

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148526.1 + phase: 0
(598 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
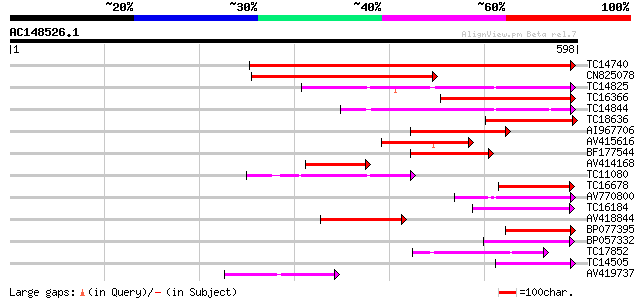
Score E
Sequences producing significant alignments: (bits) Value
TC14740 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial... 507 e-144
CN825078 219 8e-58
TC14825 similar to UP|Q8S4W7 (Q8S4W7) GAI-like protein 1, partia... 178 2e-45
TC16366 similar to PIR|E96542|E96542 scarecrow-like protein [imp... 174 4e-44
TC14844 similar to UP|Q8L553 (Q8L553) SCARECROW transcriptional ... 171 3e-43
TC18636 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial... 145 2e-35
AI967706 143 9e-35
AV415616 125 2e-29
BF177544 117 5e-27
AV414168 94 8e-20
TC11080 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin r... 87 1e-17
TC16678 similar to UP|Q8GVE2 (Q8GVE2) Chitin-inducible gibberell... 86 2e-17
AV770800 80 7e-16
TC16184 77 6e-15
AV418844 73 1e-13
BP077395 71 6e-13
BP057332 69 2e-12
TC17852 similar to UP|Q8W4I8 (Q8W4I8) SCARECROW gene regulator, ... 67 8e-12
TC14505 similar to UP|Q7Y1B6 (Q7Y1B6) GAI-like protein, partial ... 63 2e-10
AV419737 55 2e-08
>TC14740 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial (57%)
Length = 1406
Score = 507 bits (1305), Expect = e-144
Identities = 249/343 (72%), Positives = 283/343 (81%)
Frame = +1
Query: 254 KMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYF 313
KMVSVAG P QRLGAY+LEGLRAR+E SGS IYKALKCE+PT ELMS MHILYQICPY+
Sbjct: 4 KMVSVAGDPSQRLGAYLLEGLRARLELSGSLIYKALKCEQPTGKELMSYMHILYQICPYW 183
Query: 314 QFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQ 373
+FAYIS+NA I E M NESRIHIIDFQIAQG+QW LL+ AL H+PGGPPFIR+TG+DDSQ
Sbjct: 184 KFAYISANAAIAEAMANESRIHIIDFQIAQGTQWHLLIQALAHRPGGPPFIRITGVDDSQ 363
Query: 374 SFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFAL 433
SFHARGG L+IVG++L D A++ VPFEF+S M GCEV+ E+ V+ E L VNFP+ L
Sbjct: 364 SFHARGGGLEIVGERLSDFARSYGVPFEFHSAAMSGCEVERENLGVRPGEALAVNFPYVL 543
Query: 434 HHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFES 493
HH+PDESVS ENHRDRLLRLVK LSPKVV EQESNTNTSPF RF ETL YYTAMFES
Sbjct: 544 HHMPDESVSTENHRDRLLRLVKSLSPKVVTLAEQESNTNTSPFFNRFMETLEYYTAMFES 723
Query: 494 IDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLS 553
IDVA PRDDKKRI+AEQHCVARDIVN++ACEG ER ERHELFGKWK RFSMAGF LS
Sbjct: 724 IDVACPRDDKKRISAEQHCVARDIVNMVACEGAERVERHELFGKWKLRFSMAGFKQWPLS 903
Query: 554 PSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
SV+ + + LL+DFN +YR+EQ D A+ L W + + TSSAWR
Sbjct: 904 SSVMTATQNLLRDFNHNYRLEQRDGALCLGWMKRALVTSSAWR 1032
>CN825078
Length = 650
Score = 219 bits (559), Expect = 8e-58
Identities = 108/196 (55%), Positives = 139/196 (70%)
Frame = +3
Query: 256 VSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQF 315
VS+ G PIQRLGAYM+EGL AR E+SG++IY AL C EP EL+S M +L++ICPY +F
Sbjct: 63 VSINGEPIQRLGAYMVEGLVARKEASGNSIYHALNCREPEGEELLSYMQLLFEICPYLKF 242
Query: 316 AYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSF 375
Y+++N I E +NE RIHIIDFQIAQGSQWM LL AL +PGG P +R+TGIDD S
Sbjct: 243 GYMAANGAIAEACRNEDRIHIIDFQIAQGSQWMTLLQALAARPGGAPHVRITGIDDPVSK 422
Query: 376 HARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHH 435
+ARG LDIVGK+L ++ +P EF+ V ++G +V + +++ E L VNFP LHH
Sbjct: 423 YARGDGLDIVGKRLALMSEKFGIPVEFHGVPVFGPDVTRDMLDIRPGEALAVNFPLQLHH 602
Query: 436 IPDESVSMENHRDRLL 451
DESV + N RD LL
Sbjct: 603 TADESVDVSNPRDGLL 650
>TC14825 similar to UP|Q8S4W7 (Q8S4W7) GAI-like protein 1, partial (47%)
Length = 1392
Score = 178 bits (452), Expect = 2e-45
Identities = 96/294 (32%), Positives = 161/294 (54%), Gaps = 5/294 (1%)
Frame = +2
Query: 308 QICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVT 367
+ CPY +FA+ ++N I E Q + R+H+IDF I QG QW L+ AL +PGGPP R+T
Sbjct: 2 ETCPYLKFAHFTANQAILEAFQGKDRVHVIDFAINQGMQWPALMQALALRPGGPPAFRLT 181
Query: 368 GIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSV---KMYGCEVQLEDFEVQHDEV 424
GI ++ L VG KL A+T V FE+ + + + D +E
Sbjct: 182 GIGPPAPDNS--DHLQQVGWKLAQLAETIHVQFEYRGFVANSLADLDASMLDLRPPEEES 355
Query: 425 LVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETL 484
+ VN F LH + ++E ++ +++ + P+++ VEQE+ N + FL RF E+L
Sbjct: 356 VAVNSVFELHKLLARPGALE----KVFSVIRQVQPEILTVVEQEAGHNGASFLDRFTESL 523
Query: 485 NYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSM 544
+YY+A+F+S++ + + + + E + + + I N++ACEG +R ERHE +W+ RF+
Sbjct: 524 HYYSALFDSLEGSSAVESQDKAMTEIY-LGKQICNVVACEGSDRIERHETLTQWRTRFNS 700
Query: 545 AGFVPLLLSPSVIDSVRTLLKDF--NKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
AG P+ L + LL F YR+E+ + + L W ++ + +SAW+
Sbjct: 701 AGLSPVHLGSNAFKQASMLLALFAGGDGYRVEENNGCLTLGWHTRPLIATSAWQ 862
>TC16366 similar to PIR|E96542|E96542 scarecrow-like protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(27%)
Length = 711
Score = 174 bits (441), Expect = 4e-44
Identities = 80/142 (56%), Positives = 104/142 (72%)
Frame = +3
Query: 455 KILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVA 514
K LSPKVV VEQE NTN +PFL RF ET+NYY A+FESIDV LPR+ K+RIN EQHC+A
Sbjct: 6 KCLSPKVVTLVEQEFNTNNAPFLQRFVETMNYYQAVFESIDVVLPREHKERINVEQHCLA 185
Query: 515 RDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIE 574
R++VN++ACEG ER ERHEL KW+ RF+ AGF P L+ + S++ LL+ + Y +E
Sbjct: 186 REVVNLVACEGAERVERHELLNKWRMRFASAGFTPYPLNSYINSSIKDLLESYRGHYTLE 365
Query: 575 QTDVAINLAWKSKVMCTSSAWR 596
+ D A+ L W ++V+ S AWR
Sbjct: 366 ERDGALFLGWMNQVLVASCAWR 431
>TC14844 similar to UP|Q8L553 (Q8L553) SCARECROW transcriptional
regulator-like, partial (49%)
Length = 1051
Score = 171 bits (433), Expect = 3e-43
Identities = 100/248 (40%), Positives = 144/248 (57%), Gaps = 1/248 (0%)
Frame = +3
Query: 350 LLHALKHKPGGPPF-IRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMY 408
LLH L + G P ++V + + + +L VG L A+ ++ FEF V
Sbjct: 3 LLHELSARLAGKPVTVKVIAVAEDGADE----RLKTVGDMLGQQAERLRIGFEFKVVSCK 170
Query: 409 GCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQE 468
E+ E + DE L VNF F L +PDESVS EN RD LLR VK LSP+VV VEQE
Sbjct: 171 LAELTREALGCEPDETLAVNFAFKLFRMPDESVSTENPRDELLRRVKALSPRVVTLVEQE 350
Query: 469 SNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDER 528
NTNT+PF+PR E+ +YY+A+F+SI+ + R+ +R+ EQ ++R + N +ACEG +R
Sbjct: 351 LNTNTAPFVPRVTESCSYYSALFDSIESTVARESLERVKIEQG-LSRKVGNSVACEGRDR 527
Query: 529 FERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKV 588
ER E+FGKW+AR SMAGF + V +S+ L N+ +++ + I W +
Sbjct: 528 VERCEVFGKWRARMSMAGFRLRPVGQRVAESITAQLGAENR-VTVKEENGGICCGWVGRT 704
Query: 589 MCTSSAWR 596
+ +SAWR
Sbjct: 705 LTVASAWR 728
>TC18636 similar to UP|Q8W584 (Q8W584) AT4g17230/dl4650c, partial (10%)
Length = 813
Score = 145 bits (365), Expect = 2e-35
Identities = 65/97 (67%), Positives = 79/97 (81%)
Frame = +1
Query: 502 DKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVR 561
DK I+AEQHCVARDIVN+IACEG ER E HELFGKW++RFSM GFVP LS SV SVR
Sbjct: 1 DKNSISAEQHCVARDIVNMIACEGXERVEXHELFGKWRSRFSMXGFVPCPLSSSVTASVR 180
Query: 562 TLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWRCY 598
+L +FN++YR+E DVA+ L WK++ MCT+SAWRC+
Sbjct: 181 NILNEFNENYRLEHKDVALYLTWKNRAMCTASAWRCF 291
>AI967706
Length = 357
Score = 143 bits (360), Expect = 9e-35
Identities = 66/106 (62%), Positives = 84/106 (78%)
Frame = +2
Query: 423 EVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAE 482
E LVVNF F LHH+ DESVS N RD+LLR+VK L+PK+V VEQ+ NTNTSPFLPRF
Sbjct: 38 EALVVNFAFQLHHMRDESVSTVNERDQLLRMVKSLNPKLVTVVEQDMNTNTSPFLPRFII 217
Query: 483 TLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDER 528
YY+A+F+S+D LPR+ + R+N E+ C+ARDIVNI+ACEG++R
Sbjct: 218 AYEYYSAVFDSLDATLPRESQDRVNVERQCLARDIVNIVACEGEDR 355
>AV415616
Length = 316
Score = 125 bits (314), Expect = 2e-29
Identities = 61/103 (59%), Positives = 74/103 (71%), Gaps = 6/103 (5%)
Frame = +2
Query: 393 AKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMEN------H 446
AK+C VPFEF+++ EVQL+D E++ +E + VNF LHH+PDESV +N H
Sbjct: 8 AKSCNVPFEFHAIGTSPSEVQLQDLEIRPEEAIAVNFAMMLHHVPDESVDTQNQGWSQNH 187
Query: 447 RDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTA 489
R+RLLRL K LSPKVV VEQESNTN PFLPRF ET+NYY A
Sbjct: 188 RNRLLRLAKCLSPKVVTLVEQESNTNNLPFLPRFVETMNYYLA 316
>BF177544
Length = 318
Score = 117 bits (293), Expect = 5e-27
Identities = 55/88 (62%), Positives = 69/88 (77%)
Frame = +3
Query: 423 EVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAE 482
E LVVNF F LHH+PDESVS N RD+LLRLVK L+PK+V VEQ+ NTNT+PFL RF E
Sbjct: 54 EALVVNFAFQLHHMPDESVSTVNERDQLLRLVKSLNPKLVTVVEQDVNTNTAPFLQRFVE 233
Query: 483 TLNYYTAMFESIDVALPRDDKKRINAEQ 510
YY+A+FES+D+ LPR+ + R+N E+
Sbjct: 234 AYKYYSAVFESLDITLPRESQDRMNVER 317
>AV414168
Length = 208
Score = 93.6 bits (231), Expect = 8e-20
Identities = 39/68 (57%), Positives = 54/68 (79%)
Frame = +1
Query: 313 FQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDS 372
F+F Y+S+N I E M+ E +HIIDFQI+QG+QW+ L+ AL H+PGGPP +R+TG+DD+
Sbjct: 1 FKFGYMSANGAIAEAMKEEREVHIIDFQISQGTQWVSLIQALAHRPGGPPKVRITGVDDT 180
Query: 373 QSFHARGG 380
S +ARGG
Sbjct: 181 FSSYARGG 204
>TC11080 similar to UP|Q84TQ7 (Q84TQ7) GIA/RGA-like gibberellin response
modulator, partial (33%)
Length = 556
Score = 86.7 bits (213), Expect = 1e-17
Identities = 55/180 (30%), Positives = 87/180 (47%), Gaps = 1/180 (0%)
Frame = +2
Query: 250 KVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQI 309
K +G + + ++++ +Y + L R IY +S+ + MH Y+
Sbjct: 41 KHIGVVAASQAGAMRKVASYFAQALARR-------IYGIFPDTVDSSLHDVLHMHF-YES 196
Query: 310 CPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGI 369
CPY +FA+ ++N I E R+H++DF + QG QW L+ AL +PGGPP R+TGI
Sbjct: 197 CPYLKFAHFTANQAILEAFATAGRVHVVDFGLKQGMQWPALMQALALRPGGPPTFRLTGI 376
Query: 370 DDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYG-CEVQLEDFEVQHDEVLVVN 428
Q ++ L VG KL A+T V FEF ++ E++ E + VN
Sbjct: 377 GPPQPDNS--DALQQVGWKLAQLAQTIGVQFEFRGFVCNSLADLDPNMLEIRPGEAVAVN 550
>TC16678 similar to UP|Q8GVE2 (Q8GVE2) Chitin-inducible
gibberellin-responsive protein, partial (14%)
Length = 740
Score = 85.9 bits (211), Expect = 2e-17
Identities = 39/80 (48%), Positives = 57/80 (70%)
Frame = +3
Query: 516 DIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQ 575
D+VN+IACEG ER ERHELFGKWK+R +MAGF LS V ++ LL+ +++ Y + +
Sbjct: 3 DVVNVIACEGKERVERHELFGKWKSRLTMAGFRQSPLSSYVNSVIKNLLRCYSEHYTLIE 182
Query: 576 TDVAINLAWKSKVMCTSSAW 595
D A+ L WK++ + ++SAW
Sbjct: 183 KDGAMLLGWKNRSLISASAW 242
>AV770800
Length = 514
Score = 80.5 bits (197), Expect = 7e-16
Identities = 44/128 (34%), Positives = 66/128 (51%), Gaps = 1/128 (0%)
Frame = -1
Query: 470 NTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERF 529
N N L RF E L+YY+ +F+S++ DK AE + + R+I N++ CEG R
Sbjct: 514 NHNEDGLLERFTEALHYYSTVFDSLEGCPVEPDKAL--AEMY-LQREICNVVCCEGTARV 344
Query: 530 ERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKD-YRIEQTDVAINLAWKSKV 588
ERHE KW+ R AGF P+ L L F+ + Y +E+ + + L W S+
Sbjct: 343 ERHEPLAKWRERLGKAGFRPVHLGSDAFTQASMLFTLFSSEGYCVEEKEGCLTLGWHSRP 164
Query: 589 MCTSSAWR 596
+SAW+
Sbjct: 163 PLAASAWQ 140
>TC16184
Length = 702
Score = 77.4 bits (189), Expect = 6e-15
Identities = 39/108 (36%), Positives = 64/108 (59%), Gaps = 1/108 (0%)
Frame = +1
Query: 489 AMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFV 548
A F+ D + RD++ R E+ V R+ +N+IACEG ER ER E + +W+ R + AGF
Sbjct: 1 ATFDMFDXVISRDNQWRRLIERESVGREAMNVIACEGLERVERPETYKRWQVRNTRAGFK 180
Query: 549 PLLLSPSVIDSVRTLLKD-FNKDYRIEQTDVAINLAWKSKVMCTSSAW 595
L L+ ++D RT LK ++KD+ ++ + + WK ++M S+ W
Sbjct: 181 QLPLNEGLMDKFRTKLKKWYHKDFVFDEDNNWMLQGWKGRIMYASTCW 324
>AV418844
Length = 284
Score = 73.2 bits (178), Expect = 1e-13
Identities = 36/90 (40%), Positives = 55/90 (61%)
Frame = +1
Query: 329 QNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKK 388
+NES +HIIDF I G QW L+ +L + GGPP +RVTGI+ + +++ G++
Sbjct: 1 KNESSVHIIDFGICYGFQWPCLIKSLSERHGGPPRLRVTGIELPRPGFRPAERVEETGRR 180
Query: 389 LEDCAKTCKVPFEFNSVKMYGCEVQLEDFE 418
LE+ K KVPFE+N + ++LED +
Sbjct: 181 LENYCKKFKVPFEYNCLAKKWETIKLEDLK 270
>BP077395
Length = 397
Score = 70.9 bits (172), Expect = 6e-13
Identities = 31/73 (42%), Positives = 50/73 (68%)
Frame = -3
Query: 524 EGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLA 583
EG++R ER+E+ GKW+AR +MAGF P +S +V D +R L+K + Y+I++ A++
Sbjct: 395 EGEDRIERYEVAGKWRARMTMAGFTPSPMSANVGDEIRKLIKLYCDRYKIKEEMGALHFG 216
Query: 584 WKSKVMCTSSAWR 596
W+ K + +SAWR
Sbjct: 215 WEDKNLIVASAWR 177
>BP057332
Length = 494
Score = 69.3 bits (168), Expect = 2e-12
Identities = 33/97 (34%), Positives = 56/97 (57%), Gaps = 1/97 (1%)
Frame = -2
Query: 500 RDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDS 559
R+DK + E+ RD VN+IACEG ER ER E + +W+ R GF L L+P +
Sbjct: 493 REDKYXLMFEEGLFGRDAVNVIACEGAERVERPETYKQWQVRNRRPGFKQLPLAPELSSR 314
Query: 560 VRTLL-KDFNKDYRIEQTDVAINLAWKSKVMCTSSAW 595
V+ ++ K+++KD+ +++ + WK +++ S W
Sbjct: 313 VKEMVKKEYHKDFVVDEDGKWVLQGWKGRILHAVSCW 203
>TC17852 similar to UP|Q8W4I8 (Q8W4I8) SCARECROW gene regulator, partial
(26%)
Length = 456
Score = 67.0 bits (162), Expect = 8e-12
Identities = 46/146 (31%), Positives = 73/146 (49%), Gaps = 2/146 (1%)
Frame = +2
Query: 425 LVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETL 484
L VNF L+++ DE+ + + LRL K L P++V E E++ F+ RF L
Sbjct: 2 LAVNFMLQLYNLLDETPTAV---ETALRLAKSLKPRIVTLGEYEASLTRVGFVSRFKAAL 172
Query: 485 NYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACE--GDERFERHELFGKWKARF 542
+++A+FES++ LP D +R E + R I +I E G R ER E +W+
Sbjct: 173 KHFSALFESLEPNLPSDSPERFQVESLLLGRRIAGVIGPELPGCVR-ERMEDKEQWRVLM 349
Query: 543 SMAGFVPLLLSPSVIDSVRTLLKDFN 568
GF + LS I + LL +++
Sbjct: 350 ESCGFESVSLSHYAISQAKILLWNYS 427
>TC14505 similar to UP|Q7Y1B6 (Q7Y1B6) GAI-like protein, partial (16%)
Length = 670
Score = 62.8 bits (151), Expect = 2e-10
Identities = 29/86 (33%), Positives = 46/86 (52%), Gaps = 2/86 (2%)
Frame = +1
Query: 513 VARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDF--NKD 570
+ I N++ACEG +R ERHE +W+ R AGF P+ L + LL F
Sbjct: 64 LGEQICNVVACEGVDRVERHETLVQWRTRMGSAGFNPVHLGSNAFKQASMLLALFAGGDG 243
Query: 571 YRIEQTDVAINLAWKSKVMCTSSAWR 596
YR+E+ + + L W ++ + +SAW+
Sbjct: 244 YRVEENNGCLMLGWHTRPLIATSAWK 321
>AV419737
Length = 410
Score = 55.5 bits (132), Expect = 2e-08
Identities = 37/121 (30%), Positives = 58/121 (47%)
Frame = +3
Query: 227 EELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIY 286
++L + A+ + G+ A G + + L +S G P R YM E L++ + S+G +
Sbjct: 60 DQLYKTAELIEAGNPVHAQGILAR-LNHQLSPNGKPFHRAAFYMKEALQSMLHSNG---H 227
Query: 287 KALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQ 346
L + I + A +I P QFA + N + E ++ RIHIIDF I G Q
Sbjct: 228 NFLTFSPISFIFKIGAYKSFSEISPVLQFANFTCNQALIEAVERFDRIHIIDFDIGFGVQ 407
Query: 347 W 347
W
Sbjct: 408 W 410
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,023,548
Number of Sequences: 28460
Number of extensions: 131049
Number of successful extensions: 718
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 704
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 707
length of query: 598
length of database: 4,897,600
effective HSP length: 95
effective length of query: 503
effective length of database: 2,193,900
effective search space: 1103531700
effective search space used: 1103531700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148526.1


