

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
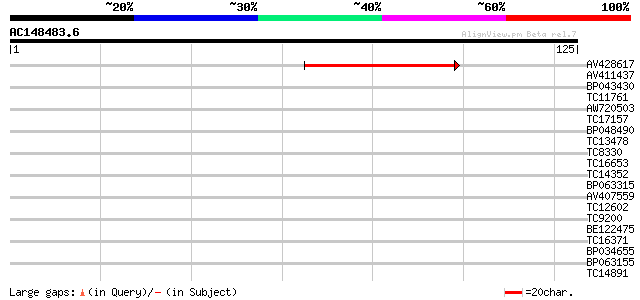
Query= AC148483.6 + phase: 0
(125 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV428617 51 7e-08
AV411437 28 0.35
BP043430 28 0.45
TC11761 28 0.59
AW720503 27 1.0
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 27 1.3
BP048490 27 1.3
TC13478 similar to UP|Q8MRP3 (Q8MRP3) GH08189p, partial (11%) 26 1.7
TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodu... 26 1.7
TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfat... 26 2.3
TC14352 similar to UP|ADF2_PETHY (Q9FVI1) Actin-depolymerizing f... 26 2.3
BP063315 26 2.3
AV407559 26 2.3
TC12602 similar to GB|AAO63373.1|28950899|BT005309 At2g27770 {Ar... 25 2.9
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 25 2.9
BE122475 25 2.9
TC16371 weakly similar to UP|P92169 (P92169) Serine repeat antig... 25 3.8
BP034655 25 3.8
BP063155 25 3.8
TC14891 25 3.8
>AV428617
Length = 355
Score = 50.8 bits (120), Expect = 7e-08
Identities = 23/34 (67%), Positives = 29/34 (84%)
Frame = +3
Query: 66 KLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNII 99
KLP VKAGDPV+HEPA+EV+ SEIKS+++ II
Sbjct: 252 KLPDTVKAGDPVLHEPAQEVNPSEIKSERVHKII 353
>AV411437
Length = 417
Score = 28.5 bits (62), Expect = 0.35
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 2/57 (3%)
Frame = -1
Query: 9 WRLRAFPMPVLSNGVVS--LSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKT 63
W + P P +SN S +SSSS+C+N S+ S S+ SS A L +T
Sbjct: 192 WASSSSPPPSVSNSSSSESVSSSSNCSNSPSTPSSSSSSSSSSSSSAIVSCAELPET 22
>BP043430
Length = 493
Score = 28.1 bits (61), Expect = 0.45
Identities = 19/37 (51%), Positives = 24/37 (64%)
Frame = +3
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETAL 59
V SLSSSSS N+ + L S+ S S+ SS SS T+L
Sbjct: 228 VQSLSSSSSLNSSLSLPSS--SSLLSSSSSSSSSTSL 332
Score = 25.8 bits (55), Expect = 2.3
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +3
Query: 20 SNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALL 60
S+ ++S SSSSS + + S+ S +LSSP + T +L
Sbjct: 282 SSSLLSSSSSSSSSTSLSFPSSSLSLPLPSLSSPLALTGIL 404
>TC11761
Length = 711
Score = 27.7 bits (60), Expect = 0.59
Identities = 18/63 (28%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Frame = +1
Query: 11 LRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSK-FGSTLSSPSSETALLRKTVNKLPY 69
+++ P+P LSN ++SS+ ++ + SS + ++ FG L S +LRK ++++
Sbjct: 97 IKSSPIPFLSNKPAAVSSNGRKSSYVMASSRREAENFGGRLVDES--MIILRKRIHEMSM 270
Query: 70 IVK 72
I K
Sbjct: 271 IEK 279
>AW720503
Length = 539
Score = 26.9 bits (58), Expect = 1.0
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 5/65 (7%)
Frame = -3
Query: 7 LVWRLRAFPMPVLSNGVVSL---SSSSSCNNKIQLSSTKFSKFGSTLSS--PSSETALLR 61
++ L AF + LS +SL SSS S + LSS+ SKF ++SS S T+ LR
Sbjct: 420 IICSLEAFTLATLSCNSLSLIMYSSSFSTKKLLNLSSSFSSKFLKSVSSLDISPATSSLR 241
Query: 62 KTVNK 66
+ +
Sbjct: 240 RLAGR 226
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 26.6 bits (57), Expect = 1.3
Identities = 17/43 (39%), Positives = 25/43 (57%)
Frame = -3
Query: 16 MPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
+P+L S SSSSS ++ SS+ S S+ SSPSS ++
Sbjct: 903 LPLLLLLFPSPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSS 775
Score = 25.4 bits (54), Expect = 2.9
Identities = 21/55 (38%), Positives = 28/55 (50%)
Frame = -3
Query: 11 LRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVN 65
L FP P S+ S SSSSS ++ SS+ S S+ SS SS++ VN
Sbjct: 891 LLLFPSPSSSSSSSSSSSSSSSSS--SSSSSSSSSPSSSSSSISSKSLCSASPVN 733
>BP048490
Length = 452
Score = 26.6 bits (57), Expect = 1.3
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 5/57 (8%)
Frame = -2
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKF--GSTLSSPSSETALLRK---TVNKLPYIVKAG 74
+V SSSSS ++++ LSST F +LS SS + L T L +V AG
Sbjct: 241 LVDFSSSSSLSSELSLSSTSILSFLTSESLSDESSSSESLLADFLTAEGLATVVVAG 71
>TC13478 similar to UP|Q8MRP3 (Q8MRP3) GH08189p, partial (11%)
Length = 531
Score = 26.2 bits (56), Expect = 1.7
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 1/56 (1%)
Frame = +3
Query: 27 SSSSSCNNKIQLSSTKFSKFGST-LSSPSSETALLRKTVNKLPYIVKAGDPVIHEP 81
SSS++ NK +LS + + FGST L++ ++ A +++LP +HEP
Sbjct: 156 SSSNAS*NKQRLSQDQKTTFGSTSLTATATAAATTHHNLHRLPKGNPHHTQ*LHEP 323
>TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodulin-like
protein {Arabidopsis thaliana;} , partial (72%)
Length = 1174
Score = 26.2 bits (56), Expect = 1.7
Identities = 17/48 (35%), Positives = 24/48 (49%)
Frame = -1
Query: 15 PMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRK 62
P P S + S SSS SCN ++S+ S +T PS ++ RK
Sbjct: 439 PSPSTSRSITSTSSSLSCNAPRRVSTATRSARDTT---PSLSRSMRRK 305
>TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfate
reductase, partial (69%)
Length = 1242
Score = 25.8 bits (55), Expect = 2.3
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +3
Query: 24 VSLSSSSSCNNKIQLSSTKFSKFGS 48
+S+SSSSS ++ Q S KFS+ S
Sbjct: 72 ISISSSSSSSSTFQSSQPKFSQIAS 146
>TC14352 similar to UP|ADF2_PETHY (Q9FVI1) Actin-depolymerizing factor 2
(ADF 2), partial (96%)
Length = 1311
Score = 25.8 bits (55), Expect = 2.3
Identities = 23/87 (26%), Positives = 33/87 (37%), Gaps = 12/87 (13%)
Frame = +3
Query: 12 RAFPMPVLSNGVVSLSSSSSC------------NNKIQLSSTKFSKFGSTLSSPSSETAL 59
++ P+P NGV+S+S S N K + T F T PS E
Sbjct: 72 KSLPLPGNKNGVISISLISEVI*EDPKCLD*TQNQKKRFLRTIFIDISPTYILPSQECGS 251
Query: 60 LRKTVNKLPYIVKAGDPVIHEPAREVD 86
+K L V G P ++P +D
Sbjct: 252 NQKRGKILCPPVHPG*PCYYQPGYYMD 332
>BP063315
Length = 440
Score = 25.8 bits (55), Expect = 2.3
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = -2
Query: 33 NNKIQLSSTKFSKFGSTLSSPSSETAL 59
N++ + SST S+F S+LSS SS +L
Sbjct: 112 NSRTRNSSTSVSRFFSSLSSSSSSLSL 32
>AV407559
Length = 429
Score = 25.8 bits (55), Expect = 2.3
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = +1
Query: 26 LSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIH 79
++SSSS N+ S TLSSPS LL ++ P+ P++H
Sbjct: 118 MTSSSSTNHSHSAHHHALSLNPQTLSSPS----LLHSSIQTTPFRKLTHQPLLH 267
>TC12602 similar to GB|AAO63373.1|28950899|BT005309 At2g27770 {Arabidopsis
thaliana;}, partial (14%)
Length = 728
Score = 25.4 bits (54), Expect = 2.9
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = -1
Query: 5 RRLVWRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTK 42
R L+ + FP +L + V S+SSS S + STK
Sbjct: 137 RSLLETIEKFPFMILRSSVSSISSSLSPRRRPSSESTK 24
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 25.4 bits (54), Expect = 2.9
Identities = 21/56 (37%), Positives = 32/56 (56%)
Frame = -3
Query: 3 AARRLVWRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
+ARRL+ + +FPM S + S SSSS+ S++ S S+ SSPSS ++
Sbjct: 398 SARRLM--ICSFPMGSRSFWIGSSSSSSA-----SASTSSSSSSSSSSSSPSSSSS 252
>BE122475
Length = 378
Score = 25.4 bits (54), Expect = 2.9
Identities = 11/34 (32%), Positives = 20/34 (58%)
Frame = -1
Query: 15 PMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGS 48
P P LS +S+S S + + +SST+ ++ G+
Sbjct: 264 PSPTLSRFTISISLSVTTRKGVSISSTRQAEVGT 163
>TC16371 weakly similar to UP|P92169 (P92169) Serine repeat antigen
(Fragment), partial (13%)
Length = 500
Score = 25.0 bits (53), Expect = 3.8
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = -2
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSS 55
+ S SSS+S N+ SST++S S S+ SS
Sbjct: 163 IPSESSSASSNSSSSFSSTRWSLSSSEASTSSS 65
>BP034655
Length = 517
Score = 25.0 bits (53), Expect = 3.8
Identities = 16/38 (42%), Positives = 22/38 (57%)
Frame = -3
Query: 20 SNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSET 57
S+ S SSSSS ++ SS+ S S+ SS SSE+
Sbjct: 242 SSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSES 129
>BP063155
Length = 462
Score = 25.0 bits (53), Expect = 3.8
Identities = 10/19 (52%), Positives = 15/19 (78%)
Frame = -3
Query: 81 PAREVDHSEIKSDKIQNII 99
PA+ +H +IKS +IQN+I
Sbjct: 370 PAKVREHCQIKSTEIQNLI 314
>TC14891
Length = 1172
Score = 25.0 bits (53), Expect = 3.8
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = -2
Query: 37 QLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDP 76
Q + T S GSTL+S S+T KTV YI K P
Sbjct: 562 QATLTSSSALGSTLTSSISDTLYTSKTV----YIGKVVSP 455
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,129,988
Number of Sequences: 28460
Number of extensions: 27520
Number of successful extensions: 242
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 233
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 242
length of query: 125
length of database: 4,897,600
effective HSP length: 80
effective length of query: 45
effective length of database: 2,620,800
effective search space: 117936000
effective search space used: 117936000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 50 (23.9 bits)
Medicago: description of AC148483.6


