits (307), Expect = 1e-28
Identities = 64/147 (43%), Positives = 89/147 (60%), Gaps = 1/147 (0%)
Frame = +2
Query: 208 MYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHAR 267
+Y A G+ P LP+ +DVGTNN++LL+D Y+GLRQ R G+EY ++ EFM AV
Sbjct: 5 LYSALGGVRPSACLPVTIDVGTNNEQLLNDEFYIGLRQKRATGKEYYDLLHEFMTAVKQN 184
Query: 268 W-PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDF 326
+ K +VQFEDF AFE L +Y +FNDDIQGTA V LAG++ ++ G L D
Sbjct: 185 YGEKVLVQFEDFANHNAFELLAKYGTTHLVFNDDIQGTASVVLAGVVAALKLIGGTLGD- 361
Query: 327 AKQKIVIVGAGSAGLGVLNMAVQAVSR 353
+ +GAG AG G+ + +S+
Sbjct: 362 --HTFLFLGAGEAGTGIAELIALEMSK 436
>TC11364 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent malic
enzyme {Phaseolus vulgaris;} , partial (27%)
Length = 490
Score = 107 bits (268), Expect = 4e-24
Identities = 63/172 (36%), Positives = 96/172 (55%), Gaps = 1/172 (0%)
Frame = +1
Query: 287 LKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNM 346
L RY +FNDDIQGTA V LAGLL +++ G L+D + +GAG AG G+ +
Sbjct: 4 LDRYSSSHLVFNDDIQGTASVVLAGLLASLKLVGGTLAD---HTFLFLGAGEAGTGIAEL 174
Query: 347 AVQAVSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASI 405
+S+ + + + +L+D GL+ R +L P+A ++ L
Sbjct: 175 IALEISKQTKAPVEETRKKVWLVDSKGLIVRSRLESLQHFKKPWAHEHEPVKAL------ 336
Query: 406 IEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAEC 457
++ VK +KP VL+G SGVG F ++V++AM S++ KP I A+SNPT +EC
Sbjct: 337 LDAVKAIKPTVLIGSSGVGKTFTKEVVEAM-ASLNEKPLILALSNPTSQSEC 489
>TC17304 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent malic
enzyme {Phaseolus vulgaris;} , partial (23%)
Length = 606
Score = 92.4 bits (228), Expect = 2e-19
Identities = 54/142 (38%), Positives = 78/142 (54%), Gaps = 1/142 (0%)
Frame = +2
Query: 461 DAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGML 520
+A+ + +FASGSPF+ V G G+ QANN Y+FPG GLG ++SGA + D ML
Sbjct: 2 EAYTWSKGKAIFASGSPFDPVKYG-GKVYAPGQANNAYIFPGFGLGLIISGAIRVRDEML 178
Query: 521 QAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKEL 580
AASE LA+ +++E+ KG++YP IR ++A + A+V GLA
Sbjct: 179 LAASEALAAQVSQENYDKGLIYPPFPNIRKISAHIAASVAAKVYELGLA----------- 325
Query: 581 EHMSE-DDTVEYVRGNMWYPEY 601
H+ D V+Y M+ P Y
Sbjct: 326 SHLPRPKDLVKYAESCMYSPGY 391
>TC16102 homologue to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzyme
(Fragment) , partial (18%)
Length = 564
Score = 82.4 bits (202), Expect = 2e-16
Identities = 38/55 (69%), Positives = 45/55 (81%)
Frame = +1
Query: 554 EVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEYCPLVHEK 608
EVGAAVLRAA+ E LA+G VG +EL HMS+++TVEYVR NMW+P Y PLVHEK
Sbjct: 1 EVGAAVLRAAVEEDLADGRDDVGPRELAHMSKEETVEYVRHNMWFPVYSPLVHEK 165
>AV407305
Length = 414
Score = 81.3 bits (199), Expect = 4e-16
Identities = 44/133 (33%), Positives = 71/133 (53%), Gaps = 1/133 (0%)
Frame = +1
Query: 270 KAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQ 329
K ++QFEDF AF+ L++YR +FNDDIQGTA V LAGL+ ++ G L+D
Sbjct: 22 KVLIQFEDFANHNAFDLLEKYRSTHLVFNDDIQGTASVVLAGLVAALKLVGGNLAD---H 192
Query: 330 KIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVP 388
+ + +GAG AG G+ + S+ + + +L+D GL+ + R +L P
Sbjct: 193 RFLFLGAGEAGTGIAELIALETSKQTNSPLEEVRKNIWLVDSKGLIVSSRKESLQHFKKP 372
Query: 389 FAKNPRDLEGLAE 401
+A ++ L +
Sbjct: 373 WAHEHEPVKNLVD 411
>TC8784 similar to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic protein (Malic
enzyme) , partial (18%)
Length = 729
Score = 72.4 bits (176), Expect = 2e-13
Identities = 35/77 (45%), Positives = 48/77 (61%)
Frame = +2
Query: 493 QANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVT 552
QANN Y+FPG GLG ++SG + D +L AASE LA +T+E KG+++P IR ++
Sbjct: 35 QANNAYIFPGFGLGLIMSGTIRVHDDLLLAASEALAEQVTQEHFDKGLIFPPFTNIRKIS 214
Query: 553 AEVGAAVLRAAIAEGLA 569
A + A V A GLA
Sbjct: 215 AHIAAKVAAKAYELGLA 265
>TC10689 UP|Q9ATF6 (Q9ATF6) Ribosomal protein L17, complete
Length = 485
Score = 29.6 bits (65), Expect = 1.5
Identities = 15/39 (38%), Positives = 19/39 (48%)
Frame = -2
Query: 479 ENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITD 517
EN L R N + +L P IGL L G HH++D
Sbjct: 301 ENPVLSAPRLALANDDSRQHLLPQIGLSLLDGGHHHVSD 185
>AV768017
Length = 314
Score = 29.6 bits (65), Expect = 1.5
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = -2
Query: 590 EYVRGNMWYPEYCPLVHEK 608
EYV+ NMW PEY LV+++
Sbjct: 304 EYVKNNMWCPEYPTLVYKQ 248
>TC16034 homologue to UP|FKB2_VICFA (Q41649) FK506-binding protein 2
precursor (Peptidyl-prolyl cis-trans isomerase)
(PPIase) (Rotamase) (15 kDa FKBP) (FKBP-15) , partial
(91%)
Length = 592
Score = 29.3 bits (64), Expect = 1.9
Identities = 13/32 (40%), Positives = 21/32 (65%), Gaps = 2/32 (6%)
Frame = -3
Query: 116 TLYYRALIDN--IKEFAPIIYTPTVGLVCQNY 145
T+ ++A I+N I EF+PI+Y + VC N+
Sbjct: 245 TISFKARIENSSISEFSPIVYFDFITFVCLNF 150
>TC14500 UP|Q9S7B1 (Q9S7B1) Nodule INCEPTION protein, complete
Length = 3001
Score = 29.3 bits (64), Expect = 1.9
Identities = 16/55 (29%), Positives = 24/55 (43%)
Frame = +3
Query: 544 SIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWY 598
S DC TAE ++ + +G GVG +E E + D +V G W+
Sbjct: 192 SSDCGSVTTAEADHHIIEELLVQGCWVEVSGVGVREGELQLQQDESSFVVGKRWW 356
>TC13373 similar to UP|Q9ASP8 (Q9ASP8) AT3g55400/T22E16_60, partial (16%)
Length = 508
Score = 27.3 bits (59), Expect = 7.3
Identities = 21/79 (26%), Positives = 34/79 (42%)
Frame = +1
Query: 384 PAAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKP 443
P PF +P E L +SII + K + H+L+ L + + + M S S P
Sbjct: 40 PYKPPFPFSPSVPELLTSTSSIICIWVKRRRHILVELCSAAPQASIPLPQTMNPSSSPTP 219
Query: 444 AIFAMSNPTMNAECTAIDA 462
+ + P M + + I A
Sbjct: 220 LYYVNAPPHMGSAYSTIAA 276
>TC14833 similar to UP|Q9ZPK4 (Q9ZPK4) Glutamyl-tRNA reductase (GluTR) ,
partial (72%)
Length = 1538
Score = 26.9 bits (58), Expect = 9.5
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = +3
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLG 342
+H+F ++N D AGL V +G+ L+ +++V VG G G G
Sbjct: 87 KHRFLLYNKDATQHIFEVSAGLDSLVLGEGQILAQV--KQVVKVGQGVTGFG 236
>BF177625
Length = 284
Score = 26.9 bits (58), Expect = 9.5
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +1
Query: 129 FAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
FAP T V L L+R+ R +YF AK +G + + YN
Sbjct: 100 FAPNTITYNVMLDVFGKAKLFRKVRRLYFMAKKQGLVDVITYN 228
>AV424046
Length = 400
Score = 26.9 bits (58), Expect = 9.5
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = -1
Query: 390 AKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVG 424
A NP+ + GL G ++EVV ++ HVL L VG
Sbjct: 367 AANPQ-VVGLLHGNQVLEVVGVIRRHVLAPLQVVG 266
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,788,946
Number of Sequences: 28460
Number of extensions: 123641
Number of successful extensions: 534
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 524
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 526
length of query: 608
length of database: 4,897,600
effective HSP length: 96
effective length of query: 512
effective length of database: 2,165,440
effective search space: 1108705280
effective search space used: 1108705280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148817.2


