

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148651.5 + phase: 0
(496 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
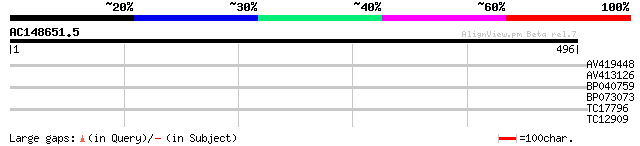
Score E
Sequences producing significant alignments: (bits) Value
AV419448 29 1.5
AV413126 29 2.0
BP040759 28 4.5
BP073073 27 7.7
TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 27 7.7
TC12909 27 7.7
>AV419448
Length = 382
Score = 29.3 bits (64), Expect = 1.5
Identities = 12/34 (35%), Positives = 21/34 (61%)
Frame = +1
Query: 141 VPLGLNEFKSRAISSKVKSGTGQSRSVIHRLEPG 174
VPL ++ FKS ++S S + +++H+L PG
Sbjct: 232 VPLTVSNFKSLCLASSDPSSSSYKNTLVHKLFPG 333
>AV413126
Length = 370
Score = 28.9 bits (63), Expect = 2.0
Identities = 12/29 (41%), Positives = 16/29 (54%)
Frame = -2
Query: 234 TIEIANFEHHSSNLKDFEIHGSLNFPTNV 262
T I NF + DF +HG + FPTN+
Sbjct: 177 TTPIQNFPIPCFLVPDFAVHGGIKFPTNI 91
>BP040759
Length = 482
Score = 27.7 bits (60), Expect = 4.5
Identities = 23/81 (28%), Positives = 36/81 (44%), Gaps = 1/81 (1%)
Frame = +1
Query: 298 HYGSEFYCTLSVVEVFGVDAVERMLEDLINTQDNLLASGEGNADKTILPHPDPAVIEHVH 357
H+ F+ TL ++V + R+ L +LAS +K +P P V H H
Sbjct: 193 HHQRAFHFTLRTIQVMK*VTILRIA--LGANHQPVLASSFPQRNKRTSHYPAPWVEVHSH 366
Query: 358 KKPLEGINSVPA-SDISSSKH 377
+ P + VP+ SD S + H
Sbjct: 367 QAPSTALVPVPSQSDTSYAAH 429
>BP073073
Length = 490
Score = 26.9 bits (58), Expect = 7.7
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -3
Query: 471 ASDLISWKSQVSSQLNHLIQD 491
+SD +SWK S+LNH + D
Sbjct: 188 SSDQVSWKCNCCSKLNHSVTD 126
>TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 594
Score = 26.9 bits (58), Expect = 7.7
Identities = 17/43 (39%), Positives = 21/43 (48%)
Frame = +1
Query: 153 ISSKVKSGTGQSRSVIHRLEPGGAEYNYASASKGAKVLGSNKE 195
ISSKV GT R+ + + GGA + K VLG KE
Sbjct: 445 ISSKVVKGTKTGRATVIAMFLGGANEVVSILGKEFPVLGLKKE 573
>TC12909
Length = 609
Score = 26.9 bits (58), Expect = 7.7
Identities = 12/40 (30%), Positives = 22/40 (55%)
Frame = +3
Query: 223 IMELSEETLVDTIEIANFEHHSSNLKDFEIHGSLNFPTNV 262
+ ++S E + + + + S NLK+ +I GS N PT +
Sbjct: 39 LYQVSFEDMKEILVVLRLITSSPNLKELQISGSSNIPTAI 158
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,830,234
Number of Sequences: 28460
Number of extensions: 100202
Number of successful extensions: 418
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 417
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 418
length of query: 496
length of database: 4,897,600
effective HSP length: 94
effective length of query: 402
effective length of database: 2,222,360
effective search space: 893388720
effective search space used: 893388720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148651.5


