

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.5 - phase: 0
(122 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
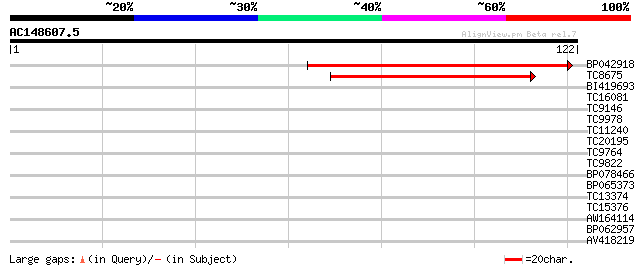
Sequences producing significant alignments: (bits) Value
BP042918 70 1e-13
TC8675 weakly similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter... 44 1e-05
BI419693 28 0.57
TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), par... 27 0.75
TC9146 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pr... 25 0.90
TC9978 similar to UP|Q84TF0 (Q84TF0) At2g37790, partial (36%) 27 1.3
TC11240 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transfera... 26 1.7
TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%) 26 2.2
TC9764 25 2.8
TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 25 3.7
BP078466 25 4.8
BP065373 25 4.8
TC13374 similar to UP|Q9FJK7 (Q9FJK7) Cyclin C-like protein, par... 25 4.8
TC15376 weakly similar to UP|Q7SYH2 (Q7SYH2) Cystatin domain fet... 24 6.3
AW164114 24 6.3
BP062957 24 8.3
AV418219 24 8.3
>BP042918
Length = 412
Score = 70.1 bits (170), Expect = 1e-13
Identities = 28/57 (49%), Positives = 38/57 (66%)
Frame = -3
Query: 65 VSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
+ RLPVFYK R +LF P W Y +P ++L+IP++ E VWV +TYY GF P R+
Sbjct: 407 IQRLPVFYKHRDHLFHPAWTYTVPNFLLRIPISIFESLVWVAITYYTTGFAPEASRF 237
>TC8675 weakly similar to UP|Q9LFH0 (Q9LFH0) PDR9 ABC transporter, partial
(13%)
Length = 727
Score = 43.5 bits (101), Expect = 1e-05
Identities = 16/44 (36%), Positives = 31/44 (70%)
Frame = +3
Query: 70 VFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIG 113
V Y++R + PWAY+ ++++P +F++ A++VI+TY +IG
Sbjct: 18 VLYRERFAGMYSPWAYSFAQVLIEVPYSFIQAALYVIITYPMIG 149
>BI419693
Length = 618
Score = 27.7 bits (60), Expect = 0.57
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +1
Query: 78 LFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPY 117
L +P A +P W +PL + + + VILTYY I D Y
Sbjct: 280 LRYPMKANLIPFWA--VPLIAILLPLVVILTYYFIRNDVY 393
>TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), partial (37%)
Length = 1189
Score = 27.3 bits (59), Expect = 0.75
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +3
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 291 YPSWLLVLGAGTLPFGTLFIELFFILSSIWLGRFYYVFGF 410
>TC9146 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane
protein-like, partial (31%)
Length = 980
Score = 25.0 bits (53), Expect(2) = 0.90
Identities = 10/28 (35%), Positives = 16/28 (56%)
Frame = +2
Query: 87 LPAWILKIPLTFVEVAVWVILTYYVIGF 114
LP + I L F+ ++W+ YY+ GF
Sbjct: 191 LPFGAVFIELFFILTSIWLNQFYYIFGF 274
Score = 20.4 bits (41), Expect(2) = 0.90
Identities = 7/18 (38%), Positives = 10/18 (54%)
Frame = +3
Query: 76 GYLFFPPWAYALPAWILK 93
G +F PW+ W+LK
Sbjct: 33 GLVFQCPWSLWAVTWVLK 86
>TC9978 similar to UP|Q84TF0 (Q84TF0) At2g37790, partial (36%)
Length = 813
Score = 26.6 bits (57), Expect = 1.3
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +1
Query: 87 LPAWILKIPLTFVEVAVWVILTYYVIG 113
L +I K LTF +A+WV+ T +++G
Sbjct: 499 LELYIYKNNLTFYNLALWVLDTLFLVG 579
>TC11240 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transferase, partial
(31%)
Length = 528
Score = 26.2 bits (56), Expect = 1.7
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -2
Query: 92 LKIPLTFVEVAVWVILTYYVIGFD 115
LK L+ + A W++L Y IGFD
Sbjct: 173 LKYSLSILNNASWILLAAYHIGFD 102
>TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%)
Length = 591
Score = 25.8 bits (55), Expect = 2.2
Identities = 20/73 (27%), Positives = 35/73 (47%)
Frame = +1
Query: 34 DSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILK 93
DS A I++ AL ++++ I V LS+V+S L + + W + P W+L
Sbjct: 160 DSTAEARIFMLALMLLFLLLIGIAVFLLSLVLSLLLLIC-----V*CWCWCWCCPCWLLM 324
Query: 94 IPLTFVEVAVWVI 106
+ L + A V+
Sbjct: 325 LSLLVLIYAAAVL 363
>TC9764
Length = 534
Score = 25.4 bits (54), Expect = 2.8
Identities = 18/66 (27%), Positives = 32/66 (48%)
Frame = +2
Query: 46 LFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWV 105
L G +V+L V + + R+ +F Q G + P L AW L++ T+ + WV
Sbjct: 137 LLEGSMVVLVYPVLQAMSI--RI*LF*TQIGVVAESP---VLQAWFLQLVSTYQFLKKWV 301
Query: 106 ILTYYV 111
L +++
Sbjct: 302 TLFFFI 319
>TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (40%)
Length = 915
Score = 25.0 bits (53), Expect = 3.7
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = +1
Query: 7 VYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAH 38
VYIF LC + +++ + EMH S +H
Sbjct: 664 VYIFFLCHVYFQSLVICSSNYLDEMHHISFSH 759
>BP078466
Length = 530
Score = 24.6 bits (52), Expect = 4.8
Identities = 11/27 (40%), Positives = 16/27 (58%), Gaps = 4/27 (14%)
Frame = -3
Query: 70 VFYKQR----GYLFFPPWAYALPAWIL 92
VFYK+ Y F+P +Y+L W+L
Sbjct: 90 VFYKELFCILHYCFYPSLSYSLKQWLL 10
>BP065373
Length = 516
Score = 24.6 bits (52), Expect = 4.8
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +1
Query: 47 FYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFP 81
FYG ++L I MV++ LPVF G L FP
Sbjct: 361 FYGXKLVLHISYPVHRMVITGLPVF--PHGTLPFP 459
>TC13374 similar to UP|Q9FJK7 (Q9FJK7) Cyclin C-like protein, partial (64%)
Length = 569
Score = 24.6 bits (52), Expect = 4.8
Identities = 20/59 (33%), Positives = 31/59 (51%)
Frame = +2
Query: 59 AELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPY 117
AE S V +RL VFY ++ Y + Y + IL++ + +E L YY++ F PY
Sbjct: 365 AEESTVQARLLVFYIKKLYAD-DKYRYEIKD-ILEMEMKILEA-----LNYYLVVFHPY 520
>TC15376 weakly similar to UP|Q7SYH2 (Q7SYH2) Cystatin domain fetuin-like
protein, partial (6%)
Length = 582
Score = 24.3 bits (51), Expect = 6.3
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = -2
Query: 44 GALFYGCIVILFIGVAELSMVVSRLPVFYKQRGY 77
GALF GC+ + A +++ S V Y + GY
Sbjct: 158 GALFVGCLTPVH*NKASATIISSNGGVGYGREGY 57
>AW164114
Length = 297
Score = 24.3 bits (51), Expect = 6.3
Identities = 9/36 (25%), Positives = 15/36 (41%)
Frame = -1
Query: 83 WAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYI 118
W ++ AW + AVW +L+ F P +
Sbjct: 195 WVFSFAAWAPWAVFSLAAWAVWAVLSLAAWAFSPML 88
>BP062957
Length = 269
Score = 23.9 bits (50), Expect = 8.3
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -2
Query: 53 ILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYAL 87
+ F G +L + LPVFY L FPP+ Y +
Sbjct: 118 VTFSGPVQLDLFP--LPVFYLFWKLLVFPPYFYPI 20
>AV418219
Length = 305
Score = 23.9 bits (50), Expect = 8.3
Identities = 8/11 (72%), Positives = 8/11 (72%)
Frame = +1
Query: 80 FPPWAYALPAW 90
FPPW AL AW
Sbjct: 238 FPPWDVALKAW 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.336 0.149 0.486
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,337,303
Number of Sequences: 28460
Number of extensions: 31864
Number of successful extensions: 271
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 271
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 271
length of query: 122
length of database: 4,897,600
effective HSP length: 79
effective length of query: 43
effective length of database: 2,649,260
effective search space: 113918180
effective search space used: 113918180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC148607.5


