

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
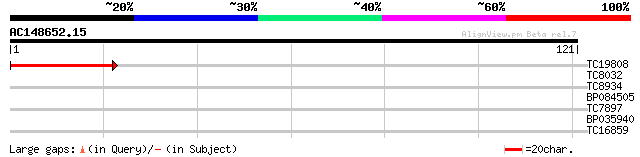
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19808 homologue to UP|Q9LH76 (Q9LH76) Genomic DNA, chromosome ... 43 1e-05
TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-... 37 0.001
TC8934 similar to GB|AAQ22601.1|33589670|BT010132 At2g27890 {Ara... 26 2.1
BP084505 25 3.6
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 25 4.7
BP035940 25 4.7
TC16859 weakly similar to UP|Q8W3M4 (Q8W3M4) Extensin-like prote... 24 8.1
>TC19808 homologue to UP|Q9LH76 (Q9LH76) Genomic DNA, chromosome 3, BAC
clone: T21E2 (AT3g14790/T21E2_4), partial (19%)
Length = 515
Score = 43.1 bits (100), Expect = 1e-05
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +2
Query: 1 MHFAAQTHVDNSYENSFEFTQTS 23
MHFAAQTHVDNS+ NSFEFT+ +
Sbjct: 395 MHFAAQTHVDNSFGNSFEFTKNN 463
>TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-like
protein, partial (97%)
Length = 2230
Score = 37.0 bits (84), Expect = 0.001
Identities = 26/106 (24%), Positives = 42/106 (39%)
Frame = +3
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ L++ S N + S+ YG ++
Sbjct: 663 MHLAAQAGVRYAMENPHSYVHSNIAALVTLMEACKSANPQPAIVWASSSSVYGLNEKVPF 842
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ Q + Y+A K E + +Y YGL + R VY
Sbjct: 843 SESDRTDQ--PASLYAATKKAGEEITHTYNHIYGLSITGLRFFTVY 974
>TC8934 similar to GB|AAQ22601.1|33589670|BT010132 At2g27890 {Arabidopsis
thaliana;}, partial (9%)
Length = 601
Score = 25.8 bits (55), Expect = 2.1
Identities = 19/76 (25%), Positives = 32/76 (42%), Gaps = 5/76 (6%)
Frame = -3
Query: 3 FAAQTHVDNSYENSFEFTQTSFMELMSFR-----SLQVSKNQVKGFIHVSTDKFYGETDE 57
F + VD S +F + + FR SL S N ++ +H+ K D+
Sbjct: 251 FHSSNKVDQSNNLAFTVFRVCSCPVYIFRLGKVISLNESFNHLQLGLHIQRHKMM*PIDQ 72
Query: 58 NAVVGNHEASQLLETN 73
N + HE+S + T+
Sbjct: 71 NLQIQ*HESSSFITTS 24
>BP084505
Length = 291
Score = 25.0 bits (53), Expect = 3.6
Identities = 15/51 (29%), Positives = 23/51 (44%)
Frame = +3
Query: 34 QVSKNQVKGFIHVSTDKFYGETDENAVVGNHEASQLLETNPYSAMKIGAEM 84
+VS ++ HVS FY +T + N S L+ Y+ KI E+
Sbjct: 93 KVSFHRAPNLYHVSAHNFYLQTSVTNMKTNKNKSLKLKIRSYALAKI*KEI 245
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein 2
precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 24.6 bits (52), Expect = 4.7
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -3
Query: 22 TSFMELMSFRSLQVSKNQVKGFI 44
T F+ ++F +LQ SKN +KG +
Sbjct: 1468 TMFLYFLTFPTLQQSKNIIKGSV 1400
>BP035940
Length = 396
Score = 24.6 bits (52), Expect = 4.7
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +2
Query: 16 SFEFTQTSFMELMSFRS 32
SF+FT +SF+ L SFR+
Sbjct: 329 SFDFTASSFVALSSFRT 379
>TC16859 weakly similar to UP|Q8W3M4 (Q8W3M4) Extensin-like protein,
partial (13%)
Length = 745
Score = 23.9 bits (50), Expect = 8.1
Identities = 8/20 (40%), Positives = 15/20 (75%)
Frame = +2
Query: 42 GFIHVSTDKFYGETDENAVV 61
GF++++ +KFYG+ E+ V
Sbjct: 2 GFLNLAQNKFYGKVPESVCV 61
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,675,464
Number of Sequences: 28460
Number of extensions: 14743
Number of successful extensions: 64
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 64
length of query: 121
length of database: 4,897,600
effective HSP length: 79
effective length of query: 42
effective length of database: 2,649,260
effective search space: 111268920
effective search space used: 111268920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 49 (23.5 bits)
Medicago: description of AC148652.15


