

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.6 + phase: 0
(243 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
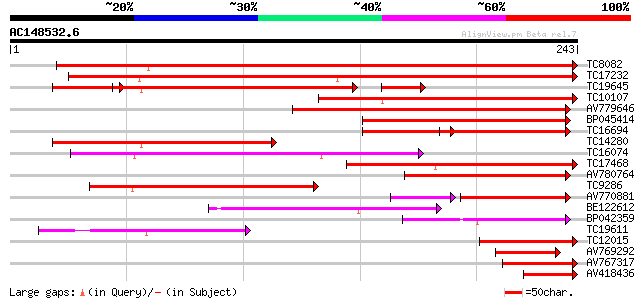
Score E
Sequences producing significant alignments: (bits) Value
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 236 3e-63
TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 226 3e-60
TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing pro... 141 9e-43
TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), p... 158 7e-40
AV779646 158 7e-40
BP045414 120 3e-28
TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein... 87 2e-25
TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-contain... 108 6e-25
TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein... 103 2e-23
TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 97 2e-21
AV780764 95 1e-20
TC9286 similar to UP|O65009 (O65009) BURP domain containing prot... 79 9e-16
AV770881 60 3e-14
BE122612 68 1e-12
BP042359 67 4e-12
TC19611 59 8e-10
TC12015 weakly similar to UP|O65009 (O65009) BURP domain contain... 54 2e-08
AV769292 48 2e-06
AV767317 40 5e-04
AV418436 39 8e-04
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 236 bits (601), Expect = 3e-63
Identities = 108/225 (48%), Positives = 150/225 (66%), Gaps = 2/225 (0%)
Frame = +1
Query: 21 ASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATF--LPREVANSIPFSSNKVENIL 78
AS LH+ FF E LH+ K M K ++ LPR++ IPFS +K++ +L
Sbjct: 955 ASDHHLHEISARVGFFQEDQLHSDNKFYMHLNKIRSSIPLLPRQITQQIPFSLDKMKEVL 1134
Query: 79 NYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEA 138
S+K S +E +++ I CE P ++GEEK C TSLESM+DF TSKLG NV A+ST+
Sbjct: 1135 EMLSVKPNSKNAEIMEKTISGCEAPAMDGEEKYCATSLESMMDFVTSKLGKNVHAMSTDV 1314
Query: 139 NKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVK 198
KE+ Q ++ GVKKL+++ ++ CH L+YPY VF CH+L+ T Y++P+EG DG +VK
Sbjct: 1315 QKETKLQNLLVKDGVKKLADDHVIACHPLAYPYVVFGCHELQKTSAYFVPLEGEDGVRVK 1494
Query: 199 TAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
A+CH DTS+W P ++AF VLKV+PGTVPVCHF GH++W SK
Sbjct: 1495 AVAVCHKDTSKWDPNYVAFRVLKVKPGTVPVCHFFPEGHLLWLSK 1629
>TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (33%)
Length = 966
Score = 226 bits (576), Expect = 3e-60
Identities = 106/222 (47%), Positives = 150/222 (66%), Gaps = 4/222 (1%)
Frame = +3
Query: 26 LHDYQNLDLFFFEKDLHNGTKLNMQFTKT--GATFLPREVANSIPFSSNKVENILNYFSI 83
LH+ +FFFE++LH GTKL++Q +K LPR++A+ IPFS K++ IL +
Sbjct: 108 LHEKTKAGVFFFEENLHPGTKLDLQLSKRKPAVPILPRKIADQIPFSLEKLKEILEILVV 287
Query: 84 KQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEAN--KE 141
K S +E +++ +G CE P++EGEE+ C TSLES+VDF SKLG NV A+STE KE
Sbjct: 288 KPDSKNTEIVEKTMGGCELPSMEGEEQMCATSLESLVDFIASKLGKNVLAISTEVELEKE 467
Query: 142 SDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAA 201
+ Q ++ GVKKL+++ + VCH ++YPY V + HKL T Y++P+EG DG +VK A
Sbjct: 468 TQSQNLLVKDGVKKLADDGMAVCHPMTYPYPVCFSHKLVKTSAYFVPLEGEDGARVKAVA 647
Query: 202 ICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+CH DTS+W P H++F L V+PGTVPVCH L H++W K
Sbjct: 648 VCHRDTSKWDPNHVSFPDLNVKPGTVPVCHVLPERHLLWVPK 773
>TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (38%)
Length = 619
Score = 141 bits (356), Expect(3) = 9e-43
Identities = 73/110 (66%), Positives = 85/110 (76%), Gaps = 5/110 (4%)
Frame = +3
Query: 45 TKLNMQFTKTG-----ATFLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGY 99
TKLN+QFTKT FLPR+VA+SIPFSSNKV+++LN FS+K GS E+E +K I
Sbjct: 120 TKLNLQFTKTSNDAAATFFLPRQVADSIPFSSNKVDDVLNKFSLKPGSEEAEIVKNTIAE 299
Query: 100 CETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYII 149
CE P + GEEK CVTSLESMVDFTTSKLG NVEAVST+ NKE + QQY I
Sbjct: 300 CEEPGVVGEEKRCVTSLESMVDFTTSKLGKNVEAVSTDVNKEKELQQYTI 449
Score = 41.6 bits (96), Expect(3) = 9e-43
Identities = 18/31 (58%), Positives = 22/31 (70%)
Frame = +2
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNM 49
Y AS TQLH+ N+ LFF EKDLH G K+ +
Sbjct: 41 YAASETQLHENPNVALFFLEKDLHQGNKIEL 133
Score = 26.9 bits (58), Expect(3) = 9e-43
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +2
Query: 160 KIVVCHLLSYPYAVFYCHK 178
K+++ H +YPYAVF C K
Sbjct: 482 KLLLFHKENYPYAVFXCXK 538
>TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(65%)
Length = 575
Score = 158 bits (400), Expect = 7e-40
Identities = 72/112 (64%), Positives = 88/112 (78%), Gaps = 1/112 (0%)
Frame = +2
Query: 133 AVSTEANKESDKQQYIIAKGVKKLSE-NKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEG 191
AVST +KE++KQ+Y IA GVKKL +K VVCH +YPYAVFYCHK +T+ Y +P+EG
Sbjct: 2 AVSTVVDKETEKQRYTIATGVKKLEAGDKAVVCHKQNYPYAVFYCHKSDSTRAYSVPLEG 181
Query: 192 IDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+G +VK A+CH DTSQW+PKHLAF VLKV+PGTVPVCHFL HV+W SK
Sbjct: 182 DNGVRVKAVAVCHTDTSQWNPKHLAFQVLKVKPGTVPVCHFLPEDHVVWVSK 337
>AV779646
Length = 510
Score = 158 bits (400), Expect = 7e-40
Identities = 67/119 (56%), Positives = 89/119 (74%)
Frame = -3
Query: 122 FTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRA 181
F TSKLGN+V A+STE KE+ ++ GV+KL+++ I VCH ++YPY VF+CHKL+
Sbjct: 508 FITSKLGNDVHAMSTEVTKETKLLYLLVKDGVEKLADDYITVCHPMAYPYVVFWCHKLQK 329
Query: 182 TKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
T Y++P+EG DG +VK A+CH DTS+W P H AF LKV+PGTVPVCHFL GH++W
Sbjct: 328 TDAYFVPLEGEDGVRVKAVAVCHKDTSKWDPNHAAFQDLKVKPGTVPVCHFLPEGHLLW 152
>BP045414
Length = 478
Score = 120 bits (300), Expect = 3e-28
Identities = 51/89 (57%), Positives = 65/89 (72%)
Frame = -2
Query: 152 GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWS 211
GVKKL ++ I VCH L+YPYAVF+CHKL T Y++P+EG DG +V A+C DTS+W
Sbjct: 471 GVKKLPDDGIAVCHPLTYPYAVFFCHKLVKTSAYFVPLEGEDGARV*PVAVCPRDTSKWD 292
Query: 212 PKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
P H++F L VQPGTVPVCH L H++W
Sbjct: 291 PNHVSFPDLHVQPGTVPVCHVLPERHLLW 205
>TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2, partial
(12%)
Length = 488
Score = 87.4 bits (215), Expect(2) = 2e-25
Identities = 34/56 (60%), Positives = 43/56 (76%)
Frame = +1
Query: 185 YYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
Y++P+EG DG +V A+CH DTS+W P H AF LKV+PGTVPVCHFL GH++W
Sbjct: 106 YFVPLEGEDGVRVNHVAVCHQDTSKWDPNHAAFPDLKVKPGTVPVCHFLPEGHLLW 273
Score = 43.9 bits (102), Expect(2) = 2e-25
Identities = 18/40 (45%), Positives = 26/40 (65%)
Frame = +3
Query: 152 GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEG 191
GV+KL+++ I VCH ++YPY VF+CH L T + G
Sbjct: 6 GVEKLADDYITVCHPMAYPYLVFWCH*LPKTHALFCSSRG 125
>TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (54%)
Length = 723
Score = 108 bits (271), Expect = 6e-25
Identities = 57/98 (58%), Positives = 69/98 (70%), Gaps = 2/98 (2%)
Frame = +2
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVEN 76
Y A+ TQLH N+ LFF EK+LH GTK+N+ FTKT ATFLPR VA+SIPFSS KVE
Sbjct: 428 YAATETQLHSDPNVALFFLEKNLHPGTKMNLHFTKTSNKATFLPRIVADSIPFSSEKVEL 607
Query: 77 ILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVT 114
LN +S+K GS E++ +K I CE I+GEEK C T
Sbjct: 608 ALNKYSVKAGSEEAQVMKNTIDECEQKGIQGEEKYCAT 721
>TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(6%)
Length = 844
Score = 103 bits (258), Expect = 2e-23
Identities = 59/158 (37%), Positives = 85/158 (53%), Gaps = 7/158 (4%)
Frame = +2
Query: 27 HDYQNLDLFFFEKDLHNGTKLNMQFT----KTGATFLPREVANSIPFSSNKVENILNYFS 82
H L +FF KDL G K+ + F+ T FLPRE AN IPFSS ++ ++LN+FS
Sbjct: 353 HMDPELHVFFIPKDLKVGKKMPIYFSMKDSSTSPKFLPREEANQIPFSSKQLPSLLNFFS 532
Query: 83 IKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVE---AVSTEAN 139
I + S +++ +K + CE +EGE K C TSLES++D G + + +T
Sbjct: 533 IPKNSPQAKAMKYTLEQCEFEPMEGETKFCATSLESLLDSARHLFGLSPQFKVLTTTHLT 712
Query: 140 KESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCH 177
K Q VK+ ++ CH + YP+AVFYCH
Sbjct: 713 KSHVLLQNYTILEVKENRVPNVIGCHPMPYPFAVFYCH 826
>TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (11%)
Length = 652
Score = 97.4 bits (241), Expect = 2e-21
Identities = 42/100 (42%), Positives = 61/100 (61%), Gaps = 1/100 (1%)
Frame = +3
Query: 145 QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRA-TKVYYLPMEGIDGTKVKTAAIC 203
Q Y I + + ++S K+V CH + YPYAVFYCH + +VY + + G +G KV+ +C
Sbjct: 3 QNYTILENIMRISAPKMVACHTMPYPYAVFYCHSQESENRVYKVSLVGDNGDKVEAMVVC 182
Query: 204 HNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
H DTS W H++F +L V PG+ VCHF ++IW K
Sbjct: 183 HLDTSHWGSGHVSFQILGVTPGSSSVCHFFPADNLIWAPK 302
>AV780764
Length = 438
Score = 94.7 bits (234), Expect = 1e-20
Identities = 39/71 (54%), Positives = 50/71 (69%)
Frame = -1
Query: 170 PYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPV 229
PY VF+ H L + Y++P+EG DG +VKT A+CH TS+W P H A LKV+PGT PV
Sbjct: 378 PYVVFWFH*LPKPQAYFVPLEGEDGVRVKTIAVCHQHTSKWDPNHAALPNLKVKPGTGPV 199
Query: 230 CHFLQHGHVIW 240
CHFL GH++W
Sbjct: 198 CHFLLEGHLLW 166
>TC9286 similar to UP|O65009 (O65009) BURP domain containing protein,
partial (6%)
Length = 805
Score = 78.6 bits (192), Expect = 9e-16
Identities = 40/99 (40%), Positives = 63/99 (63%), Gaps = 1/99 (1%)
Frame = +1
Query: 35 FFFEKDLHNGTKLNMQF-TKTGATFLPREVANSIPFSSNKVENILNYFSIKQGSAESENL 93
FF DL+ G + +QF + + FL R+ N IPFSS+++ ++L FSI + S +++++
Sbjct: 445 FFALDDLYAGNVMTLQFPVQEISHFLSRKETNYIPFSSSQLPSVLQLFSIPEDSPKAKSM 624
Query: 94 KQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVE 132
+ +G CE I GE K C TSLESM++F S +G+N E
Sbjct: 625 RGTLGQCEGEPITGERKICATSLESMLEFVDSVIGSNTE 741
>AV770881
Length = 403
Score = 60.5 bits (145), Expect(2) = 3e-14
Identities = 25/47 (53%), Positives = 31/47 (65%)
Frame = -3
Query: 194 GTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
G +K +CH DTS+W P H AF LKV+PGTVPV FL G ++W
Sbjct: 311 GLGLKPLLVCHQDTSKWDPNHAAFPDLKVKPGTVPVSQFLPEGPLLW 171
Score = 33.5 bits (75), Expect(2) = 3e-14
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = -1
Query: 164 CHLLSYPYAVFYCHKLRATKVYYLPMEG 191
CH ++YPY VF+CH L T + G
Sbjct: 403 CHPMAYPYGVFWCHGLPKTHAIFCSSRG 320
>BE122612
Length = 386
Score = 68.2 bits (165), Expect = 1e-12
Identities = 39/101 (38%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Frame = +2
Query: 86 GSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQ 145
GS E + ++ ++ CE GE K CV S+E MVDF TS LG NV +TE S+K+
Sbjct: 23 GSME-KMMRDSLQECERVPSRGETKRCVGSIEDMVDFATSVLGRNVVVRTTENVNGSNKK 199
Query: 146 QYI-IAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVY 185
+ +GV + V CH +PY ++YCH L +VY
Sbjct: 200 VMVGRVQGVNGGKVTRSVSCHQSLFPYLLYYCHSLPKVRVY 322
>BP042359
Length = 404
Score = 66.6 bits (161), Expect = 4e-12
Identities = 31/75 (41%), Positives = 43/75 (57%), Gaps = 3/75 (4%)
Frame = -3
Query: 169 YPYAVFYCHKLRATKVYYLPMEGIDGTKVKT---AAICHNDTSQWSPKHLAFHVLKVQPG 225
YPY ++YCH + +VY + +D TK+K AICH DTS W P+H AF L PG
Sbjct: 402 YPYLLYYCHSVPQVRVYEAEILDVD-TKLKINHGVAICHLDTSAWGPEHGAFVALGAGPG 226
Query: 226 TVPVCHFLQHGHVIW 240
+ VCH++ + W
Sbjct: 225 KIEVCHWIFQNDMTW 181
>TC19611
Length = 533
Score = 58.9 bits (141), Expect = 8e-10
Identities = 36/93 (38%), Positives = 49/93 (51%), Gaps = 2/93 (2%)
Frame = -3
Query: 13 LFIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGAT--FLPREVANSIPFS 70
L Y Y A TQ FF E LH G K N+ K + +LPRE+A+ IPFS
Sbjct: 264 LLFYAYYVAKETQSRS------FFEEYQLHAGNKFNVNLNKIKSVVPWLPREIADHIPFS 103
Query: 71 SNKVENILNYFSIKQGSAESENLKQAIGYCETP 103
+K+ IL S+K S +E +++ IG CE+P
Sbjct: 102 LDKMNEILEMLSVKLESKNAEMVEKTIGDCESP 4
>TC12015 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (12%)
Length = 464
Score = 53.9 bits (128), Expect = 2e-08
Identities = 19/42 (45%), Positives = 27/42 (64%)
Frame = +1
Query: 202 ICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+CH DTS W+P H+ F L ++ G PVCHF H++W +K
Sbjct: 10 VCHLDTSDWTPSHIIFKQLGIEAGKAPVCHFFPIKHLMWVAK 135
>AV769292
Length = 250
Score = 47.8 bits (112), Expect = 2e-06
Identities = 20/28 (71%), Positives = 23/28 (81%)
Frame = -3
Query: 209 QWSPKHLAFHVLKVQPGTVPVCHFLQHG 236
+W+ KHLAF VLKV+PGTV VCHFL G
Sbjct: 248 KWNLKHLAFQVLKVKPGTVSVCHFLPQG 165
>AV767317
Length = 226
Score = 39.7 bits (91), Expect = 5e-04
Identities = 17/32 (53%), Positives = 21/32 (65%)
Frame = -1
Query: 212 PKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
P ++AF VLKV+PGT PV HF G W S+
Sbjct: 226 PNYVAFPVLKVKPGTGPVGHFFPQGPFFWLSQ 131
>AV418436
Length = 280
Score = 38.9 bits (89), Expect = 8e-04
Identities = 15/23 (65%), Positives = 17/23 (73%)
Frame = -1
Query: 221 KVQPGTVPVCHFLQHGHVIWFSK 243
KV+PGTV VCHFL HV+W K
Sbjct: 280 KVKPGTVSVCHFLPQDHVVWVPK 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,466,818
Number of Sequences: 28460
Number of extensions: 60365
Number of successful extensions: 376
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 364
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 365
length of query: 243
length of database: 4,897,600
effective HSP length: 88
effective length of query: 155
effective length of database: 2,393,120
effective search space: 370933600
effective search space used: 370933600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148532.6


