

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.13 + phase: 0
(303 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
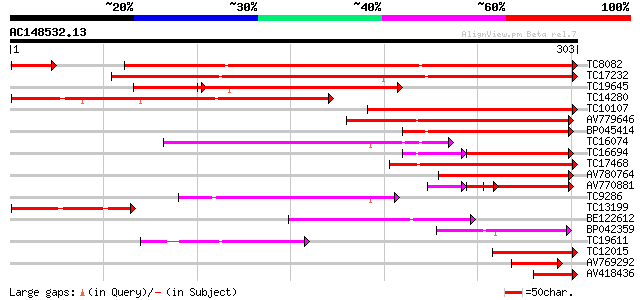
Score E
Sequences producing significant alignments: (bits) Value
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 243 4e-65
TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 238 7e-64
TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing pro... 160 1e-54
TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-contain... 202 7e-53
TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), p... 194 1e-50
AV779646 156 4e-39
BP045414 122 1e-28
TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein... 105 1e-23
TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein... 91 4e-23
TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 101 1e-22
AV780764 94 4e-20
AV770881 62 1e-16
TC9286 similar to UP|O65009 (O65009) BURP domain containing prot... 78 2e-15
TC13199 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing pro... 75 2e-14
BE122612 72 1e-13
BP042359 65 1e-11
TC19611 64 4e-11
TC12015 weakly similar to UP|O65009 (O65009) BURP domain contain... 60 3e-10
AV769292 51 3e-07
AV418436 45 2e-05
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 243 bits (619), Expect = 4e-65
Identities = 120/242 (49%), Positives = 158/242 (64%)
Frame = +1
Query: 62 VNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQ 121
+ + VG+ N AS+ LH+ FF E LH K + +K S+ LPRQ
Sbjct: 910 LRIYVGKESQHENSGASDHHLHEISARVGFFQEDQLHSDNKFYMHLNKIRSSIP-LLPRQ 1086
Query: 122 VANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDF 181
+ IPFS +KM+ ++ L++K SK +I++ TIS CE + GEEK C TSLESM+DF
Sbjct: 1087 ITQQIPFSLDKMKEVLEMLSVKPNSKNAEIMEKTISGCEAPAMDGEEKYCATSLESMMDF 1266
Query: 182 TTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT 241
TSKLGKNV A+ST+V KE+ LQ + GVKKL + + + CH YPY VF CH+
Sbjct: 1267 VTSKLGKNVHAMSTDVQKETKLQNLLVKDGVKKLADDH-VIACHPLAYPYVVFGCHELQK 1443
Query: 242 TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
T AY VPLEG DG RVKA+AVCH DTS+W+P ++AF+VLKV+PGTVPVCH PE H++W+
Sbjct: 1444 TSAYFVPLEGEDGVRVKAVAVCHKDTSKWDPNYVAFRVLKVKPGTVPVCHFFPEGHLLWL 1623
Query: 302 RK 303
K
Sbjct: 1624 SK 1629
Score = 43.1 bits (100), Expect = 6e-05
Identities = 16/24 (66%), Positives = 21/24 (86%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLL 25
LPP++YWK MLP++PMPKA+ LL
Sbjct: 97 LPPEIYWKMMLPSTPMPKAVIELL 168
>TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (33%)
Length = 966
Score = 238 bits (608), Expect = 7e-64
Identities = 124/252 (49%), Positives = 166/252 (65%), Gaps = 3/252 (1%)
Frame = +3
Query: 55 YEGGGTDVNVGVGRSPFIYNYAAS-ETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSN 113
Y+ G ++ + + + Y AS LH+K +FF E++LH GTKL+LQ SK
Sbjct: 24 YKIGAKEIQLHDDTNLYGYKSGASLHKLLHEKTKAGVFFFEENLHPGTKLDLQLSKRKP- 200
Query: 114 AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVT 173
A LPR++A+ IPFS K++ I+ L +K SK +IV+ T+ CE ++GEE++C T
Sbjct: 201 AVPILPRKIADQIPFSLEKLKEILEILVVKPDSKNTEIVEKTMGGCELPSMEGEEQMCAT 380
Query: 174 SLESMVDFTTSKLGKNVEAVSTEVN--KESNLQQYTIASGVKKLGEKNKAVVCHKENYPY 231
SLES+VDF SKLGKNV A+STEV KE+ Q + GVKKL + AV CH YPY
Sbjct: 381 SLESLVDFIASKLGKNVLAISTEVELEKETQSQNLLVKDGVKKLADDGMAV-CHPMTYPY 557
Query: 232 AVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCH 291
V + HK T AY VPLEG DG+RVKA+AVCH DTS+W+P H++F L V+PGTVPVCH
Sbjct: 558 PVCFSHKLVKTSAYFVPLEGEDGARVKAVAVCHRDTSKWDPNHVSFPDLNVKPGTVPVCH 737
Query: 292 LLPEDHVVWIRK 303
+LPE H++W+ K
Sbjct: 738 VLPERHLLWVPK 773
>TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (38%)
Length = 619
Score = 160 bits (406), Expect(2) = 1e-54
Identities = 81/112 (72%), Positives = 98/112 (87%), Gaps = 2/112 (1%)
Frame = +3
Query: 101 TKLNLQFSKTTSNAAT--FLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISE 158
TKLNLQF+KT+++AA FLPRQVA+SIPFSSNK++ ++NK ++K GS+ +IVKNTI+E
Sbjct: 120 TKLNLQFTKTSNDAAATFFLPRQVADSIPFSSNKVDDVLNKFSLKPGSEEAEIVKNTIAE 299
Query: 159 CEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIAS 210
CEE G+ GEEK CVTSLESMVDFTTSKLGKNVEAVST+VNKE LQQYTI +
Sbjct: 300 CEEPGVVGEEKRCVTSLESMVDFTTSKLGKNVEAVSTDVNKEKELQQYTIXA 455
Score = 68.9 bits (167), Expect(2) = 1e-54
Identities = 31/39 (79%), Positives = 33/39 (84%)
Frame = +2
Query: 67 GRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNL 105
G SPF YNYAASETQLH+ PNVALFFLEKDLH G K+ L
Sbjct: 17 GHSPFDYNYAASETQLHENPNVALFFLEKDLHQGNKIEL 133
>TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (54%)
Length = 723
Score = 202 bits (513), Expect = 7e-53
Identities = 114/216 (52%), Positives = 140/216 (64%), Gaps = 44/216 (20%)
Frame = +2
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGT---------------------WVD 40
LPP+LYWKS+LP +PMPKAIT++L+P +W E+K T VD
Sbjct: 83 LPPELYWKSVLPTTPMPKAITDILYP--HWVEDKSTNVGVRKGGVNVDAGKGKPGHTKVD 256
Query: 41 VGKGGVDVGVRKGYYEG-GGTDVNVGVGR----------------------SPFIYNYAA 77
VGKGGV+V KG G GGT+VNVG G+ SPF Y YAA
Sbjct: 257 VGKGGVNVNAGKGSGGGHGGTNVNVGGGKGGGVSVHTGHKGKPVFVNVGSKSPFDYVYAA 436
Query: 78 SETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYII 137
+ETQLH PNVALFFLEK+LH GTK+NL F+KT SN ATFLPR VA+SIPFSS K+E +
Sbjct: 437 TETQLHSDPNVALFFLEKNLHPGTKMNLHFTKT-SNKATFLPRIVADSIPFSSEKVELAL 613
Query: 138 NKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVT 173
NK ++K GS+ Q++KNTI ECE++GI+GEEK C T
Sbjct: 614 NKYSVKAGSEEAQVMKNTIDECEQKGIQGEEKYCAT 721
>TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(65%)
Length = 575
Score = 194 bits (494), Expect = 1e-50
Identities = 88/112 (78%), Positives = 102/112 (90%)
Frame = +2
Query: 192 AVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEG 251
AVST V+KE+ Q+YTIA+GVKKL +KAVVCHK+NYPYAVFYCHK+D+T+AYSVPLEG
Sbjct: 2 AVSTVVDKETEKQRYTIATGVKKLEAGDKAVVCHKQNYPYAVFYCHKSDSTRAYSVPLEG 181
Query: 252 ADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
+G RVKA+AVCHTDTS+WNPKHLAFQVLKV+PGTVPVCH LPEDHVVW+ K
Sbjct: 182 DNGVRVKAVAVCHTDTSQWNPKHLAFQVLKVKPGTVPVCHFLPEDHVVWVSK 337
>AV779646
Length = 510
Score = 156 bits (395), Expect = 4e-39
Identities = 73/121 (60%), Positives = 88/121 (72%)
Frame = -3
Query: 181 FTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTD 240
F TSKLG +V A+STEV KE+ L + GV+KL + + VCH YPY VF+CHK
Sbjct: 508 FITSKLGNDVHAMSTEVTKETKLLYLLVKDGVEKLAD-DYITVCHPMAYPYVVFWCHKLQ 332
Query: 241 TTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
T AY VPLEG DG RVKA+AVCH DTS+W+P H AFQ LKV+PGTVPVCH LPE H++W
Sbjct: 331 KTDAYFVPLEGEDGVRVKAVAVCHKDTSKWDPNHAAFQDLKVKPGTVPVCHFLPEGHLLW 152
Query: 301 I 301
+
Sbjct: 151 L 149
>BP045414
Length = 478
Score = 122 bits (305), Expect = 1e-28
Identities = 56/91 (61%), Positives = 68/91 (74%)
Frame = -2
Query: 211 GVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEW 270
GVKKL + AV CH YPYAVF+CHK T AY VPLEG DG+RV +AVC DTS+W
Sbjct: 471 GVKKLPDDGIAV-CHPLTYPYAVFFCHKLVKTSAYFVPLEGEDGARV*PVAVCPRDTSKW 295
Query: 271 NPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
+P H++F L VQPGTVPVCH+LPE H++W+
Sbjct: 294 DPNHVSFPDLHVQPGTVPVCHVLPERHLLWV 202
>TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(6%)
Length = 844
Score = 105 bits (261), Expect = 1e-23
Identities = 63/160 (39%), Positives = 92/160 (57%), Gaps = 5/160 (3%)
Frame = +2
Query: 83 HDKPNVALFFLEKDLHHGTKLNLQFS-KTTSNAATFLPRQVANSIPFSSNKMEYIINKLN 141
H P + +FF+ KDL G K+ + FS K +S + FLPR+ AN IPFSS ++ ++N +
Sbjct: 353 HMDPELHVFFIPKDLKVGKKMPIYFSMKDSSTSPKFLPREEANQIPFSSKQLPSLLNFFS 532
Query: 142 IKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVE---AVSTEVN 198
I K S + +K T+ +CE + ++GE K C TSLES++D G + + +T +
Sbjct: 533 IPKNSPQAKAMKYTLEQCEFEPMEGETKFCATSLESLLDSARHLFGLSPQFKVLTTTHLT 712
Query: 199 KES-NLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCH 237
K LQ YTI VK+ N + CH YP+AVFYCH
Sbjct: 713 KSHVLLQNYTILE-VKENRVPN-VIGCHPMPYPFAVFYCH 826
>TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2, partial
(12%)
Length = 488
Score = 90.9 bits (224), Expect(2) = 4e-23
Identities = 38/57 (66%), Positives = 45/57 (78%)
Frame = +1
Query: 245 YSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
Y VPLEG DG RV +AVCH DTS+W+P H AF LKV+PGTVPVCH LPE H++W+
Sbjct: 106 YFVPLEGEDGVRVNHVAVCHQDTSKWDPNHAAFPDLKVKPGTVPVCHFLPEGHLLWL 276
Score = 33.1 bits (74), Expect(2) = 4e-23
Identities = 16/34 (47%), Positives = 20/34 (58%)
Frame = +3
Query: 211 GVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKA 244
GV+KL + + VCH YPY VF+CH T A
Sbjct: 6 GVEKLAD-DYITVCHPMAYPYLVFWCH*LPKTHA 104
>TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (11%)
Length = 652
Score = 101 bits (252), Expect = 1e-22
Identities = 45/101 (44%), Positives = 62/101 (60%), Gaps = 1/101 (0%)
Frame = +3
Query: 204 QQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT-TKAYSVPLEGADGSRVKAIAV 262
Q YTI + ++ K V CH YPYAVFYCH ++ + Y V L G +G +V+A+ V
Sbjct: 3 QNYTILENIMRISAP-KMVACHTMPYPYAVFYCHSQESENRVYKVSLVGDNGDKVEAMVV 179
Query: 263 CHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
CH DTS W H++FQ+L V PG+ VCH P D+++W K
Sbjct: 180 CHLDTSHWGSGHVSFQILGVTPGSSSVCHFFPADNLIWAPK 302
>AV780764
Length = 438
Score = 93.6 bits (231), Expect = 4e-20
Identities = 42/72 (58%), Positives = 50/72 (69%)
Frame = -1
Query: 230 PYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPV 289
PY VF+ H +AY VPLEG DG RVK IAVCH TS+W+P H A LKV+PGT PV
Sbjct: 378 PYVVFWFH*LPKPQAYFVPLEGEDGVRVKTIAVCHQHTSKWDPNHAALPNLKVKPGTGPV 199
Query: 290 CHLLPEDHVVWI 301
CH L E H++W+
Sbjct: 198 CHFLLEGHLLWL 163
>AV770881
Length = 403
Score = 62.4 bits (150), Expect(3) = 1e-16
Identities = 26/48 (54%), Positives = 34/48 (70%)
Frame = -3
Query: 254 GSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
G +K + VCH DTS+W+P H AF LKV+PGTVPV LPE ++W+
Sbjct: 311 GLGLKPLLVCHQDTSKWDPNHAAFPDLKVKPGTVPVSQFLPEGPLLWL 168
Score = 29.6 bits (65), Expect(3) = 1e-16
Identities = 13/17 (76%), Positives = 14/17 (81%)
Frame = -2
Query: 245 YSVPLEGADGSRVKAIA 261
Y VPLEG DG RVKA+A
Sbjct: 339 YFVPLEGEDGVRVKAVA 289
Score = 29.6 bits (65), Expect(3) = 1e-16
Identities = 11/21 (52%), Positives = 12/21 (56%)
Frame = -1
Query: 224 CHKENYPYAVFYCHKTDTTKA 244
CH YPY VF+CH T A
Sbjct: 403 CHPMAYPYGVFWCHGLPKTHA 341
>TC9286 similar to UP|O65009 (O65009) BURP domain containing protein,
partial (6%)
Length = 805
Score = 78.2 bits (191), Expect = 2e-15
Identities = 45/122 (36%), Positives = 67/122 (54%), Gaps = 4/122 (3%)
Frame = +1
Query: 91 FFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQ 150
FF DL+ G + LQF + FL R+ N IPFSS+++ ++ +I + S +
Sbjct: 445 FFALDDLYAGNVMTLQFP--VQEISHFLSRKETNYIPFSSSQLPSVLQLFSIPEDSPKAK 618
Query: 151 IVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVE----AVSTEVNKESNLQQY 206
++ T+ +CE + I GE K+C TSLESM++F S +G N E S S LQ+Y
Sbjct: 619 SMRGTLGQCEGEPITGERKICATSLESMLEFVDSVIGSNTEGDLLTTSHPSPSGSPLQKY 798
Query: 207 TI 208
TI
Sbjct: 799 TI 804
>TC13199 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (19%)
Length = 426
Score = 74.7 bits (182), Expect = 2e-14
Identities = 40/66 (60%), Positives = 47/66 (70%)
Frame = +2
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+LYWKS LP +PMPKAIT+LLH W+E+K T V VGKGGV+V G + GGT
Sbjct: 119 LTPELYWKSKLPTTPMPKAITDLLH--SDWTEDKSTSVAVGKGGVNVDA--GKTKPGGTA 286
Query: 62 VNVGVG 67
VNVG G
Sbjct: 287 VNVGKG 304
>BE122612
Length = 386
Score = 72.0 bits (175), Expect = 1e-13
Identities = 37/101 (36%), Positives = 57/101 (55%), Gaps = 1/101 (0%)
Frame = +2
Query: 150 QIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTE-VNKESNLQQYTI 208
+++++++ ECE +GE K CV S+E MVDF TS LG+NV +TE VN +
Sbjct: 35 KMMRDSLQECERVPSRGETKRCVGSIEDMVDFATSVLGRNVVVRTTENVNGSNKKVMVGR 214
Query: 209 ASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPL 249
GV G+ ++V CH+ +PY ++YCH + Y L
Sbjct: 215 VQGVNG-GKVTRSVSCHQSLFPYLLYYCHSLPKVRVYQADL 334
>BP042359
Length = 404
Score = 65.1 bits (157), Expect = 1e-11
Identities = 27/75 (36%), Positives = 42/75 (56%), Gaps = 3/75 (4%)
Frame = -3
Query: 229 YPYAVFYCHKTDTTKAYSVPLEGADGSRVK---AIAVCHTDTSEWNPKHLAFQVLKVQPG 285
YPY ++YCH + Y + D +++K +A+CH DTS W P+H AF L PG
Sbjct: 402 YPYLLYYCHSVPQVRVYEAEILDVD-TKLKINHGVAICHLDTSAWGPEHGAFVALGAGPG 226
Query: 286 TVPVCHLLPEDHVVW 300
+ VCH + ++ + W
Sbjct: 225 KIEVCHWIFQNDMTW 181
>TC19611
Length = 533
Score = 63.5 bits (153), Expect = 4e-11
Identities = 36/90 (40%), Positives = 50/90 (55%)
Frame = -3
Query: 71 FIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSS 130
F Y A ETQ FF E LH G K N+ +K S +LPR++A+ IPFS
Sbjct: 258 FYAYYVAKETQSRS------FFEEYQLHAGNKFNVNLNKIKS-VVPWLPREIADHIPFSL 100
Query: 131 NKMEYIINKLNIKKGSKGVQIVKNTISECE 160
+KM I+ L++K SK ++V+ TI +CE
Sbjct: 99 DKMNEILEMLSVKLESKNAEMVEKTIGDCE 10
Score = 33.1 bits (74), Expect = 0.060
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = -2
Query: 2 LPPQLYWKSMLPNSPMPKAITNL 24
LP +L WK LP +P+PKAIT L
Sbjct: 445 LPAELDWKRKLPATPVPKAITEL 377
>TC12015 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (12%)
Length = 464
Score = 60.5 bits (145), Expect = 3e-10
Identities = 21/45 (46%), Positives = 30/45 (66%)
Frame = +1
Query: 259 AIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
A+ VCH DTS+W P H+ F+ L ++ G PVCH P H++W+ K
Sbjct: 1 ALGVCHLDTSDWTPSHIIFKQLGIEAGKAPVCHFFPIKHLMWVAK 135
>AV769292
Length = 250
Score = 50.8 bits (120), Expect = 3e-07
Identities = 21/27 (77%), Positives = 24/27 (88%)
Frame = -3
Query: 269 EWNPKHLAFQVLKVQPGTVPVCHLLPE 295
+WN KHLAFQVLKV+PGTV VCH LP+
Sbjct: 248 KWNLKHLAFQVLKVKPGTVSVCHFLPQ 168
>AV418436
Length = 280
Score = 44.7 bits (104), Expect = 2e-05
Identities = 17/23 (73%), Positives = 20/23 (86%)
Frame = -1
Query: 281 KVQPGTVPVCHLLPEDHVVWIRK 303
KV+PGTV VCH LP+DHVVW+ K
Sbjct: 280 KVKPGTVSVCHFLPQDHVVWVPK 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,611,989
Number of Sequences: 28460
Number of extensions: 75267
Number of successful extensions: 379
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 350
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 361
length of query: 303
length of database: 4,897,600
effective HSP length: 90
effective length of query: 213
effective length of database: 2,336,200
effective search space: 497610600
effective search space used: 497610600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148532.13


