

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
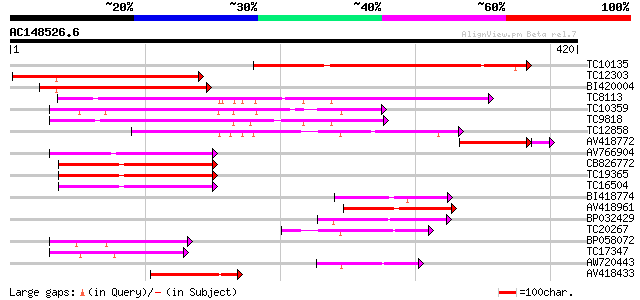
Query= AC148526.6 + phase: 0 /pseudo
(420 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10135 189 1e-48
TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transpo... 172 9e-44
BI420004 162 1e-40
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 116 8e-27
TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partia... 102 1e-22
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 102 1e-22
TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter,... 99 1e-21
AV418772 88 5e-20
AV766904 83 1e-16
CB826772 77 5e-15
TC19365 weakly similar to UP|O23213 (O23213) Sugar transporter l... 74 3e-14
TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporte... 69 2e-12
BI418774 65 3e-11
AV418961 62 2e-10
BP032429 54 6e-08
TC20267 similar to UP|Q7XA52 (Q7XA52) Monosaccharide transporter... 53 8e-08
BP058072 49 1e-06
TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter... 49 1e-06
AW720443 49 2e-06
AV418433 46 1e-05
>TC10135
Length = 1501
Score = 189 bits (479), Expect = 1e-48
Identities = 103/208 (49%), Positives = 137/208 (65%), Gaps = 2/208 (0%)
Frame = +2
Query: 181 VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDV 240
+YR+ +EEE+K IL KIY P +V+ E++A+ ES+E+E E S+ LK
Sbjct: 179 LYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLK----TTT 346
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVS 300
VRRGLYAG+ +Q QQ VGINT+MYYSPTIVQ AG ASN TA LSL+TSGLNA G+I+S
Sbjct: 347 VRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILS 526
Query: 301 MVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTT 360
+ ID+ GR+KL LISL G+ +SL L+V F ++ HAP++S ++ + N+TC +
Sbjct: 527 IYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHY--NNTCPDFRA 700
Query: 361 APNHLSWNCMQCL--HEDCAFCANSQNE 386
A N W CM CL C FCA S N+
Sbjct: 701 AVNPGRWTCMTCLKASPSCGFCAASPNK 784
>TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transporter like
protein {Arabidopsis thaliana;}, partial (25%)
Length = 545
Score = 172 bits (436), Expect = 9e-44
Identities = 85/144 (59%), Positives = 111/144 (77%), Gaps = 3/144 (2%)
Frame = +3
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG P AD + F EC + ++PY++R A S GGLLFGYDTGVISGA LYI+D+F+ V
Sbjct: 114 EGGVPEADMSAFRECLSLSWKNPYVLRRAFSAGIGGLLFGYDTGVISGALLYIKDDFKAV 293
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV 119
D+K WLQE IVS A AGAIIGAA GG++ND+ GR+ I++AD +F+ G+++MAAAP P +
Sbjct: 294 DRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAI 473
Query: 120 IIIGRLLVGLGVGAASMTAPLYIS 143
++ GR+ VG GVG ASM +PLYIS
Sbjct: 474 LLFGRVFVGFGVGMASMASPLYIS 545
>BI420004
Length = 547
Score = 162 bits (409), Expect = 1e-40
Identities = 82/130 (63%), Positives = 104/130 (79%), Gaps = 3/130 (2%)
Frame = +1
Query: 23 ESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAII 79
++PY++ + + GGLLFGYDTGVISGA LYI+D+FE+V + LQETIVSMA GAII
Sbjct: 157 KNPYILGVTAAAGIGGLLFGYDTGVISGALLYIKDDFEEVRRSSLLQETIVSMALVGAII 336
Query: 80 GAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAP 139
GAA GG++ND GRKK L ADVVF G+LVMAAAP + +I GRLLVGLG+G AS+TAP
Sbjct: 337 GAATGGWINDAFGRKKATLSADVVFTLGSLVMAAAPDAYFVIFGRLLVGLGIGVASVTAP 516
Query: 140 LYISEASPAK 149
+YI+E+SP++
Sbjct: 517 VYIAESSPSE 546
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (93%)
Length = 1865
Score = 116 bits (290), Expect = 8e-27
Identities = 100/365 (27%), Positives = 161/365 (43%), Gaps = 42/365 (11%)
Frame = +2
Query: 36 LLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKK 95
++ GYDTGV+SGA L+I+++ D + QE + + + A++G G +D +GR+
Sbjct: 152 IISGYDTGVMSGALLFIKEDIGISDTQ---QEVLAGILNICALVGCLAAGKTSDYIGRRY 322
Query: 96 TILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGAL- 154
TI +A ++F+ GA+ M P +++ GR + GLGVG A TAP+Y +E S A RG L
Sbjct: 323 TIFLASILFLVGAVFMGYGPNFAILMFGRCVCGLGVGFALTTAPVYSAELSSASTRGFLT 502
Query: 155 ----VC--------RLHQWFTHN-----RWSISL-------LPHQFGIYQ-------VYR 183
VC + +F W + L L GI +
Sbjct: 503 SLPEVCIGLGIFIGYISNYFLGKLALTLGWRLMLGLAAIPSLGLALGILTMPESPRWLVM 682
Query: 184 QSKEEEAKQILSKIYRPGEVEEEMKAMHESIEA---EKAEDGLIGH-SLAQKLKGAWS-- 237
Q + AK++L ++ E E E++ + A EK D + + +G W
Sbjct: 683 QGRLGCAKKVLLEVSNTTE-EAELRFRDIIVSAGFDEKCNDEFVKQPQKSHHGEGVWKEL 859
Query: 238 ----NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLN 293
VRR L A + + + GI +M Y P I + AG+ + ++ T
Sbjct: 860 FLRPTPPVRRMLIAAVGIHFFEHATGIEAVMLYGPRIFKKAGVTTKDRLLLATIGTGLTK 1039
Query: 294 AVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNS 353
+S L+DR GRR+L+ IS+ G+ L L + + + L SLS
Sbjct: 1040ITFLTISTFLLDRVGRRRLLQISVAGMIFGLTILGFSLTMVEYSSEKLVWALSLSIVATY 1219
Query: 354 TCKAY 358
T A+
Sbjct: 1220TYVAF 1234
>TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partial (90%)
Length = 1099
Score = 102 bits (254), Expect = 1e-22
Identities = 85/303 (28%), Positives = 133/303 (43%), Gaps = 53/303 (17%)
Frame = +2
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKPWLQETI---------------VS 71
LA +GGL+FGYD GV G + ++ F V KK L++ + S
Sbjct: 227 LAATGGLMFGYDVGVSGGVTSMPHFLHKFFPDVYKKTVLEKELDSNYCKYDNQGLQLFTS 406
Query: 72 MASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGV 131
++ F Y +GR+ T+L+A V F+AG + A ++I+GR+L+G GV
Sbjct: 407 SLYLAGLVSTFFASYTTRTLGRRMTMLIAGVFFIAGVVFNVTAQNLAMLIVGRILLGCGV 586
Query: 132 GAASMTAPLYISEASPAKIRGA-------------LVCRLHQWFTH---NRWSISLLPHQ 175
G A+ PL++SE +P++IRGA L L + T+ W L
Sbjct: 587 GFANQAVPLFLSEIAPSRIRGALNILFQLNITIGILFANLINYGTNKIKGGWGWRLSLGL 766
Query: 176 FGIYQV----------------YRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKA 219
GI V + + EE K +L KI VE E E +EA +
Sbjct: 767 AGIPAVLLTVGALLVVDTPNSLIERGRTEEGKAVLKKIRGTDNVEPEFL---ELVEASR- 934
Query: 220 EDGLIGHSLAQKLKGAWSNDVVRRG---LYAGITVQVVQQIVGINTIMYYSPTIVQFAGI 276
+A+++K + N + RR + I +Q+ QQ GIN IM+Y+P + G
Sbjct: 935 --------VAKEVKDPFRNLLKRRNRPQIIISIALQIFQQFTGINAIMFYAPVLFNTLGF 1090
Query: 277 ASN 279
++
Sbjct: 1091KND 1099
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 102 bits (254), Expect = 1e-22
Identities = 77/295 (26%), Positives = 137/295 (46%), Gaps = 44/295 (14%)
Frame = +2
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
LA +L GYD GV+SGA++YI+ + + D K E ++ + + ++IG+ G +D
Sbjct: 254 LASMTSILLGYDIGVMSGAAIYIKRDLKVSDVKI---EILLGIINLYSLIGSGLAGRTSD 424
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR+ TI+ A +F GAL+M +P W ++ GR + G+G+G A M AP+Y +E SPA
Sbjct: 425 WIGRRYTIVFAGAIFFVGALLMGFSPNYWFLMFGRFIAGIGIGYALMIAPVYTAEVSPAS 604
Query: 150 IRGALVCRLHQWFTH-------NRWSISLLPHQFG-------------IYQVYRQSKEEE 189
RG L + + ++ S L + G I V + E
Sbjct: 605 SRGFLTSFPEVFINGGILLGYISNFAFSKLSLKVGWRMMLGVGALPSVILGVGVLAMPES 784
Query: 190 AKQILSKIYRPGEVEEEMKAMHESIEA------------------EKAEDGLIGHSLAQK 231
+ ++ + G + + +K ++++ ++ E D ++ S
Sbjct: 785 PRWLVMR----GRLGDAIKVLNKTSDSPEEAQLRLADIKRAAGIPESCTDDVVEVSKRST 952
Query: 232 LKGAWS------NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNS 280
+G W +R + A + + QQ GI+ ++ YSPTI + AGI S++
Sbjct: 953 GEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGIDAVVLYSPTIFEKAGIKSDT 1117
>TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter, partial
(51%)
Length = 882
Score = 99.0 bits (245), Expect = 1e-21
Identities = 78/286 (27%), Positives = 131/286 (45%), Gaps = 40/286 (13%)
Frame = +3
Query: 91 MGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKI 150
MG + T+++ + FV+GAL A W++I+GRLL+G G+G A+ + P+Y+SE +P K
Sbjct: 33 MGMRATMIVGGLFFVSGALFNGLADGIWMLIVGRLLLGFGIGCANQSVPIYLSEMAPYKY 212
Query: 151 RGAL-------------VCRLHQWF-----THNRWSISL----LPHQFGIY--------- 179
RG L V L ++ W +SL +P +
Sbjct: 213 RGGLNMCFQLSITIGIFVANLFNYYFAKILNGQGWRLSLGLGAVPAVIFVVGSLCLPDSP 392
Query: 180 -QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSN 238
+ + + E A+Q L KI ++E E+K + + EA + +K W
Sbjct: 393 SSLVARGRHEAARQELVKIRGTTDIEAELKDIITASEA------------LENVKHPWKT 536
Query: 239 DVVRR---GLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSG-LNA 294
+ R+ L + + QQ G+N I +Y+P + F I TA +S V G
Sbjct: 537 LLERKYRPQLVFAVCIPFFQQFTGLNVITFYAP--ILFRTIGFGPTASLMSAVIIGSFKP 710
Query: 295 VGTIVSMVLIDRFGRRKLMLIS----LIGIFVSLVTLSVTFNQAAH 336
V T++S+ ++D+FGRR L L LI + + ++VTF + +
Sbjct: 711 VSTLISIFVVDKFGRRTLFLEGGAQMLICQIIMTIAIAVTFGTSGN 848
>AV418772
Length = 231
Score = 87.8 bits (216), Expect(2) = 5e-20
Identities = 36/53 (67%), Positives = 45/53 (83%)
Frame = +3
Query: 334 AAHHAPSLSIQDSLSFGGNSTCKAYTTAPNHLSWNCMQCLHEDCAFCANSQNE 386
AAHH+P+++ QD+L FGGNSTC+AYT APN SW+CMQCL DCAFCAN ++E
Sbjct: 6 AAHHSPAVNNQDTLIFGGNSTCQAYTKAPNFSSWSCMQCLQADCAFCANGKSE 164
Score = 26.6 bits (57), Expect(2) = 5e-20
Identities = 9/17 (52%), Positives = 10/17 (57%)
Frame = +1
Query: 387 EHVWQQKRISEACVVHK 403
EH W Q R+ E C HK
Sbjct: 175 EHAWLQTRV*EVCAAHK 225
>AV766904
Length = 611
Score = 82.8 bits (203), Expect = 1e-16
Identities = 46/125 (36%), Positives = 75/125 (59%)
Frame = +3
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
LA +L GYD GV+SGA +YI+ + + D + + I+++ S IG+ G +D
Sbjct: 108 LASMTSILLGYDIGVMSGAVIYIKRDLKISDVQIEILSGIMNIYSP---IGSYIAGRTSD 278
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR+ TI++A ++F GA++M +P ++ GR + G+G+G A + AP+Y SE SP
Sbjct: 279 WIGRRYTIVLAGLIFFVGAILMGFSPNYAFLMFGRFVAGVGIGFAFLIAPVYTSEISPTL 458
Query: 150 IRGAL 154
RG L
Sbjct: 459 SRGFL 473
>CB826772
Length = 490
Score = 77.0 bits (188), Expect = 5e-15
Identities = 38/118 (32%), Positives = 72/118 (60%)
Frame = +1
Query: 37 LFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKT 96
++GY+ GV++GA L+I+++ + D + I M A + A G ++D +GR+ T
Sbjct: 94 IYGYEVGVMTGALLFIKEDLQISDLQLQFLVGIFHMCGLPASMAA---GRISDYIGRRYT 264
Query: 97 ILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGAL 154
I++A + F++G+++ +P+ +++IG + G+G G + APLY +E SP RG+L
Sbjct: 265 IILASIAFLSGSILTGYSPSYLILMIGNFISGIGAGFTLIVAPLYCAEISPPSHRGSL 438
>TC19365 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (24%)
Length = 441
Score = 74.3 bits (181), Expect = 3e-14
Identities = 43/118 (36%), Positives = 73/118 (61%)
Frame = +2
Query: 37 LFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKT 96
+FGY TGV++GA L+I++E D + L I+++ + A + A G +D +GR+ T
Sbjct: 83 IFGYVTGVMAGALLFIKEELGINDMQVQLLGGILNVCALPACMVA---GRTSDYVGRRYT 253
Query: 97 ILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGAL 154
I+++ V+ + G+++M P+ +++IGR GLGVG A + AP+Y +E S RG L
Sbjct: 254 IILSAVILLLGSILMGYGPSFPILMIGRCTAGLGVGLALIIAPVYSTEISLPSNRGFL 427
>TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporter, partial
(37%)
Length = 903
Score = 68.6 bits (166), Expect = 2e-12
Identities = 35/118 (29%), Positives = 71/118 (59%)
Frame = +2
Query: 37 LFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKT 96
L+GY V++G L+I+++ + D + ++ M++ A + A G + D +GR+ T
Sbjct: 125 LYGYVMAVMTGVLLFIKEDLQLSDLQVQFSVGLLHMSALPACMTA---GRIADYIGRRYT 295
Query: 97 ILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGAL 154
I++ ++F+ G+++M P+ +++IG + G+G+G + + AP+Y +E SP RG L
Sbjct: 296 IILTAIIFLVGSVLMGYGPSYPILMIGLSISGIGLGFSLIVAPVYCAEISPPSHRGFL 469
>BI418774
Length = 554
Score = 64.7 bits (156), Expect = 3e-11
Identities = 45/91 (49%), Positives = 53/91 (57%), Gaps = 3/91 (3%)
Frame = +3
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLN---AVGT 297
VRR L A I V V QQI GI I YSP + + GI S S L L T GL AV T
Sbjct: 60 VRRILIAAIGVHVFQQICGIEGICIYSPRVFEKTGITSKSN---LLLATVGLGISQAVFT 230
Query: 298 IVSMVLIDRFGRRKLMLISLIGIFVSLVTLS 328
+S L+DR GRR L+LIS G+ V+L+ LS
Sbjct: 231 FISAFLLDRVGRRILLLISSGGVVVTLLGLS 323
>AV418961
Length = 394
Score = 61.6 bits (148), Expect = 2e-10
Identities = 33/84 (39%), Positives = 52/84 (61%)
Frame = +3
Query: 248 GITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRF 307
G + + QQ+ GIN ++YYS ++ + AGIAS+ A AL N GT V+ L+DR
Sbjct: 81 GAALFLFQQLAGINAVVYYSTSVFRSAGIASDVAASAL---VGASNVFGTAVASSLMDRQ 251
Query: 308 GRRKLMLISLIGIFVSLVTLSVTF 331
GR+ L++ S G+ S++ LS++F
Sbjct: 252 GRKSLLITSFSGMAASMLLLSLSF 323
>BP032429
Length = 468
Score = 53.5 bits (127), Expect = 6e-08
Identities = 28/102 (27%), Positives = 56/102 (54%), Gaps = 3/102 (2%)
Frame = -2
Query: 229 AQKLKGAWSN---DVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFAL 285
++K++ W N V + + + QQ+ GIN IM+Y+P + + G ++ A
Sbjct: 458 SKKVEHPWKNITKPVYKPQMVFCSLIPFFQQLTGINVIMFYAPVLFKTLGFGDDA-ALMS 282
Query: 286 SLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
++++ G+N + T VS+ +D+FGRR L L + + +S + +
Sbjct: 281 AVISGGVNVLATFVSIFSVDKFGRRLLFLEGGVQMLISQIAV 156
>TC20267 similar to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(31%)
Length = 525
Score = 53.1 bits (126), Expect = 8e-08
Identities = 34/116 (29%), Positives = 60/116 (51%), Gaps = 3/116 (2%)
Frame = +2
Query: 202 EVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRR---GLYAGITVQVVQQIV 258
+V+EE + E+ EA + +++ W N + R+ L I + QQ
Sbjct: 113 DVDEEFSDLVEASEA------------SMQVEHPWRNLLQRKYRPHLTMAILIPFFQQFT 256
Query: 259 GINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLML 314
GIN IM+Y+P + G +++ + +++T +N V T VS+ +D++GRRKL L
Sbjct: 257 GINVIMFYAPVLFSSIGFKDDASLMS-AVITGVVNVVATCVSIYGVDKWGRRKLFL 421
>BP058072
Length = 520
Score = 49.3 bits (116), Expect = 1e-06
Identities = 31/123 (25%), Positives = 59/123 (47%), Gaps = 17/123 (13%)
Frame = -1
Query: 30 LALSGGLLFGYDTGVISGA------------SLYIRDEFEQVDKKPWLQETIV-----SM 72
+A GGL+FGYD G+ G S+Y++ + + K ++ V S
Sbjct: 451 VAAMGGLIFGYDIGISGGVTSMDPFLRKFFHSVYLKKQEKSSPNKYCQYDSQVLTIFTSS 272
Query: 73 ASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
A++ + + + K ++ + + FV AL+ A W++I+GR+L+G+G+G
Sbjct: 271 LYLAALLSSLVASTITRRFSPKLSMFVGGMHFVVVALLNGLAQRVWMLIVGRILLGIGIG 92
Query: 133 AAS 135
A+
Sbjct: 91 FAN 83
>TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter (STP11
protein), partial (30%)
Length = 663
Score = 49.3 bits (116), Expect = 1e-06
Identities = 34/121 (28%), Positives = 57/121 (47%), Gaps = 18/121 (14%)
Frame = +3
Query: 30 LALSGGLLFGYDTGVISGASLY--------------IRDEFEQVDKKPWLQETIVSMASA 75
+A GGLLFGYD G+ G + ++DE + ++++ ++
Sbjct: 252 VAAMGGLLFGYDLGITGGVTSMEPFLVKFFPGVYKQMKDESGHESEYCKFDNELLTLFTS 431
Query: 76 G----AIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGV 131
A+I + F +GRK ++ + F+ GAL+ A ++IIGRLL+G GV
Sbjct: 432 SLYLAALIASFFASTTTRMLGRKTSMFAGGLFFLVGALLNGFAVNIEMLIIGRLLLGFGV 611
Query: 132 G 132
G
Sbjct: 612 G 614
>AW720443
Length = 406
Score = 48.5 bits (114), Expect = 2e-06
Identities = 27/82 (32%), Positives = 45/82 (53%), Gaps = 3/82 (3%)
Frame = +2
Query: 228 LAQKLKGAWSNDVVRRG---LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFA 284
+A+++K + N + R+ L I +Q+ QQ GIN IM+Y+P + G N A
Sbjct: 56 VAKQVKHPFRNILKRKNRPQLVIAIALQIFQQFTGINAIMFYAPVLFNTFGF-KNDAALY 232
Query: 285 LSLVTSGLNAVGTIVSMVLIDR 306
+++ +N V TIVS+ +DR
Sbjct: 233 SAVIVGAVNCVSTIVSIYSVDR 298
>AV418433
Length = 401
Score = 46.2 bits (108), Expect = 1e-05
Identities = 25/68 (36%), Positives = 42/68 (61%)
Frame = +3
Query: 105 VAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALVCRLHQWFTH 164
VAG LV+ + P ++ IGR+ G G+G S P++I+E +P ++RGAL L+Q+
Sbjct: 3 VAGWLVIYFSQGPVLLDIGRVATGYGMGVFSYVVPVFIAEIAPKELRGALT-TLNQFMLV 179
Query: 165 NRWSISLL 172
+ S+S +
Sbjct: 180 SAMSVSFV 203
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,045,882
Number of Sequences: 28460
Number of extensions: 98610
Number of successful extensions: 586
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 564
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 571
length of query: 420
length of database: 4,897,600
effective HSP length: 93
effective length of query: 327
effective length of database: 2,250,820
effective search space: 736018140
effective search space used: 736018140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148526.6


