

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148525.11 + phase: 0 /pseudo
(88 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
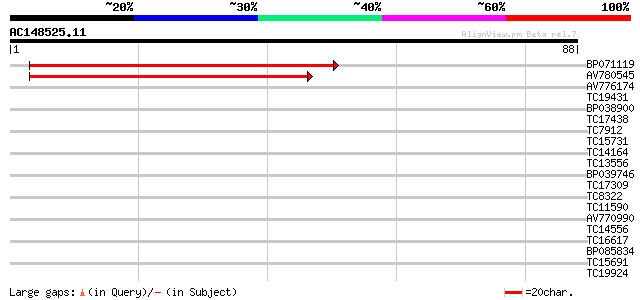
Score E
Sequences producing significant alignments: (bits) Value
BP071119 55 2e-09
AV780545 41 4e-05
AV776174 27 0.48
TC19431 weakly similar to UP|AAR20721 (AAR20721) At5g43820, part... 26 1.4
BP038900 26 1.4
TC17438 weakly similar to GB|AAP21348.1|30102860|BT006540 At3g23... 25 1.8
TC7912 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic p... 25 1.8
TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Pr... 25 3.1
TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, ... 25 3.1
TC13556 similar to UP|Q9M729 (Q9M729) Actin, partial (37%) 25 3.1
BP039746 24 4.1
TC17309 24 4.1
TC8322 weakly similar to UP|ATP7_ARATH (Q9SJ12) Probable ATP syn... 24 4.1
TC11590 24 4.1
AV770990 24 5.4
TC14556 homologue to UP|Q9ZTW5 (Q9ZTW5) GDP-mannose pyrophosphor... 24 5.4
TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protei... 24 5.4
BP085834 23 7.0
TC15691 similar to UP|Q944G3 (Q944G3) Acetyl Co-A acetyltransfer... 23 7.0
TC19924 23 7.0
>BP071119
Length = 412
Score = 55.1 bits (131), Expect = 2e-09
Identities = 23/48 (47%), Positives = 33/48 (67%)
Frame = +2
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLS 51
+GL+W+ + +DQ ++S ERWH TSSFHLP EM +TL+D+S
Sbjct: 266 AGLRWVMKCTSPSVDQSIISAFVERWHPETSSFHLPWGEMTITLEDVS 409
>AV780545
Length = 555
Score = 40.8 bits (94), Expect = 4e-05
Identities = 17/44 (38%), Positives = 28/44 (63%)
Frame = +1
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTL 47
+G +L+ + + +D L+S + ERW T++FHL + EM VTL
Sbjct: 424 AGFGYLRSIPAISLDNPLISALVERWRRETNTFHLNVGEMTVTL 555
>AV776174
Length = 330
Score = 27.3 bits (59), Expect = 0.48
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +2
Query: 44 IVTLDDLSCLLHLSIESHMLYHN 66
IVT++ LSCLLHL H+ YH+
Sbjct: 53 IVTVNALSCLLHL---PHLKYHS 112
>TC19431 weakly similar to UP|AAR20721 (AAR20721) At5g43820, partial (39%)
Length = 603
Score = 25.8 bits (55), Expect = 1.4
Identities = 8/43 (18%), Positives = 23/43 (52%)
Frame = -2
Query: 27 ERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYL 69
+ WH V + + + +++ ++ SC+ H+ +H H +++
Sbjct: 227 DHWHWVLRPWLIKLRPLMILINTQSCVFHVETTAHPSNH*VWV 99
>BP038900
Length = 471
Score = 25.8 bits (55), Expect = 1.4
Identities = 15/49 (30%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Frame = -2
Query: 23 SVIAERWHEVTSSFHLP--IWEMIVTLDDLSCLLHLSIESHMLYHNIYL 69
S++ RW E LP + + +D + CL+H+S S + H+I L
Sbjct: 164 SLLKSRWKE*VRVEDLPC*LSSLNSYMDQIYCLIHVSTTSITVPHHIIL 18
>TC17438 weakly similar to GB|AAP21348.1|30102860|BT006540 At3g23220
{Arabidopsis thaliana;}, partial (41%)
Length = 575
Score = 25.4 bits (54), Expect = 1.8
Identities = 12/44 (27%), Positives = 23/44 (52%), Gaps = 3/44 (6%)
Frame = +3
Query: 28 RWHE---VTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIY 68
RWH ++ S+ LP W + + D SC L ++++ + + Y
Sbjct: 114 RWHHSFVLS*SWTLPFWRGMPEVSDFSCTSLLELDTNFNFISNY 245
>TC7912 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
partial (95%)
Length = 1045
Score = 25.4 bits (54), Expect = 1.8
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 5/60 (8%)
Frame = +3
Query: 6 LQWL---QHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTL--DDLSCLLHLSIES 60
L WL + S LV GL S +A WH S+ W++ T+ +L L LS+ S
Sbjct: 291 LDWLLVARSPSSLVSSTGLHSFLAP*WHVFYSTMSQEAWKLPSTVWPLELELLKELSLRS 470
>TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Probable
serine/threonine-protein kinase PK5) , partial (17%)
Length = 940
Score = 24.6 bits (52), Expect = 3.1
Identities = 13/44 (29%), Positives = 22/44 (49%), Gaps = 4/44 (9%)
Frame = -2
Query: 30 HEVTSSFH----LPIWEMIVTLDDLSCLLHLSIESHMLYHNIYL 69
H+ +S FH P D+ +LS ++H+L+H IY+
Sbjct: 357 HQESSKFH*LLKNPNNSFCTPEFDMKSSFNLSEDTHILFHRIYI 226
>TC14164 weakly similar to UP|Q8VYH6 (Q8VYH6) AT4g17280/dl4675c, partial (44%)
Length = 1606
Score = 24.6 bits (52), Expect = 3.1
Identities = 11/37 (29%), Positives = 19/37 (50%)
Frame = -1
Query: 30 HEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHN 66
H + + HL +W +T+D + + HL + L HN
Sbjct: 1339 HSLLITLHL-LWPRTITIDPICSMCHLVL*FPPLQHN 1232
>TC13556 similar to UP|Q9M729 (Q9M729) Actin, partial (37%)
Length = 497
Score = 24.6 bits (52), Expect = 3.1
Identities = 12/39 (30%), Positives = 22/39 (55%), Gaps = 1/39 (2%)
Frame = -2
Query: 8 WLQHVSYLVIDQGLLSVIAERWHE-VTSSFHLPIWEMIV 45
W+ V Y +QG L + +T+S ++PIW++I+
Sbjct: 379 WIISVRYFHTNQGWLE*NLWTFEPLITNSDYMPIWQLII 263
>BP039746
Length = 377
Score = 24.3 bits (51), Expect = 4.1
Identities = 11/28 (39%), Positives = 16/28 (56%), Gaps = 1/28 (3%)
Frame = +3
Query: 30 HEVTSSFH-LPIWEMIVTLDDLSCLLHL 56
HE+ S F IW I+ + + CLLH+
Sbjct: 156 HEIASIFFAFKIWICIIMVRVICCLLHI 239
>TC17309
Length = 523
Score = 24.3 bits (51), Expect = 4.1
Identities = 7/16 (43%), Positives = 12/16 (74%)
Frame = +3
Query: 40 IWEMIVTLDDLSCLLH 55
IW +++ DLSC++H
Sbjct: 423 IWSCRISIPDLSCMIH 470
>TC8322 weakly similar to UP|ATP7_ARATH (Q9SJ12) Probable ATP synthase 24
kDa subunit, mitochondrial precursor , partial (94%)
Length = 1089
Score = 24.3 bits (51), Expect = 4.1
Identities = 9/19 (47%), Positives = 11/19 (57%)
Frame = +2
Query: 26 AERWHEVTSSFHLPIWEMI 44
A+ WH V SF P W M+
Sbjct: 65 AQPWHSVLVSFPNPNWPMV 121
>TC11590
Length = 412
Score = 24.3 bits (51), Expect = 4.1
Identities = 13/46 (28%), Positives = 19/46 (41%)
Frame = +1
Query: 7 QWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSC 52
QW Q ++ Q L+ H++ S IW +V D SC
Sbjct: 118 QWQQTQVMMIQSQPLIIPFWSTEHQINGSGPSMIWARLVRNGDSSC 255
>AV770990
Length = 180
Score = 23.9 bits (50), Expect = 5.4
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = -2
Query: 40 IWEMIVTLDDLSCLLHL 56
I E ++TLD + CL+HL
Sbjct: 131 ILECVITLDFIFCLVHL 81
>TC14556 homologue to UP|Q9ZTW5 (Q9ZTW5) GDP-mannose pyrophosphorylase,
complete
Length = 1521
Score = 23.9 bits (50), Expect = 5.4
Identities = 11/35 (31%), Positives = 20/35 (56%)
Frame = +1
Query: 38 LPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRS 72
L +WE++ L L +++ESH L+ + L+ S
Sbjct: 400 LNLWELLAPLL*LGIS**MALESHFLFSTVMLSAS 504
>TC16617 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (42%)
Length = 551
Score = 23.9 bits (50), Expect = 5.4
Identities = 12/42 (28%), Positives = 20/42 (47%), Gaps = 5/42 (11%)
Frame = +3
Query: 6 LQWLQHVSYLVIDQGLLSVIA-----ERWHEVTSSFHLPIWE 42
+QW + + + V Q +A E WH + + LP+WE
Sbjct: 15 IQWRRFLWHSVSTQSHFRTMADTTGLEEWHLLVWAKKLPLWE 140
>BP085834
Length = 457
Score = 23.5 bits (49), Expect = 7.0
Identities = 11/24 (45%), Positives = 14/24 (57%), Gaps = 4/24 (16%)
Frame = -3
Query: 36 FHLP----IWEMIVTLDDLSCLLH 55
F LP IW + + DLSCL+H
Sbjct: 134 FFLPL**EIWSWRIPILDLSCLIH 63
>TC15691 similar to UP|Q944G3 (Q944G3) Acetyl Co-A acetyltransferase,
partial (15%)
Length = 459
Score = 23.5 bits (49), Expect = 7.0
Identities = 7/12 (58%), Positives = 10/12 (83%)
Frame = +1
Query: 5 GLQWLQHVSYLV 16
GLQW H+SY++
Sbjct: 55 GLQWSSHLSYII 90
>TC19924
Length = 525
Score = 23.5 bits (49), Expect = 7.0
Identities = 14/44 (31%), Positives = 21/44 (46%), Gaps = 3/44 (6%)
Frame = +3
Query: 40 IWEMIVTLDDLSCLLHLSIESH---MLYHNIYLTRSEAVDLMVE 80
IWE+ D+SCL +L ++ H I +SEA + E
Sbjct: 48 IWELAFKHSDVSCLPYLP*KAAK**AFLHKILFPKSEAFNFTEE 179
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,865,696
Number of Sequences: 28460
Number of extensions: 26030
Number of successful extensions: 166
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 166
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 166
length of query: 88
length of database: 4,897,600
effective HSP length: 64
effective length of query: 24
effective length of database: 3,076,160
effective search space: 73827840
effective search space used: 73827840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 48 (23.1 bits)
Medicago: description of AC148525.11


