

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.15 - phase: 0
(174 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
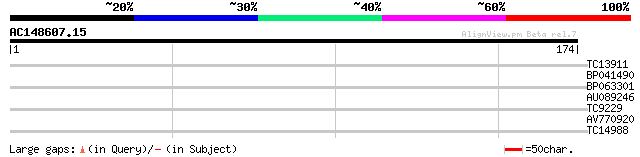
TC13911 homologue to UP|MPK2_ARATH (Q39022) Mitogen-activated pr... 25 5.6
BP041490 25 5.6
BP063301 25 5.6
AU089246 25 7.3
TC9229 similar to UP|Q9SKU1 (Q9SKU1) At2g20760 protein (At2g2076... 25 7.3
AV770920 25 7.3
TC14988 homologue to UP|Q8L7J5 (Q8L7J5) Pyruvate kinase , parti... 25 9.6
>TC13911 homologue to UP|MPK2_ARATH (Q39022) Mitogen-activated protein
kinase homolog 2 (MAP kinase 2) (AtMPK2) , partial
(14%)
Length = 499
Score = 25.4 bits (54), Expect = 5.6
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +3
Query: 121 YFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFC 162
+FH++ L G ++ Y DFI+ LSK+ +LS K C
Sbjct: 216 FFHIKDLGGGCYLSF--YTDFIQ*RPKLSKLGLVLSLKSCCC 335
>BP041490
Length = 545
Score = 25.4 bits (54), Expect = 5.6
Identities = 12/39 (30%), Positives = 19/39 (47%)
Frame = -1
Query: 36 NRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADN 74
N PLS + T E+ + A + + + Q E H D + N
Sbjct: 224 NSPLSHINTESESPSSAAIAAIEVSQHSSIETHEDISQN 108
>BP063301
Length = 462
Score = 25.4 bits (54), Expect = 5.6
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +1
Query: 62 GVEGEVHPDGADNSPKPASAGL 83
G+ GE+ PDGA +P P G+
Sbjct: 130 GLGGEILPDGAARAPCPLHRGI 195
>AU089246
Length = 392
Score = 25.0 bits (53), Expect = 7.3
Identities = 16/34 (47%), Positives = 17/34 (49%)
Frame = +3
Query: 14 RILENPNQQLDRYFNSFKELASNRPLSELRTADE 47
R E+ Q LDR S A N PL RTADE
Sbjct: 207 RRFESFQQSLDRTIPSTSSHAINFPLLADRTADE 308
>TC9229 similar to UP|Q9SKU1 (Q9SKU1) At2g20760 protein
(At2g20760/F5H14.27), partial (31%)
Length = 756
Score = 25.0 bits (53), Expect = 7.3
Identities = 21/73 (28%), Positives = 31/73 (41%), Gaps = 3/73 (4%)
Frame = +1
Query: 60 DQGV---EGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETT 116
D+GV +G V P D P+ G E + + EE K+ KE KII
Sbjct: 385 DEGVFVSDGPVLPPPGDMEPEE---GYALREWRRQNAILLEEKEKREKEMRLKIIEEAEE 555
Query: 117 IRRPYFHVRPLNI 129
+ ++ R LN+
Sbjct: 556 YKVAFYEKRKLNV 594
>AV770920
Length = 426
Score = 25.0 bits (53), Expect = 7.3
Identities = 9/12 (75%), Positives = 9/12 (75%)
Frame = +1
Query: 158 KGTFCSIFSCYR 169
KG CSIF CYR
Sbjct: 169 KGVSCSIFKCYR 204
>TC14988 homologue to UP|Q8L7J5 (Q8L7J5) Pyruvate kinase , partial (61%)
Length = 1179
Score = 24.6 bits (52), Expect = 9.6
Identities = 11/37 (29%), Positives = 22/37 (58%)
Frame = -2
Query: 39 LSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNS 75
L +RTA++A G++ +G+D + ++ P D+S
Sbjct: 581 LLAVRTAEDARLSNGLIGKGVDLIISLKIAP*SRDDS 471
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,629,973
Number of Sequences: 28460
Number of extensions: 28562
Number of successful extensions: 151
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 151
length of query: 174
length of database: 4,897,600
effective HSP length: 84
effective length of query: 90
effective length of database: 2,506,960
effective search space: 225626400
effective search space used: 225626400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148607.15


