

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.14 - phase: 0
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
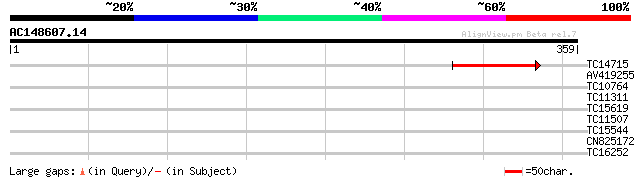
Score E
Sequences producing significant alignments: (bits) Value
TC14715 95 2e-20
AV419255 28 2.4
TC10764 27 4.1
TC11311 similar to UP|Q9FPH4 (Q9FPH4) AT4g33900, partial (30%) 27 4.1
TC15619 similar to UP|Q8GTX9 (Q8GTX9) Cell cycle control crn (Cr... 27 4.1
TC11507 weakly similar to UP|Q9HD28 (Q9HD28) Retinitis pigmentos... 27 5.3
TC15544 weakly similar to PIR|E88979|E88979 protein F37B4.8 [imp... 26 9.0
CN825172 26 9.0
TC16252 26 9.0
>TC14715
Length = 637
Score = 94.7 bits (234), Expect = 2e-20
Identities = 41/56 (73%), Positives = 42/56 (74%)
Frame = +2
Query: 281 YSAQSPAQSVAGAYPNGPNQWPNYGVQPQTWPATTQAQGQQWPAGYTQQQVNWFFP 336
YS QSPAQSV GAYPN PNQW NYG QPQTWPA Q QGQQWPAGY QQ +P
Sbjct: 5 YSVQSPAQSVMGAYPNAPNQWTNYGAQPQTWPAVPQIQGQQWPAGYNQQASYGTYP 172
>AV419255
Length = 421
Score = 28.1 bits (61), Expect = 2.4
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +2
Query: 199 ESLVVKFITPNPENPGVASATEREELSNIFLE 230
ESL +F+TPN + G A+ E + IFLE
Sbjct: 14 ESLPGQFVTPNSPSLGAAAQIEDDFFCGIFLE 109
>TC10764
Length = 555
Score = 27.3 bits (59), Expect = 4.1
Identities = 15/35 (42%), Positives = 19/35 (53%), Gaps = 3/35 (8%)
Frame = -3
Query: 318 QGQQWPAGYTQQ---QVNWFFPSTIFLVCCFPSVQ 349
QG Q P QQ Q+N FP+ LVCC +V+
Sbjct: 124 QGDQTPHPNAQQLTHQINLDFPTRRLLVCCHVTVK 20
>TC11311 similar to UP|Q9FPH4 (Q9FPH4) AT4g33900, partial (30%)
Length = 523
Score = 27.3 bits (59), Expect = 4.1
Identities = 12/22 (54%), Positives = 14/22 (63%), Gaps = 1/22 (4%)
Frame = +1
Query: 325 GYTQQQVNWFFPS-TIFLVCCF 345
GYT +NWF P IFL+ CF
Sbjct: 349 GYTTN*INWFHPV*QIFLLICF 414
>TC15619 similar to UP|Q8GTX9 (Q8GTX9) Cell cycle control crn (Crooked neck)
protein-like, partial (26%)
Length = 822
Score = 27.3 bits (59), Expect = 4.1
Identities = 17/64 (26%), Positives = 33/64 (51%), Gaps = 3/64 (4%)
Frame = +3
Query: 102 IIRHANMEHRLGKLEDAFSLYEQAIA---IEKGKEHSQTLPMLFAQYSRFVYLASGNSEK 158
++ + +E +G E +YE+AIA + K + Q L+ Y+ + L +G+ E+
Sbjct: 12 LVDYIRLEESVGNKERVRDVYERAIANVPPAEEKLYWQRYIYLWINYALYEELDAGDVER 191
Query: 159 AREI 162
RE+
Sbjct: 192 TREV 203
>TC11507 weakly similar to UP|Q9HD28 (Q9HD28) Retinitis pigmentosa GTPase
regulator (Fragment), partial (3%)
Length = 744
Score = 26.9 bits (58), Expect = 5.3
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = -2
Query: 84 RAAYQLVHTEISPGLLEAIIRHANMEHRLGKLE 116
RA Y HT+ISP LL A+ E R+G +E
Sbjct: 311 RARYTTTHTQISPPLLLAV------ESRIGAVE 231
>TC15544 weakly similar to PIR|E88979|E88979 protein F37B4.8 [imported] -
Caenorhabditis elegans {Caenorhabditis elegans;} ,
partial (3%)
Length = 615
Score = 26.2 bits (56), Expect = 9.0
Identities = 14/48 (29%), Positives = 25/48 (51%)
Frame = +3
Query: 145 YSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKR 192
+S+ +A+GN KAR+ + ++ LSK +H ++ P P R
Sbjct: 195 HSQLTSIANGNGSKARKPISKSAKSPELSK----NSIHSSSLGPSPSR 326
>CN825172
Length = 655
Score = 26.2 bits (56), Expect = 9.0
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 278 SRAYSAQSPAQSVAGAYPNGPNQWPNYGVQPQ 309
SR+ S+ SP+ V P P W N V+PQ
Sbjct: 99 SRSSSSTSPSVRVEIVSPEQPRCWSNLVVKPQ 194
>TC16252
Length = 550
Score = 26.2 bits (56), Expect = 9.0
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = -3
Query: 86 AYQLVHTEISPGLLEAIIRHANMEHRLGKLE 116
A+ L+H+ + PG L+ H N+ H LG+LE
Sbjct: 218 AWNLLHSLVHPG*LDQSCGHRNLLH-LGELE 129
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,107,139
Number of Sequences: 28460
Number of extensions: 82302
Number of successful extensions: 417
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 417
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 417
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148607.14


