

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.13 - phase: 0 /pseudo
(395 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
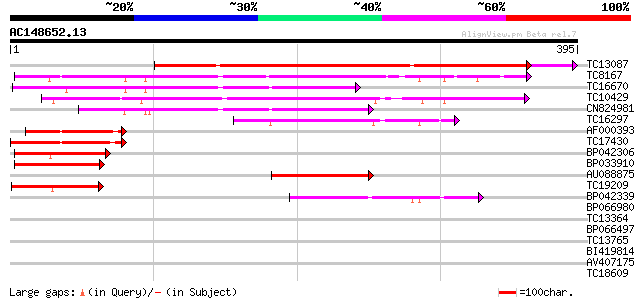
Score E
Sequences producing significant alignments: (bits) Value
TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like prote... 275 5e-77
TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine prote... 216 7e-57
TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 208 1e-54
TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%) 199 5e-52
CN824981 177 4e-45
TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type pro... 100 3e-22
AF000393 83 7e-17
TC17430 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like pro... 80 8e-16
BP042306 73 9e-14
BP033910 72 1e-13
AU088875 67 7e-12
TC19209 similar to GB|BAB11244.1|10177874|AB010074 serine protea... 65 3e-11
BP042339 65 3e-11
BP066980 35 0.017
TC13364 similar to UP|Q9LUM3 (Q9LUM3) Subtilisin proteinase-like... 33 0.083
BP066497 30 0.91
TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine end... 30 0.91
BI419814 29 1.6
AV407175 29 1.6
TC18609 weakly similar to UP|Q7YRB0 (Q7YRB0) Block of proliferat... 28 3.5
>TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like protein, partial
(31%)
Length = 1036
Score = 275 bits (703), Expect(2) = 5e-77
Identities = 145/263 (55%), Positives = 183/263 (69%), Gaps = 1/263 (0%)
Frame = +2
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAF 161
GH +HTASTA G V SFYG A+GTARG VPS+RIAAYK+C N C ILAAF
Sbjct: 38 GHGSHTASTAAGNTVIGASFYGLAQGTARGAVPSARIAAYKVC--YPNHGCDDANILAAF 211
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGA 221
DDAI+DGV++++VSLG + D L+D +AIGSFHAME+GILT QAAGN GP S+CS A
Sbjct: 212 DDAIADGVNILSVSLGTNYCVDLLSDTVAIGSFHAMERGILTLQAAGNSGPASPSICSVA 391
Query: 222 PWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASP 281
PWL++VAA++IDRQFIDKV+LGNG T +G+S+N SNG K PIA NA G +
Sbjct: 392 PWLLTVAASTIDRQFIDKVLLGNGMTLIGRSVNAVVSNGTKFPIAKWNASDERCKGFS-- 565
Query: 282 EMCDCIDENMVKGKLVLCGSRNGKDLAYANGAIGSI-HNVTKSQLGASFVTPRPSLNLKT 340
++C C+ VKGK+VLC R + AY +GA+GSI ++T SFVT PS+NL
Sbjct: 566 DLCQCLGSESVKGKVVLCEGRMHDEAAYLDGAVGSITEDLTNPDNDVSFVTYFPSVNLNP 745
Query: 341 NDFVHIQSYTNSSKYPVADFLFS 363
D+ +Q+YTNS+K P A+ L S
Sbjct: 746 KDYAIVQNYTNSAKNPKAEILKS 814
Score = 29.6 bits (65), Expect(2) = 5e-77
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +1
Query: 361 LFSWPKSFGPINYETRYKCPRSGNLSRIFTSSVTL 395
L W KS ++ET +CPR G L + T S TL
Sbjct: 856 LIPWAKSLDSGDHETGCECPRGGYLGCVVT*SGTL 960
>TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (65%)
Length = 2760
Score = 216 bits (549), Expect = 7e-57
Identities = 148/400 (37%), Positives = 208/400 (52%), Gaps = 40/400 (10%)
Frame = +3
Query: 4 VISVFPSEEFHIQTTRSWDFLGL---------RQSIKRDQIIETDLVIGVIDIRIWPESE 54
V+ ++ +H+ TTR+ FLGL ++++ DQ D++IGV+D +WPES
Sbjct: 357 VLGLYEDTLYHLHTTRTPQFLGLDTQTGLWEGHRTLELDQA-SRDVIIGVLDTGVWPESP 533
Query: 55 SFNDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDG----------------DV 95
SFND G+ IP +WRG C +FS CN K+IGAR + G
Sbjct: 534 SFNDAGMPEIPSRWRGECENATDFSSSLCNRKLIGARSFSRGFHMAAGNDGGFGKEREPP 713
Query: 96 SARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGD 155
S R S GH THTASTA G V + S G+A GTARG P +R+A YK+C + C
Sbjct: 714 SPRDSDGHGTHTASTAAGSHVGNASLLGYASGTARGMAPQARVATYKVCW---SDGCFAS 884
Query: 156 AILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPS 215
ILA D AI DGVDV+++SLG + + D IAIG+F AME+GI + +AGN GP +
Sbjct: 885 DILAGMDRAIRDGVDVLSLSLGGG-SGPYFRDTIAIGAFAAMERGIFVSCSAGNSGPSKA 1061
Query: 216 SVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLA 275
S+ + APWL++V A ++DR F +LGN K F G S+ G + P+ + ++
Sbjct: 1062SLANVAPWLMTVGAGTLDRDFPASALLGNKKRFAGVSLYSGKGMGAE-PVGLVYSK---- 1226
Query: 276 GGNASPEMC--DCIDENMVKGKLVLCGS------RNGKDLAYANGAIGSIHNVTKSQ--- 324
G N S +C +D +V+GK+VLC GK + A G IG I T +
Sbjct: 1227GSNQSGILCLPGSLDPAVVRGKVVLCDRGLNARVEKGKVVKEA-GGIGMILTNTAANGEE 1403
Query: 325 -LGASFVTPRPSLNLKTNDFVHIQSYTNSSKYPVADFLFS 363
+ S + P ++ D I+ Y S P A FS
Sbjct: 1404LVADSHLLPAVAVGRIVGD--QIREYVTSDPNPTAVLSFS 1517
>TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (42%)
Length = 1183
Score = 208 bits (530), Expect = 1e-54
Identities = 116/260 (44%), Positives = 157/260 (59%), Gaps = 18/260 (6%)
Frame = +2
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIET--DLVIGVIDIRIWPESESFNDKG 60
GV++V P +F + TTR+ FLGL +S ++++GV+D +WPES+SF+D G
Sbjct: 416 GVLAVLPERKFELHTTRTPLFLGLDKSADMFPASGAGGEVIVGVLDTGVWPESKSFDDTG 595
Query: 61 LGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDG-------------DVSARYSFGHR 104
GP+P W+G C G NF+ CN K+IGAR++ G S R GH
Sbjct: 596 FGPVPSTWKGECESGTNFTAANCNKKLIGARYFSKGAEAMLGPIDETAESKSPRDDDGHG 775
Query: 105 THTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDA 164
THTASTA G V S +G+A GTARG P +R+AAYK+C G C ILAA D A
Sbjct: 776 THTASTAAGSVVSGASLFGYAAGTARGMAPHARVAAYKVCW---KGGCFSSDILAAIDKA 946
Query: 165 ISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWL 224
I+D V+V+++SLG A D+ D +AIG+F AMEKGIL + +AGN GP S+ + APW+
Sbjct: 947 IADNVNVLSLSLGGGMA-DYYRDSVAIGAFAAMEKGILVSCSAGNAGPSAYSLSNVAPWI 1123
Query: 225 VSVAATSIDRQFIDKVILGN 244
+V A ++DR F V LGN
Sbjct: 1124TTVGAGTLDRDFPALVELGN 1183
>TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%)
Length = 1742
Score = 199 bits (507), Expect = 5e-52
Identities = 136/366 (37%), Positives = 192/366 (52%), Gaps = 26/366 (7%)
Frame = +3
Query: 23 FLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFNDKGLGPIPKKWRGVCAGGGNFSC 80
FLGL+Q + ++ ++IGV+D I P SF+D G+ P P KW+G C +C
Sbjct: 414 FLGLQQDTGVWKESNFGKGVIIGVLDSGITPGHPSFSDVGIPPPPPKWKGRCDLNVT-AC 590
Query: 81 NNKIIGARFY--------GDGDVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGG 132
NNK+IGAR + G + GH THTASTA G V G AKGTA G
Sbjct: 591 NNKLIGARAFNLAAEAMNGKKAEAPIDEDGHGTHTASTAAGAFVNYAEVLGNAKGTAAGM 770
Query: 133 VPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIG 192
P + +A YK+C D C ILAA D A+ DGVDVI++SLG F ND AIG
Sbjct: 771 APHAHLAIYKVCFGED---CPESDILAALDAAVEDGVDVISISLGLSEPPPFFNDSTAIG 941
Query: 193 SFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKS 252
+F AM+KGI + AAGN GP SS+ + APW+++V A++IDR+ + LGNG+ F G+S
Sbjct: 942 AFAAMQKGIFVSCAAGNSGPFNSSIVNAAPWILTVGASTIDRRIVATAKLGNGQEFDGES 1121
Query: 253 I----NITPSNGKKIPIAVRNAQACLAGGNASPEMCDC----IDENMVKGKLVLCGS--- 301
+ + TP+ +P+A AG N E C +D++ +GK+VLC
Sbjct: 1122VFQPSSFTPT---LLPLA-------YAGKNGKEESAFCANGSLDDSAFRGKVVLCERGGG 1271
Query: 302 ----RNGKDLAYANGAIGSIHNVTKSQLGASF-VTPRPSLNLKTNDFVHIQSYTNSSKYP 356
G+++ A GA + N + S V P+ ++ + I++Y NS+ P
Sbjct: 1272IARIAKGEEVKRAGGAAMILMNDETNAFSLSADVHALPATHVSYAAGIEIKAYINSTATP 1451
Query: 357 VADFLF 362
A LF
Sbjct: 1452TATILF 1469
>CN824981
Length = 726
Score = 177 bits (448), Expect = 4e-45
Identities = 99/221 (44%), Positives = 128/221 (57%), Gaps = 16/221 (7%)
Frame = +1
Query: 49 IWPESESFNDKGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGDG--------DVS- 96
+WPE +S +D GL P+P W+G C G N SCN K+IGARF+ G DVS
Sbjct: 7 VWPELKSLDDTGLSPVPSTWKGQCEAGNNMNSSSCNRKLIGARFFSKGYEATLGPIDVST 186
Query: 97 ----ARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRC 152
AR GH +HT +TA G V S +G A GTARG +R+AAYK+C G C
Sbjct: 187 ESRSARDDDGHGSHTLTTAAGSAVAGASLFGLASGTARGMATQARVAAYKVCW---LGGC 357
Query: 153 SGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGP 212
I A D AI DGV++I++S+G A D+ D IAIG+F A GIL + +AGN GP
Sbjct: 358 FSSDIAAGIDKAIEDGVNIISMSIGGSSA-DYFRDIIAIGAFTANSHGILVSTSAGNGGP 534
Query: 213 IPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI 253
PSS+ + APW+ +V A +IDR F + LGN T G S+
Sbjct: 535 SPSSLSNTAPWITTVGAGTIDRDFPAYITLGNNITHTGASL 657
>TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease,
partial (10%)
Length = 574
Score = 100 bits (250), Expect = 3e-22
Identities = 64/165 (38%), Positives = 96/165 (57%), Gaps = 8/165 (4%)
Frame = +2
Query: 157 ILAAFDDAISDGVDVITVSLGPEH---ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPI 213
+LAA D AI+DGVDV +VS G A + D ++IGS HA+ K IL +AGN GP
Sbjct: 5 VLAAIDQAITDGVDVNSVSAGGRSIISAEEIFTDEVSIGSVHALAKNILVVASAGNDGPA 184
Query: 214 PSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKS--INITPSNGKKIPIAVRNAQ 271
+V + APW+ +VAA++IDR+F + LGN + G S +N+ P+ ++ + +
Sbjct: 185 LGTVLNVAPWVFTVAASTIDREFSSTITLGNNQQITGASLFVNLPPN---QLFSLILSTD 355
Query: 272 ACLAGG-NASPEMC--DCIDENMVKGKLVLCGSRNGKDLAYANGA 313
A LA N ++C +D VKGK+V+C +R GK + A G+
Sbjct: 356 AKLANATNRDAQLCRPGTLDPAKVKGKVVVC-TRGGKIKSVAEGS 487
>AF000393
Length = 411
Score = 83.2 bits (204), Expect = 7e-17
Identities = 35/70 (50%), Positives = 52/70 (74%)
Frame = +2
Query: 12 EFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKGLGPIPKKWRGV 71
++ + TT+SWDF+G Q + R + E+D+++GV+D IWPES+SF+D+G GP PKKW+G
Sbjct: 14 KYILHTTKSWDFIGFPQQVTRASM-ESDIIVGVVDTGIWPESKSFSDEGFGPPPKKWKGS 190
Query: 72 CAGGGNFSCN 81
C NF+CN
Sbjct: 191C---HNFTCN 211
>TC17430 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1,
partial (12%)
Length = 729
Score = 79.7 bits (195), Expect = 8e-16
Identities = 37/81 (45%), Positives = 55/81 (67%)
Frame = +1
Query: 1 MKGVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
M V+SV P ++ + TT+SWDF+G Q + R + E+D+++GV+D I PES+SF+D+G
Sbjct: 259 MDEVVSVIPDSKYILHTTKSWDFIGFPQQVTRASM-ESDIIVGVVDTGIGPESKSFSDEG 435
Query: 61 LGPIPKKWRGVCAGGGNFSCN 81
GP PKK +G NF+CN
Sbjct: 436 FGPPPKKGKG---SSHNFTCN 489
>BP042306
Length = 549
Score = 72.8 bits (177), Expect = 9e-14
Identities = 35/71 (49%), Positives = 47/71 (65%), Gaps = 4/71 (5%)
Frame = +3
Query: 4 VISVFPSEEFHIQTTRSWDFLGLR----QSIKRDQIIETDLVIGVIDIRIWPESESFNDK 59
V+SVF S+E + TTRSW+FLGLR S + + +I ID +WPES+SF+DK
Sbjct: 327 VVSVFVSKEHKLHTTRSWEFLGLRGNGGNSAWQKGRFGENTIIANIDTGVWPESQSFSDK 506
Query: 60 GLGPIPKKWRG 70
G+G +P KWRG
Sbjct: 507 GIGSVPAKWRG 539
>BP033910
Length = 538
Score = 72.4 bits (176), Expect = 1e-13
Identities = 30/63 (47%), Positives = 46/63 (72%)
Frame = +1
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKGLGP 63
V+SVFP+ + TTRSWDFLG+ +++R+ +E+++V+GV+D IW + SFND G GP
Sbjct: 349 VVSVFPNTRRKLHTTRSWDFLGMPLNVRRNPAMESNMVVGVLDTGIWVDCPSFNDTGYGP 528
Query: 64 IPK 66
P+
Sbjct: 529 PPR 537
>AU088875
Length = 322
Score = 66.6 bits (161), Expect = 7e-12
Identities = 30/71 (42%), Positives = 45/71 (63%)
Frame = +3
Query: 183 DFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVIL 242
+ D ++IGSFHA+ K IL +AGN GP +V + APW+ +VAA++ID +F + L
Sbjct: 6 EIFTDEVSIGSFHALAKNILVVASAGNDGPALGTVLNVAPWVFTVAASTIDXEFSSTITL 185
Query: 243 GNGKTFVGKSI 253
GN + G S+
Sbjct: 186 GNNQQITGASL 218
>TC19209 similar to GB|BAB11244.1|10177874|AB010074 serine protease-like
protein {Arabidopsis thaliana;} , partial (12%)
Length = 553
Score = 64.7 bits (156), Expect = 3e-11
Identities = 30/68 (44%), Positives = 44/68 (64%), Gaps = 4/68 (5%)
Frame = +1
Query: 2 KGVISVFPSEEFHIQTTRSWDFLGLRQ----SIKRDQIIETDLVIGVIDIRIWPESESFN 57
+GV+++FP + + TTRS FLGL + I +++ D+V+GV D IWPES+SFN
Sbjct: 343 EGVVAIFPETRYQLHTTRSPTFLGLEKMQSTEIWSEKLSNHDVVVGVSDTGIWPESQSFN 522
Query: 58 DKGLGPIP 65
D G P+P
Sbjct: 523 DTGFKPVP 546
>BP042339
Length = 447
Score = 64.7 bits (156), Expect = 3e-11
Identities = 42/141 (29%), Positives = 70/141 (48%), Gaps = 6/141 (4%)
Frame = +1
Query: 196 AMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINI 255
A++ G+ QAAGN GP P ++ S +PW+ SVAA DR++ + + LGNGK G I +
Sbjct: 1 AVKAGVFVAQAAGNGGPFPKTMVSYSPWITSVAAAIDDRRYKNHLTLGNGKILAG--IGL 174
Query: 256 TPSNGKKIPIAVRNAQACLAGGNA---SPEMC---DCIDENMVKGKLVLCGSRNGKDLAY 309
+PS + A L + SP C + +++ +++G ++LCG +
Sbjct: 175 SPSTHLNQTYKLVAANDVLLDSSLIKYSPTDCQRPEVLNKKLIEGNILLCG--YSFNFVV 348
Query: 310 ANGAIGSIHNVTKSQLGASFV 330
+I + K+ A FV
Sbjct: 349 GTASIKKVSETAKTLGAAGFV 411
>BP066980
Length = 472
Score = 35.4 bits (80), Expect = 0.017
Identities = 16/24 (66%), Positives = 19/24 (78%)
Frame = +3
Query: 4 VISVFPSEEFHIQTTRSWDFLGLR 27
V+SVF S+E I T RSW+FLGLR
Sbjct: 384 VVSVFVSKEH*IDTNRSWEFLGLR 455
>TC13364 similar to UP|Q9LUM3 (Q9LUM3) Subtilisin proteinase-like protein,
partial (13%)
Length = 514
Score = 33.1 bits (74), Expect = 0.083
Identities = 20/47 (42%), Positives = 28/47 (59%), Gaps = 4/47 (8%)
Frame = +1
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIET----DLVIGVID 46
V ++ P + + TTRS FLGL+ + + + ET DLVIGVID
Sbjct: 367 VTTLIPEQVRQLHTTRSPHFLGLKTADRAGLLHETDFGSDLVIGVID 507
>BP066497
Length = 389
Score = 29.6 bits (65), Expect = 0.91
Identities = 20/68 (29%), Positives = 33/68 (48%)
Frame = +3
Query: 288 DENMVKGKLVLCGSRNGKDLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQ 347
D+N+VKGK+VL S + G +G + T++ A++ P L D V +
Sbjct: 147 DQNLVKGKMVLGESWYPNE----TGPMGFLSTTTQTPFAAAY--PWAGSYLDFKDAVSVW 308
Query: 348 SYTNSSKY 355
Y S++Y
Sbjct: 309 KYIKSTRY 332
>TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine endopeptidase
XSP1, partial (20%)
Length = 635
Score = 29.6 bits (65), Expect = 0.91
Identities = 24/77 (31%), Positives = 35/77 (45%), Gaps = 3/77 (3%)
Frame = +1
Query: 285 DCIDENMVKGKLVLC--GSRNGKDLAYANGAIGSIHNVTK-SQLGASFVTPRPSLNLKTN 341
D + N VKGKLV C G+ + + G IGSI + L F+ P LN
Sbjct: 88 DSLQPNKVKGKLVYCKLGNWGTEGVVKKFGGIGSIMESDQYPDLAQIFMAPATILNHTIG 267
Query: 342 DFVHIQSYTNSSKYPVA 358
+ + +Y S++ P A
Sbjct: 268 E--SVTNYIKSTRSPSA 312
>BI419814
Length = 527
Score = 28.9 bits (63), Expect = 1.6
Identities = 20/81 (24%), Positives = 37/81 (44%), Gaps = 4/81 (4%)
Frame = +2
Query: 316 SIHNVTKSQLGASFVTPRPSLNLKTNDFVHIQS---YTNSSKYPVAD-FLFSWPKSFGPI 371
+I + +G V PS +L++ +F H+ + +++ + + FL S+P SF P
Sbjct: 101 TIQMLVSGAIGDVLVVLPPSASLRSPEFRHLHAPPVVSDAKNFLCSGGFLKSYPSSFNPK 280
Query: 372 NYETRYKCPRSGNLSRIFTSS 392
+Y Y G + R S
Sbjct: 281 HYNELYNSRTRGLVMRASADS 343
>AV407175
Length = 354
Score = 28.9 bits (63), Expect = 1.6
Identities = 16/43 (37%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = -2
Query: 47 IRIWPESESFNDKGLGP---IPKKWRGVCAGGGNFSCNNKIIG 86
+R W E F DKG P PK+WR GG+ IG
Sbjct: 182 VRTWRTMEGFADKGNSPGVESPKQWRCCL*NGGSCVATMAAIG 54
>TC18609 weakly similar to UP|Q7YRB0 (Q7YRB0) Block of proliferation 1
(Fragment), partial (10%)
Length = 357
Score = 27.7 bits (60), Expect = 3.5
Identities = 16/44 (36%), Positives = 21/44 (47%), Gaps = 7/44 (15%)
Frame = -3
Query: 265 IAVRNAQACLAG------GNASPEMCDCIDENMVKGKLVL-CGS 301
I ACL+ G+ PE CD IDE+ + L+L C S
Sbjct: 316 ICFNRIPACLSSISQLTLGDCLPESCDAIDESTQRSSLILFCSS 185
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,737,258
Number of Sequences: 28460
Number of extensions: 92159
Number of successful extensions: 450
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 421
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 423
length of query: 395
length of database: 4,897,600
effective HSP length: 92
effective length of query: 303
effective length of database: 2,279,280
effective search space: 690621840
effective search space used: 690621840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148652.13


