

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.11 + phase: 0
(308 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
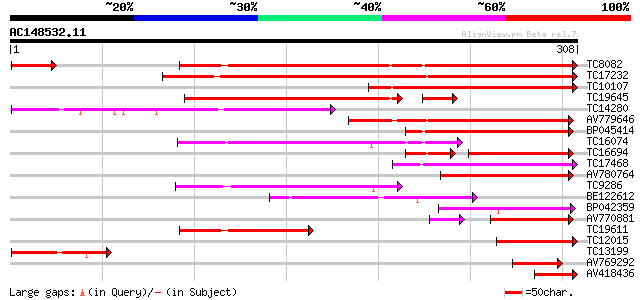
Score E
Sequences producing significant alignments: (bits) Value
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 220 3e-58
TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 216 4e-57
TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), p... 178 1e-45
TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing pro... 150 4e-41
TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-contain... 145 8e-36
AV779646 141 1e-34
BP045414 114 2e-26
TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein... 108 9e-25
TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein... 85 2e-21
TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific ... 97 3e-21
AV780764 89 9e-19
TC9286 similar to UP|O65009 (O65009) BURP domain containing prot... 85 2e-17
BE122612 72 9e-14
BP042359 69 1e-12
AV770881 59 2e-12
TC19611 61 3e-10
TC12015 weakly similar to UP|O65009 (O65009) BURP domain contain... 59 1e-09
TC13199 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing pro... 50 4e-07
AV769292 49 8e-07
AV418436 46 7e-06
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 220 bits (560), Expect = 3e-58
Identities = 108/216 (50%), Positives = 142/216 (65%)
Frame = +1
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF + LH K + + S+ LPR++ IPFS +KM+ +L S+K SK
Sbjct: 997 FFQEDQLHSDNKFYMHLNKIRSS---IPLLPRQITQQIPFSLDKMKEVLEMLSVKPNSKN 1167
Query: 153 AEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYV 212
AEI+++TIS CEA + GEEK C TSLESM+DF SKLG NV A+ST+V K + LQ +
Sbjct: 1168 AEIMEKTISGCEAPAMDGEEKYCATSLESMMDFVTSKLGKNVHAMSTDVQKETK-LQNLL 1344
Query: 213 IAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDT 272
+ GVKKL + + I CH YPY VF CH+ T Y VPLEG DG VK +AVCH DT
Sbjct: 1345 VKDGVKKLADDH-VIACHPLAYPYVVFGCHELQKTSAYFVPLEGEDGVRVKAVAVCHKDT 1521
Query: 273 SEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
S+W+P ++AF VLKV+PGTVP+CH P+ H++W+SK
Sbjct: 1522 SKWDPNYVAFRVLKVKPGTVPVCHFFPEGHLLWLSK 1629
Score = 40.0 bits (92), Expect = 5e-04
Identities = 15/24 (62%), Positives = 20/24 (82%)
Frame = +1
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLL 25
LPP++YW+ MLP++PMPKA LL
Sbjct: 97 LPPEIYWKMMLPSTPMPKAVIELL 168
>TC17232 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (33%)
Length = 966
Score = 216 bits (550), Expect = 4e-57
Identities = 108/226 (47%), Positives = 146/226 (63%), Gaps = 1/226 (0%)
Frame = +3
Query: 84 LDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNK 143
L +K +FF +++LH GTKL+LQ + LPR++A+ IPFS K++ IL
Sbjct: 108 LHEKTKAGVFFFEENLHPGTKLDLQLSK---RKPAVPILPRKIADQIPFSLEKLKEILEI 278
Query: 144 FSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDK 203
+K SK EIV++T+ CE ++GEE++C TSLES+VDF SKLG NV A+STEV+
Sbjct: 279 LVVKPDSKNTEIVEKTMGGCELPSMEGEEQMCATSLESLVDFIASKLGKNVLAISTEVEL 458
Query: 204 NSNGLQQYVIAK-GVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMV 262
Q ++ K GVKKL + VCH YPY V + HK T Y VPLEG DG V
Sbjct: 459 EKETQSQNLLVKDGVKKLADDGMA-VCHPMTYPYPVCFSHKLVKTSAYFVPLEGEDGARV 635
Query: 263 KTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
K +AVCH DTS+W+P H++F L V+PGTVP+CH+LP+ H++WV K
Sbjct: 636 KAVAVCHRDTSKWDPNHVSFPDLNVKPGTVPVCHVLPERHLLWVPK 773
>TC10107 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(65%)
Length = 575
Score = 178 bits (451), Expect = 1e-45
Identities = 82/113 (72%), Positives = 94/113 (82%)
Frame = +2
Query: 196 AVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLE 255
AVST VDK + Q+Y IA GVKKL +K +VCHK+NYPYAVFYCHK+DST YSVPLE
Sbjct: 2 AVSTVVDKETEK-QRYTIATGVKKLEAGDKAVVCHKQNYPYAVFYCHKSDSTRAYSVPLE 178
Query: 256 GVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
G +G VK +AVCHTDTS+WNPKHLAF VLKV+PGTVP+CH LP+DHVVWVSK
Sbjct: 179 GDNGVRVKAVAVCHTDTSQWNPKHLAFQVLKVKPGTVPVCHFLPEDHVVWVSK 337
>TC19645 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (38%)
Length = 619
Score = 150 bits (379), Expect(2) = 4e-41
Identities = 78/118 (66%), Positives = 95/118 (80%)
Frame = +3
Query: 96 KKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEI 155
K+ TKLNLQF +T+++ T FLPR+VA+SIPFSSNK++++LNKFS+K GS+EAEI
Sbjct: 99 KRTCIRXTKLNLQFTKTSNDAAATFFLPRQVADSIPFSSNKVDDVLNKFSLKPGSEEAEI 278
Query: 156 VKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVI 213
VK TI+ECE G+ GEEK CVTSLESMVDFT SKLG NVEAVST+V+K LQQY I
Sbjct: 279 VKNTIAECEEPGVVGEEKRCVTSLESMVDFTTSKLGKNVEAVSTDVNKEKE-LQQYTI 449
Score = 37.4 bits (85), Expect = 0.003
Identities = 16/28 (57%), Positives = 20/28 (71%)
Frame = +2
Query: 80 SQIQLDDKQNVTLFFLKKDLHHGTKLNL 107
S+ QL + NV LFFL+KDLH G K+ L
Sbjct: 50 SETQLHENPNVALFFLEKDLHQGNKIEL 133
Score = 33.9 bits (76), Expect(2) = 4e-41
Identities = 13/19 (68%), Positives = 15/19 (78%)
Frame = +2
Query: 225 KTIVCHKENYPYAVFYCHK 243
K ++ HKENYPYAVF C K
Sbjct: 482 KLLLFHKENYPYAVFXCXK 538
>TC14280 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (54%)
Length = 723
Score = 145 bits (366), Expect = 8e-36
Identities = 89/218 (40%), Positives = 121/218 (54%), Gaps = 42/218 (19%)
Frame = +2
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKAR-----------------------D 38
LPP+LYW+S+LP +PMPKA T++L+P +W ++K+ D
Sbjct: 83 LPPELYWKSVLPTTPMPKAITDILYP--HWVEDKSTNVGVRKGGVNVDAGKGKPGHTKVD 256
Query: 39 ASNGGLDVGVRKGYEGG--GTYLN-------------DEKIIPLIYFYPIPIPLN----E 79
GG++V KG GG GT +N K P+ P +
Sbjct: 257 VGKGGVNVNAGKGSGGGHGGTNVNVGGGKGGGVSVHTGHKGKPVFVNVGSKSPFDYVYAA 436
Query: 80 SQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMEN 139
++ QL NV LFFL+K+LH GTK+NL F +T+ N FLPR VA+SIPFSS K+E
Sbjct: 437 TETQLHSDPNVALFFLEKNLHPGTKMNLHFTKTS---NKATFLPRIVADSIPFSSEKVEL 607
Query: 140 ILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVT 177
LNK+S+K GS+EA+++K TI ECE GI+GEEK C T
Sbjct: 608 ALNKYSVKAGSEEAQVMKNTIDECEQKGIQGEEKYCAT 721
>AV779646
Length = 510
Score = 141 bits (356), Expect = 1e-34
Identities = 69/123 (56%), Positives = 87/123 (70%), Gaps = 1/123 (0%)
Frame = -3
Query: 185 FTISKLGNNVEAVSTEVDKNSNGLQQYVIAK-GVKKLGEKNKTIVCHKENYPYAVFYCHK 243
F SKLGN+V A+STEV K + L Y++ K GV+KL + T VCH YPY VF+CHK
Sbjct: 508 FITSKLGNDVHAMSTEVTKETKLL--YLLVKDGVEKLADDYIT-VCHPMAYPYVVFWCHK 338
Query: 244 TDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHV 303
T+ Y VPLEG DG VK +AVCH DTS+W+P H AF LKV+PGTVP+CH LP+ H+
Sbjct: 337 LQKTDAYFVPLEGEDGVRVKAVAVCHKDTSKWDPNHAAFQDLKVKPGTVPVCHFLPEGHL 158
Query: 304 VWV 306
+W+
Sbjct: 157 LWL 149
>BP045414
Length = 478
Score = 114 bits (286), Expect = 2e-26
Identities = 52/91 (57%), Positives = 64/91 (70%)
Frame = -2
Query: 216 GVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEW 275
GVKKL + VCH YPYAVF+CHK T Y VPLEG DG V +AVC DTS+W
Sbjct: 471 GVKKLPDDG-IAVCHPLTYPYAVFFCHKLVKTSAYFVPLEGEDGARV*PVAVCPRDTSKW 295
Query: 276 NPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
+P H++F L VQPGTVP+CH+LP+ H++WV
Sbjct: 294 DPNHVSFPDLHVQPGTVPVCHVLPERHLLWV 202
>TC16074 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(6%)
Length = 844
Score = 108 bits (271), Expect = 9e-25
Identities = 65/158 (41%), Positives = 93/158 (58%), Gaps = 3/158 (1%)
Frame = +2
Query: 92 LFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSK 151
+FF+ KDL G K+ + F S+ + KFLPRE AN IPFSS ++ ++LN FSI + S
Sbjct: 374 VFFIPKDLKVGKKMPIYFSMKDSSTS-PKFLPREEANQIPFSSKQLPSLLNFFSIPKNSP 550
Query: 152 EAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVE---AVSTEVDKNSNGL 208
+A+ +K T+ +CE ++GE K C TSLES++D G + + +T + K+ L
Sbjct: 551 QAKAMKYTLEQCEFEPMEGETKFCATSLESLLDSARHLFGLSPQFKVLTTTHLTKSHVLL 730
Query: 209 QQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDS 246
Q Y I + VK+ N I CH YP+AVFYCH S
Sbjct: 731 QNYTILE-VKENRVPN-VIGCHPMPYPFAVFYCHSQQS 838
>TC16694 similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2, partial
(12%)
Length = 488
Score = 85.1 bits (209), Expect(2) = 2e-21
Identities = 35/57 (61%), Positives = 44/57 (76%)
Frame = +1
Query: 250 YSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
Y VPLEG DG V +AVCH DTS+W+P H AF LKV+PGTVP+CH LP+ H++W+
Sbjct: 106 YFVPLEGEDGVRVNHVAVCHQDTSKWDPNHAAFPDLKVKPGTVPVCHFLPEGHLLWL 276
Score = 33.1 bits (74), Expect(2) = 2e-21
Identities = 15/27 (55%), Positives = 18/27 (66%)
Frame = +3
Query: 216 GVKKLGEKNKTIVCHKENYPYAVFYCH 242
GV+KL + T VCH YPY VF+CH
Sbjct: 6 GVEKLADDYIT-VCHPMAYPYLVFWCH 83
>TC17468 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (11%)
Length = 652
Score = 97.1 bits (240), Expect = 3e-21
Identities = 42/101 (41%), Positives = 60/101 (58%), Gaps = 1/101 (0%)
Frame = +3
Query: 209 QQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDS-TEVYSVPLEGVDGNMVKTIAV 267
Q Y I + + ++ K + CH YPYAVFYCH +S VY V L G +G+ V+ + V
Sbjct: 3 QNYTILENIMRISAP-KMVACHTMPYPYAVFYCHSQESENRVYKVSLVGDNGDKVEAMVV 179
Query: 268 CHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
CH DTS W H++F +L V PG+ +CH P D+++W K
Sbjct: 180 CHLDTSHWGSGHVSFQILGVTPGSSSVCHFFPADNLIWAPK 302
>AV780764
Length = 438
Score = 89.0 bits (219), Expect = 9e-19
Identities = 39/72 (54%), Positives = 49/72 (67%)
Frame = -1
Query: 235 PYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPI 294
PY VF+ H + Y VPLEG DG VKTIAVCH TS+W+P H A LKV+PGT P+
Sbjct: 378 PYVVFWFH*LPKPQAYFVPLEGEDGVRVKTIAVCHQHTSKWDPNHAALPNLKVKPGTGPV 199
Query: 295 CHILPQDHVVWV 306
CH L + H++W+
Sbjct: 198 CHFLLEGHLLWL 163
>TC9286 similar to UP|O65009 (O65009) BURP domain containing protein,
partial (6%)
Length = 805
Score = 84.7 bits (208), Expect = 2e-17
Identities = 48/126 (38%), Positives = 74/126 (58%), Gaps = 3/126 (2%)
Frame = +1
Query: 91 TLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGS 150
T FF DL+ G + LQF ++ FL R+ N IPFSS+++ ++L FSI E S
Sbjct: 439 TGFFALDDLYAGNVMTLQFPVQEISH----FLSRKETNYIPFSSSQLPSVLQLFSIPEDS 606
Query: 151 KEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEA---VSTEVDKNSNG 207
+A+ ++ T+ +CE I GE K+C TSLESM++F S +G+N E ++ + +
Sbjct: 607 PKAKSMRGTLGQCEGEPITGERKICATSLESMLEFVDSVIGSNTEGDLLTTSHPSPSGSP 786
Query: 208 LQQYVI 213
LQ+Y I
Sbjct: 787 LQKYTI 804
>BE122612
Length = 386
Score = 72.4 bits (176), Expect = 9e-14
Identities = 41/115 (35%), Positives = 62/115 (53%), Gaps = 2/115 (1%)
Frame = +2
Query: 142 NKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEV 201
N GS E ++++ ++ ECE +GE K CV S+E MVDF S LG NV +TE
Sbjct: 2 NSARADNGSME-KMMRDSLQECERVPSRGETKRCVGSIEDMVDFATSVLGRNVVVRTTE- 175
Query: 202 DKNSNGLQQYVIAKGVKKL--GEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPL 254
N NG + V+ V+ + G+ +++ CH+ +PY ++YCH VY L
Sbjct: 176 --NVNGSNKKVMVGRVQGVNGGKVTRSVSCHQSLFPYLLYYCHSLPKVRVYQADL 334
>BP042359
Length = 404
Score = 68.6 bits (166), Expect = 1e-12
Identities = 28/76 (36%), Positives = 41/76 (53%), Gaps = 2/76 (2%)
Frame = -3
Query: 234 YPYAVFYCHKTDSTEVYSVPLEGVDGNMVKT--IAVCHTDTSEWNPKHLAFYVLKVQPGT 291
YPY ++YCH VY + VD + +A+CH DTS W P+H AF L PG
Sbjct: 402 YPYLLYYCHSVPQVRVYEAEILDVDTKLKINHGVAICHLDTSAWGPEHGAFVALGAGPGK 223
Query: 292 VPICHILPQDHVVWVS 307
+ +CH + Q+ + W +
Sbjct: 222 IEVCHWIFQNDMTWTT 175
>AV770881
Length = 403
Score = 58.9 bits (141), Expect(2) = 2e-12
Identities = 23/45 (51%), Positives = 33/45 (73%)
Frame = -3
Query: 262 VKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
+K + VCH DTS+W+P H AF LKV+PGTVP+ LP+ ++W+
Sbjct: 302 LKPLLVCHQDTSKWDPNHAAFPDLKVKPGTVPVSQFLPEGPLLWL 168
Score = 28.9 bits (63), Expect(2) = 2e-12
Identities = 10/19 (52%), Positives = 11/19 (57%)
Frame = -1
Query: 229 CHKENYPYAVFYCHKTDST 247
CH YPY VF+CH T
Sbjct: 403 CHPMAYPYGVFWCHGLPKT 347
>TC19611
Length = 533
Score = 60.8 bits (146), Expect = 3e-10
Identities = 32/73 (43%), Positives = 45/73 (60%)
Frame = -3
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF + LH G K N+ + S +LPRE+A+ IPFS +KM IL S+K SK
Sbjct: 216 FFEEYQLHAGNKFNVNLNKIKSV---VPWLPREIADHIPFSLDKMNEILEMLSVKLESKN 46
Query: 153 AEIVKRTISECEA 165
AE+V++TI +CE+
Sbjct: 45 AEMVEKTIGDCES 7
>TC12015 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (12%)
Length = 464
Score = 58.5 bits (140), Expect = 1e-09
Identities = 20/44 (45%), Positives = 29/44 (65%)
Frame = +1
Query: 265 IAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
+ VCH DTS+W P H+ F L ++ G P+CH P H++WV+K
Sbjct: 4 LGVCHLDTSDWTPSHIIFKQLGIEAGKAPVCHFFPIKHLMWVAK 135
>TC13199 similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (19%)
Length = 426
Score = 50.4 bits (119), Expect = 4e-07
Identities = 25/56 (44%), Positives = 36/56 (63%), Gaps = 2/56 (3%)
Frame = +2
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDAS--NGGLDVGVRKGYEGG 55
L P+LYW+S LP +PMPKA T+LLH W+++K+ + GG++V K GG
Sbjct: 119 LTPELYWKSKLPTTPMPKAITDLLH--SDWTEDKSTSVAVGKGGVNVDAGKTKPGG 280
>AV769292
Length = 250
Score = 49.3 bits (116), Expect = 8e-07
Identities = 20/27 (74%), Positives = 23/27 (85%)
Frame = -3
Query: 274 EWNPKHLAFYVLKVQPGTVPICHILPQ 300
+WN KHLAF VLKV+PGTV +CH LPQ
Sbjct: 248 KWNLKHLAFQVLKVKPGTVSVCHFLPQ 168
>AV418436
Length = 280
Score = 46.2 bits (108), Expect = 7e-06
Identities = 18/23 (78%), Positives = 20/23 (86%)
Frame = -1
Query: 286 KVQPGTVPICHILPQDHVVWVSK 308
KV+PGTV +CH LPQDHVVWV K
Sbjct: 280 KVKPGTVSVCHFLPQDHVVWVPK 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,720,927
Number of Sequences: 28460
Number of extensions: 77382
Number of successful extensions: 362
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 347
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 353
length of query: 308
length of database: 4,897,600
effective HSP length: 90
effective length of query: 218
effective length of database: 2,336,200
effective search space: 509291600
effective search space used: 509291600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148532.11


