

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
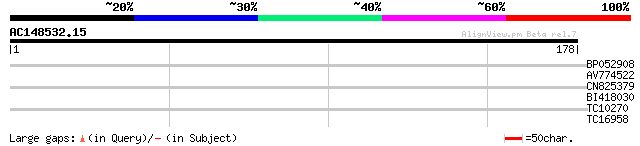
Score E
Sequences producing significant alignments: (bits) Value
BP052908 35 0.010
AV774522 30 0.31
CN825379 29 0.41
BI418030 26 3.4
TC10270 homologue to UP|Q8VXT0 (Q8VXT0) Translation initiation f... 25 5.9
TC16958 similar to UP|Q9FNG9 (Q9FNG9) Gb|AAF02137.1, partial (20%) 25 5.9
>BP052908
Length = 612
Score = 34.7 bits (78), Expect = 0.010
Identities = 22/75 (29%), Positives = 35/75 (46%), Gaps = 7/75 (9%)
Frame = -3
Query: 18 KWGQKRGKGGKKRDIQ------FYQSFTLRGVDYSLFDNVYVK-NDFAEPRIGKIIKIWE 70
K G R G K + + Y + G+ Y L D+ YVK D I +I++++E
Sbjct: 448 KKGSARNSNGAKSETENLQVKSHYSQARVDGILYDLNDDAYVKAEDGNADFIARIVEMFE 269
Query: 71 TPTLERKIKVQWFFR 85
T ++ QWF+R
Sbjct: 268 TVDDKKYFTAQWFYR 224
>AV774522
Length = 541
Score = 29.6 bits (65), Expect = 0.31
Identities = 22/82 (26%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Frame = +2
Query: 11 ESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVD--YSLFDNVYVK--NDFAEPRIGKII 66
ES+ L F + + + GKK D+ S+T+RG + D V ++ + P + ++
Sbjct: 179 ESKALVFLFQMAKTRPGKK-DVD---SYTIRGTNKVVKAGDCVLMRPSDTSKPPYVARVE 346
Query: 67 KIWETPTLERKIKVQWFFRPIE 88
KI + +++V+W++RP E
Sbjct: 347 KIEQDNRSNVRVRVRWYYRPEE 412
>CN825379
Length = 646
Score = 29.3 bits (64), Expect = 0.41
Identities = 26/88 (29%), Positives = 41/88 (46%), Gaps = 5/88 (5%)
Frame = +3
Query: 24 GKG-GKKRDIQFYQSFTLRGVDYSLFDNVYV--KNDFAEPRIGKIIKIWETPTLERKIKV 80
GKG G+KR + F G Y+L D V + + + +P + I I ++ +
Sbjct: 384 GKGRGRKRH---FHCFEFDGNQYALEDPVLLVPEGEDQKPYVAIIKDIAQSVNGSIMVTG 554
Query: 81 QWFFRPIEVSK--CLTWIKIYFNELFFA 106
QWF+RP E K +W ELF++
Sbjct: 555 QWFYRPEEAEKKGGGSWQSCDARELFYS 638
>BI418030
Length = 488
Score = 26.2 bits (56), Expect = 3.4
Identities = 9/29 (31%), Positives = 18/29 (62%)
Frame = +1
Query: 60 PRIGKIIKIWETPTLERKIKVQWFFRPIE 88
P + ++ KI + K++V+W++RP E
Sbjct: 247 PYVARVEKIEQDTRSNVKVRVRWYYRPEE 333
>TC10270 homologue to UP|Q8VXT0 (Q8VXT0) Translation initiation factor
(eIF-1A), partial (98%)
Length = 619
Score = 25.4 bits (54), Expect = 5.9
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +1
Query: 1 KSNSPPAPSPESETLEFKWGQKRGKGGKKR 30
K+ SPP PSP + + +GKGGK R
Sbjct: 67 KAVSPPPPSPTT------MPKNKGKGGKNR 138
>TC16958 similar to UP|Q9FNG9 (Q9FNG9) Gb|AAF02137.1, partial (20%)
Length = 644
Score = 25.4 bits (54), Expect = 5.9
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -1
Query: 6 PAPSPESETLEFKWGQKRGKGGKKR 30
P P+PE E +W + R GGK R
Sbjct: 263 PDPNPEFEKTVMRWRRLRRGGGKCR 189
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,740,148
Number of Sequences: 28460
Number of extensions: 56195
Number of successful extensions: 271
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 269
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 271
length of query: 178
length of database: 4,897,600
effective HSP length: 84
effective length of query: 94
effective length of database: 2,506,960
effective search space: 235654240
effective search space used: 235654240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148532.15


