

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
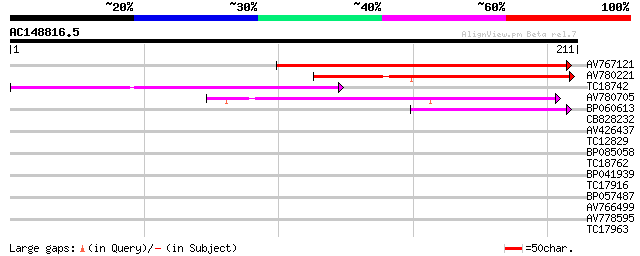
Score E
Sequences producing significant alignments: (bits) Value
AV767121 109 3e-25
AV780221 81 2e-16
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 58 1e-09
AV780705 57 2e-09
BP060613 49 5e-07
CB828232 38 0.001
AV426437 31 0.14
TC12829 28 1.5
BP085058 27 2.6
TC18762 similar to UP|Q9VVG2 (Q9VVG2) CG13731-PA, partial (3%) 26 4.5
BP041939 26 5.8
TC17916 weakly similar to PIR|B97853|B97853 NADH2 dehydrogenase ... 26 5.8
BP057487 26 5.8
AV766499 25 7.6
AV778595 25 7.6
TC17963 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zippe... 25 10.0
>AV767121
Length = 525
Score = 109 bits (273), Expect = 3e-25
Identities = 51/110 (46%), Positives = 73/110 (66%)
Frame = +2
Query: 100 LNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKK 159
L+ W + +SFGGRI L+ SVL ++P+F+LSF K+ +GV K VR+ R FLWGG K
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 160 ISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
I+ V W VC+ KE GG+ +KD+ N +LL K RWR + ++LW+ V+
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVI 331
>AV780221
Length = 461
Score = 80.9 bits (198), Expect = 2e-16
Identities = 40/100 (40%), Positives = 63/100 (63%), Gaps = 3/100 (3%)
Frame = +3
Query: 114 IVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*---FLWGGVGGGKKISWVNWKSVCQ 170
+ L+ SVL ++P+F+L F K+ G + + * F V GGKKI+WV W +VC+
Sbjct: 3 LCLIRSVLIALPLFHLPFFKLPTGGF--YTNVNN**DPFFGREVEGGKKIAWVKWSTVCR 176
Query: 171 QKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VLV 210
K+ GG+ V+++ + N +LL KWRWR++ R+ LW VL+
Sbjct: 177 PKDEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLL 296
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 58.2 bits (139), Expect = 1e-09
Identities = 38/124 (30%), Positives = 63/124 (50%)
Frame = -3
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+SHL +AD+ + ++++ + +L FQ +SGL N +KS V I V E
Sbjct: 373 ISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSE-VFILVKETTKNQIT 197
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ L G + + LG+P S + LV+ +L W+ R +S+ GR+ L+NS+
Sbjct: 196 NMLGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINSI 17
Query: 121 LNSM 124
L SM
Sbjct: 16 LFSM 5
>AV780705
Length = 524
Score = 57.0 bits (136), Expect = 2e-09
Identities = 35/135 (25%), Positives = 62/135 (45%), Gaps = 3/135 (2%)
Frame = -2
Query: 74 YLGLPV--GANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSF 131
YLGLP G N W + E + +L W ++ G+ VL+ +++ ++P + ++
Sbjct: 484 YLGLPAIWGRNKSHSLVW--IEEKVKEKLEGWKETLLNQAGKEVLIKAIIQAIPSYAMTI 311
Query: 132 LKMLVGVWKRIVRIQR*FLWGGVG-GGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLL 190
+ + + F W G G + + KE GGV +D+R N++ L
Sbjct: 310 VHFPKTFCNSLYALVADFWWKSQGEGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQNLAFL 131
Query: 191 AKWRWRLIDGREALW 205
A+ WR++ EALW
Sbjct: 130 ARQAWRVLTNPEALW 86
>BP060613
Length = 378
Score = 49.3 bits (116), Expect = 5e-07
Identities = 20/60 (33%), Positives = 33/60 (54%)
Frame = +1
Query: 150 LWGGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+W G + + W W + + K+ GG+ KD + N +LLAK WR++ +ALW +L
Sbjct: 43 VWWSSKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQIL 222
>CB828232
Length = 532
Score = 38.1 bits (87), Expect = 0.001
Identities = 22/50 (44%), Positives = 30/50 (60%)
Frame = +1
Query: 103 WSTRYISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWG 152
W T +S G R+ VL+S+P+F+LSF K+ V K +IQ FLWG
Sbjct: 388 WKTLCLS-GARVCCFKMVLSSLPLFFLSFFKIKC-VAKVCKQIQSRFLWG 531
>AV426437
Length = 303
Score = 31.2 bits (69), Expect = 0.14
Identities = 10/24 (41%), Positives = 19/24 (78%)
Frame = +1
Query: 158 KKISWVNWKSVCQQKENGGVRVKD 181
+KI+W+NW+++C+ K G+ VK+
Sbjct: 223 RKIAWINWETICKPKWLRGLGVKN 294
>TC12829
Length = 448
Score = 27.7 bits (60), Expect = 1.5
Identities = 8/21 (38%), Positives = 12/21 (57%)
Frame = +1
Query: 149 FLWGGVGGGKKISWVNWKSVC 169
F+W G + I W+ WK +C
Sbjct: 385 FIWSGDVSRRSIHWLGWKKLC 447
>BP085058
Length = 452
Score = 26.9 bits (58), Expect = 2.6
Identities = 20/105 (19%), Positives = 45/105 (42%)
Frame = +2
Query: 73 KYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFL 132
KY+ P+ + ++ + RL ++ R+ L VL+++ ++ +
Sbjct: 134 KYMSFPIHQGHV*KEDFVFTLDRVISRLAC*KAYLLNKPARLTLAKPVLSAISMYVMQLN 313
Query: 133 KMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGV 177
+ + +I + R F+W V G I V W+++ +K G+
Sbjct: 314 WLPQSICDQICTVVRHFVWENVNG---IPLVAWETIQSRKSMVGL 439
>TC18762 similar to UP|Q9VVG2 (Q9VVG2) CG13731-PA, partial (3%)
Length = 759
Score = 26.2 bits (56), Expect = 4.5
Identities = 17/54 (31%), Positives = 23/54 (42%)
Frame = -3
Query: 152 GGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALW 205
GG GGG W W+ C+ R R++ + +WRWR GR W
Sbjct: 301 GGGGGG*GSCWW-WRFRCR-------RWWRRRILRLRRRRRWRWRRRCGRWKWW 164
>BP041939
Length = 392
Score = 25.8 bits (55), Expect = 5.8
Identities = 15/46 (32%), Positives = 30/46 (64%), Gaps = 3/46 (6%)
Frame = +1
Query: 105 TRYISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRI---VRIQR 147
+R+I G R+VLL+ +LN + + +L L+ L+ +W + V+++R
Sbjct: 196 SRHIVAGRRVVLLHRILN-LYLHFLWCLQTLMIIWTSL*LYVKVKR 330
>TC17916 weakly similar to PIR|B97853|B97853 NADH2 dehydrogenase
(ubiquinone) - Rickettsia conorii
(strain Malish 7) {Rickettsia conorii;} , partial (3%)
Length = 520
Score = 25.8 bits (55), Expect = 5.8
Identities = 12/46 (26%), Positives = 24/46 (52%)
Frame = -2
Query: 118 NSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWV 163
N + + F+LSF +L GV +V + + +WG ++ +W+
Sbjct: 384 NKKIKPLYSFFLSFKFLLSGVEITLVSLIKTSIWGKEKNEEQFAWI 247
>BP057487
Length = 512
Score = 25.8 bits (55), Expect = 5.8
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +2
Query: 76 GLPVGANMRSMSTWEPLVETIGGRLNTWSTR 106
G G +M +MS ++ + +GG L TWS +
Sbjct: 287 GAANGISMTAMSLFKAIGPAVGGALLTWSQK 379
>AV766499
Length = 378
Score = 25.4 bits (54), Expect = 7.6
Identities = 10/26 (38%), Positives = 13/26 (49%)
Frame = -1
Query: 152 GGVGGGKKISWVNWKSVCQQKENGGV 177
G GG ++W N VC NGG+
Sbjct: 345 GWTSGGVSVAWSNAVRVCSLSWNGGM 268
>AV778595
Length = 594
Score = 25.4 bits (54), Expect = 7.6
Identities = 17/71 (23%), Positives = 30/71 (41%), Gaps = 1/71 (1%)
Frame = -1
Query: 99 RLNTWSTRYISFG-GRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGG 157
R+ W R +S ++ ++ S + L+ + VWK+ R + GGVG G
Sbjct: 447 RIMAWPPRLVSLDFNKLWMMGRRSWSREVRLLAAAASVTKVWKKRRECSREAVEGGVGEG 268
Query: 158 KKISWVNWKSV 168
+ +W V
Sbjct: 267 DRREMESWSLV 235
>TC17963 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zipper protein
HAT5 (HD-ZIP protein 5) (HD-ZIP protein ATHB-1), partial
(24%)
Length = 738
Score = 25.0 bits (53), Expect = 10.0
Identities = 12/29 (41%), Positives = 16/29 (54%), Gaps = 2/29 (6%)
Frame = -3
Query: 152 GGVGGGKKISWVNWKSV--CQQKENGGVR 178
G V GGK +SW K + C+ + GG R
Sbjct: 127 GRVKGGKGVSWNQEKGIRGCRNQGKGGCR 41
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,386,255
Number of Sequences: 28460
Number of extensions: 64140
Number of successful extensions: 432
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 429
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 430
length of query: 211
length of database: 4,897,600
effective HSP length: 86
effective length of query: 125
effective length of database: 2,450,040
effective search space: 306255000
effective search space used: 306255000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148816.5


