

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148655.10 + phase: 0
(250 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
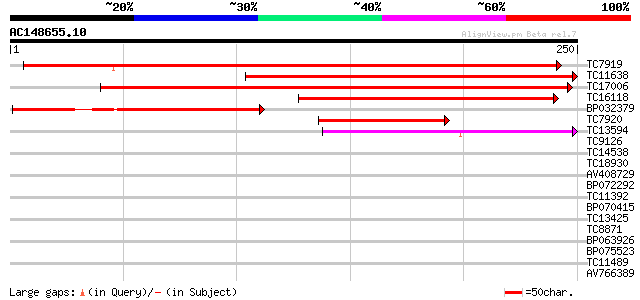
TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus cornicula... 278 8e-76
TC11638 weakly similar to GB|AAP47493.1|32331931|AY164919 RTNLB2... 238 7e-64
TC17006 similar to GB|AAP47434.1|32331813|AY164860 RTNLB33w {Gly... 180 2e-46
TC16118 similar to GB|AAP47508.1|32331961|AY164934 RTNLB5 {Medic... 137 1e-33
BP032379 128 1e-30
TC7920 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus cornicula... 74 2e-14
TC13594 homologue to GB|AAP47511.1|32331967|AY164937 RTNLB18w {M... 59 1e-09
TC9126 30 0.51
TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization ... 27 3.3
TC18930 27 3.3
AV408729 27 3.3
BP072292 27 4.3
TC11392 similar to UP|Q8BGI0 (Q8BGI0) Sterol-C5-desaturase, part... 27 4.3
BP070415 26 7.4
TC13425 weakly similar to UP|Q7XJL5 (Q7XJL5) At2g17100 protein, ... 26 7.4
TC8871 similar to UP|AAN73331 (AAN73331) ALS7 protein (Fragment)... 26 7.4
BP063926 26 7.4
BP075523 26 7.4
TC11489 25 9.7
AV766389 25 9.7
>TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus corniculatus var.
japonicus;} , complete
Length = 1219
Score = 278 bits (710), Expect = 8e-76
Identities = 136/238 (57%), Positives = 177/238 (74%), Gaps = 1/238 (0%)
Frame = +3
Query: 7 ESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAK-SNTVYRLFGRDRPVHKVLGGG 65
ES +KI +K HG DS S SDS+++ P + + V RLFGR++P+H VLGGG
Sbjct: 135 ESLMEKIAEKIHGHDSSSSSDSDNEKEKEKSSSSPSSDFKSKVSRLFGREKPIHHVLGGG 314
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFI 125
K AD+ LWRNKK + LG T +W+ FEL++Y+L+TL+CHL ILAL LFLWSN S I
Sbjct: 315 KQADVFLWRNKKISGTVLGVATVVWILFELLEYHLLTLICHLAILALTLLFLWSNGSSLI 494
Query: 126 HKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVI 185
+KSP +IP V IP+E V + AS LR +IN+ FA+LR+I +G+D+KKFL+VIAGLW SV+
Sbjct: 495 NKSPPKIPQVHIPEEPVLQLASALRADINRAFAVLRDIASGKDLKKFLSVIAGLWVFSVV 674
Query: 186 GSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
GS NFLTLFYI +V L T+P++YEK EDQVD++ EKA+ E KKQYAV DA+VLS+IP
Sbjct: 675 GSWTNFLTLFYIAFVLLHTVPVLYEKYEDQVDSIGEKALHEFKKQYAVFDAQVLSKIP 848
>TC11638 weakly similar to GB|AAP47493.1|32331931|AY164919 RTNLB21w {Vitis
vinifera;} , partial (88%)
Length = 652
Score = 238 bits (607), Expect = 7e-64
Identities = 114/146 (78%), Positives = 129/146 (88%)
Frame = +1
Query: 105 CHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIG 164
CHL IL+LA LFLWSNASVFIHKSPL IPH+ IP++C + AS +R+EINQ F +LR+IG
Sbjct: 1 CHLSILSLAGLFLWSNASVFIHKSPLHIPHIAIPEDCALQVASAIRVEINQAFVVLRQIG 180
Query: 165 TGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAM 224
TGRDIKKFL+VI LW +SV+G+ FNFLTLFYIF+VSLFTLPL+YEK EDQVDALAEKAM
Sbjct: 181 TGRDIKKFLSVIGILWVVSVVGNWFNFLTLFYIFFVSLFTLPLIYEKYEDQVDALAEKAM 360
Query: 225 IEIKKQYAVLDAKVLSQIPIAGFKKD 250
IEIKKQYAVLDAKVLSQIPI G KKD
Sbjct: 361 IEIKKQYAVLDAKVLSQIPIPGLKKD 438
>TC17006 similar to GB|AAP47434.1|32331813|AY164860 RTNLB33w {Glycine max;}
, partial (98%)
Length = 1088
Score = 180 bits (456), Expect = 2e-46
Identities = 88/208 (42%), Positives = 134/208 (64%)
Frame = +1
Query: 41 PLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNL 100
P + S+ RLFGR +PVH +LGGGK AD+LLWRNKK +A L TA+WV +E + YN
Sbjct: 178 PNSVSSQFNRLFGRQKPVHHLLGGGKSADVLLWRNKKISAGVLTGATAIWVVWEWLNYNF 357
Query: 101 ITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAIL 160
+TL+ + L + A FLW+NAS + + ++P V+P++ + A+ + E+N+ +L
Sbjct: 358 LTLLFLALALGMLAQFLWANASGVLGRKSSKVPRFVLPEDFIVNIATAVGFEVNRGLRVL 537
Query: 161 REIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALA 220
++ G ++K FL V+ LW +VIGS NFLT+ YI +V+ T P++YE+ EDQVD
Sbjct: 538 QDASCGGNLKPFLLVVVSLWAGAVIGSWCNFLTVMYIGFVAAHTFPVLYERYEDQVDDFV 717
Query: 221 EKAMIEIKKQYAVLDAKVLSQIPIAGFK 248
K + +++ QY LD+ +LS+IP FK
Sbjct: 718 YKFLGQMRNQYQKLDSGLLSKIPHGKFK 801
>TC16118 similar to GB|AAP47508.1|32331961|AY164934 RTNLB5 {Medicago
truncatula;} , partial (48%)
Length = 721
Score = 137 bits (346), Expect = 1e-33
Identities = 68/115 (59%), Positives = 86/115 (74%)
Frame = +1
Query: 128 SPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGS 187
+P +IP +P+E E AS LRIEIN FA LR+IGTG+D+KKFL I GLW +S++GS
Sbjct: 1 APPRIPVFHLPEEPFLEIASALRIEINHGFAALRDIGTGKDVKKFLIAIVGLWILSIVGS 180
Query: 188 CFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
NFLTLFYI +V L T+P +Y+K ED++D LAEKA IKKQY V DAKVLS++
Sbjct: 181 WCNFLTLFYINFVLLLTIPFLYDKYEDKIDPLAEKAAHAIKKQYVVFDAKVLSKV 345
>BP032379
Length = 441
Score = 128 bits (321), Expect = 1e-30
Identities = 62/111 (55%), Positives = 82/111 (73%)
Frame = +2
Query: 2 AAVDTESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKV 61
AA ES DKI DK HG DS S SDSE +P + K ++RLFGR++P+H V
Sbjct: 125 AAATGESLLDKISDKLHGHDSSSDSDSE-------KPSLSSVKQK-IFRLFGREKPLHSV 280
Query: 62 LGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILAL 112
LGGGKPAD+LLW+NKK +A ALG TA+W+FFEL++Y+L+TLVCH++IL++
Sbjct: 281 LGGGKPADVLLWKNKKVSAGALGVATAVWLFFELLEYHLLTLVCHILILSI 433
>TC7920 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus corniculatus var.
japonicus;} , partial (24%)
Length = 618
Score = 74.3 bits (181), Expect = 2e-14
Identities = 34/58 (58%), Positives = 46/58 (78%)
Frame = +1
Query: 137 IPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTL 194
IP+E V + AS LR +IN+ FA+LR+I +G+D+KKFL+VIAGLW SV+GS NFL +
Sbjct: 1 IPEEPVLQLASALRADINRAFAVLRDIASGKDLKKFLSVIAGLWVFSVVGSWTNFLDI 174
>TC13594 homologue to GB|AAP47511.1|32331967|AY164937 RTNLB18w {Medicago
truncatula;} , partial (41%)
Length = 556
Score = 58.5 bits (140), Expect = 1e-09
Identities = 37/113 (32%), Positives = 53/113 (46%), Gaps = 1/113 (0%)
Frame = +1
Query: 139 QECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYI- 197
+E V A+ R++IN V I +I G+D + F V+ LW +SVIGS F+F TL YI
Sbjct: 1 EEMVNNVAASFRVKINNVLLIAHDITIGQDFRVFFKVVVCLWLLSVIGSYFSFFTLAYIG 180
Query: 198 FYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIPIAGFKKD 250
+ P K + VD K Y ++D V +Q + KD
Sbjct: 181 SLYDDYDXPRCTRKYGNYVDKCCGILPPRFSKPYRIVDESVFNQXCPIMYPKD 339
>TC9126
Length = 1013
Score = 29.6 bits (65), Expect = 0.51
Identities = 11/18 (61%), Positives = 15/18 (83%)
Frame = -1
Query: 187 SCFNFLTLFYIFYVSLFT 204
S FNF L+Y+FY+SLF+
Sbjct: 872 SLFNFSLLYYLFYISLFS 819
>TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization 1 homolog a
{Mus musculus;}, partial (3%)
Length = 2631
Score = 26.9 bits (58), Expect = 3.3
Identities = 19/68 (27%), Positives = 26/68 (37%)
Frame = -3
Query: 37 RPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELM 96
RP L NT R+F P IL W NK+ + +G W+ E +
Sbjct: 928 RPFAKLRAGNTAIRVFP----------WVSPPCILPWMNKRTLSPVMGVSP*AWLHLETV 779
Query: 97 QYNLITLV 104
LIT +
Sbjct: 778 HPELITSI 755
>TC18930
Length = 726
Score = 26.9 bits (58), Expect = 3.3
Identities = 16/36 (44%), Positives = 23/36 (63%)
Frame = +1
Query: 95 LMQYNLITLVCHLMILALAALFLWSNASVFIHKSPL 130
L+Q+ L+ L+ L+IL + LFL +AS F SPL
Sbjct: 457 LLQWLLVILIPRLIIL*MWHLFLSLSASSFDFHSPL 564
>AV408729
Length = 433
Score = 26.9 bits (58), Expect = 3.3
Identities = 15/48 (31%), Positives = 25/48 (51%)
Frame = -3
Query: 116 FLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREI 163
F W N ++ IH V+PQECVF + +I +N V+ + ++
Sbjct: 167 FYW-NPALVIH---------VLPQECVFN*ILITKIVVNLVYYLSMQL 54
>BP072292
Length = 325
Score = 26.6 bits (57), Expect = 4.3
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +1
Query: 17 FHGSDSDSFSDSEDDHNINNRPPIPLAK 44
F ++S+ ++ + NIN +PPIPL K
Sbjct: 199 FQ*NNSNEITELRKE*NINYKPPIPLTK 282
>TC11392 similar to UP|Q8BGI0 (Q8BGI0) Sterol-C5-desaturase, partial (5%)
Length = 579
Score = 26.6 bits (57), Expect = 4.3
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +3
Query: 171 KFLTVIAGLWFISVIGSCFNFLTLFYIFYVS 201
+FL + W ++GSC+ F+T+F+I +S
Sbjct: 294 RFLCSLQCWW---ILGSCWRFITVFHILILS 377
>BP070415
Length = 410
Score = 25.8 bits (55), Expect = 7.4
Identities = 13/39 (33%), Positives = 23/39 (58%)
Frame = -2
Query: 181 FISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDAL 219
F++ +G+ F+F L+ I F VYEK E+++D +
Sbjct: 196 FLASLGA-FSFEDLYTIGLAFAFIAFYVYEKKEEEIDGI 83
>TC13425 weakly similar to UP|Q7XJL5 (Q7XJL5) At2g17100 protein, partial
(6%)
Length = 538
Score = 25.8 bits (55), Expect = 7.4
Identities = 11/21 (52%), Positives = 15/21 (71%), Gaps = 3/21 (14%)
Frame = +1
Query: 22 SDSF---SDSEDDHNINNRPP 39
SD F S+S+DDH+ +N PP
Sbjct: 328 SDEFPLHSESDDDHDADNEPP 390
>TC8871 similar to UP|AAN73331 (AAN73331) ALS7 protein (Fragment), partial
(5%)
Length = 1290
Score = 25.8 bits (55), Expect = 7.4
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = -1
Query: 10 DDKIEDKFHGSDSDSFSDSEDDHNINNR 37
D +E FH S S+SD +H I NR
Sbjct: 861 DK*VEQHFHNRTSCSYSDLHKNH*IGNR 778
>BP063926
Length = 487
Score = 25.8 bits (55), Expect = 7.4
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = +2
Query: 4 VDTESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVY 49
VD D ++ SD+ S SDS+DD N PP+ A Y
Sbjct: 119 VDDSDSDFEVSSDATLSDTASHSDSDDDPQTQN-PPVAEADPEAHY 253
>BP075523
Length = 466
Score = 25.8 bits (55), Expect = 7.4
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = -2
Query: 93 FELMQYNLITLVCHLMILAL 112
FEL+Q NL VC+L +L++
Sbjct: 225 FELLQPNLYRFVCYLFLLSI 166
>TC11489
Length = 402
Score = 25.4 bits (54), Expect = 9.7
Identities = 20/74 (27%), Positives = 33/74 (44%)
Frame = +1
Query: 93 FELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIE 152
FEL+ L+++ L L L + W N F ++IP V+P C+ + V
Sbjct: 64 FELLHVKLLSV*RFL--LRLFFFYKWCNKVNFK*PVDMEIPSPVLPFLCIIIIS*VPCFS 237
Query: 153 INQVFAILREIGTG 166
+++F L E G
Sbjct: 238 SSEIFPYLIEFVDG 279
>AV766389
Length = 511
Score = 25.4 bits (54), Expect = 9.7
Identities = 13/40 (32%), Positives = 22/40 (54%)
Frame = -3
Query: 204 TLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
+LP++Y+K +DQ++ QY + VLS+IP
Sbjct: 491 SLPVLYDKYQDQINDKLYLVHGIYLTQYKTIHRVVLSKIP 372
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,381,120
Number of Sequences: 28460
Number of extensions: 61932
Number of successful extensions: 558
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 553
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 556
length of query: 250
length of database: 4,897,600
effective HSP length: 88
effective length of query: 162
effective length of database: 2,393,120
effective search space: 387685440
effective search space used: 387685440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148655.10


