

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.13 + phase: 0
(63 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
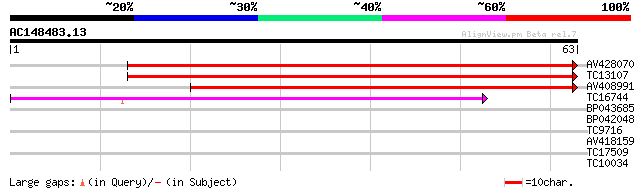
Score E
Sequences producing significant alignments: (bits) Value
AV428070 77 5e-16
TC13107 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13... 77 9e-16
AV408991 70 1e-13
TC16744 similar to GB|AAP37738.1|30725432|BT008379 At4g13350 {Ar... 42 3e-05
BP043685 36 0.002
BP042048 26 1.3
TC9716 25 3.0
AV418159 25 3.9
TC17509 23 8.6
TC10034 similar to UP|Q84PB7 (Q84PB7) Inositol phosphatase-like ... 23 8.6
>AV428070
Length = 356
Score = 77.4 bits (189), Expect = 5e-16
Identities = 31/50 (62%), Positives = 40/50 (80%)
Frame = +1
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVR
Sbjct: 166 RDLQSEPSNKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVR 315
>TC13107 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13M22.5
{Arabidopsis thaliana;}, partial (29%)
Length = 465
Score = 76.6 bits (187), Expect = 9e-16
Identities = 32/50 (64%), Positives = 39/50 (78%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R+L++ NK C DC KNP WASV+YGIF+C++CS HR LGVHISFVR
Sbjct: 62 RELQSMPGNKICVDCTQKNPQWASVSYGIFMCLECSGKHRGLGVHISFVR 211
>AV408991
Length = 435
Score = 69.7 bits (169), Expect = 1e-13
Identities = 29/43 (67%), Positives = 33/43 (76%)
Frame = +1
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
EN+ C DC AK P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 175 ENRECADCKAKGPRWASVNLGIFICMQCSGIHRSLGVHISKVR 303
>TC16744 similar to GB|AAP37738.1|30725432|BT008379 At4g13350 {Arabidopsis
thaliana;}, partial (18%)
Length = 687
Score = 41.6 bits (96), Expect = 3e-05
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Frame = +2
Query: 1 MASDSFTDKNA-VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
MAS +KN V R L N+ C +CN+ P + + F+C +CS +HR
Sbjct: 350 MASRKEDEKNERVIRGLLKLESNRRCINCNSLGPQYVCTNFWTFVCTNCSGIHR 511
>BP043685
Length = 433
Score = 35.8 bits (81), Expect = 0.002
Identities = 16/54 (29%), Positives = 25/54 (45%), Gaps = 1/54 (1%)
Frame = +1
Query: 1 MASDSFTDKNA-VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
M S +KN + R L N+ C +C+ P + + F+C CS +HR
Sbjct: 259 MGSRKEEEKNEKIIRGLMKLPPNRRCINCDNLGPQYVCTNFWTFICTTCSGIHR 420
>BP042048
Length = 520
Score = 26.2 bits (56), Expect = 1.3
Identities = 11/39 (28%), Positives = 19/39 (48%)
Frame = +3
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
N SC + KN W++ ++C A+ RS +I+
Sbjct: 18 NNSCIELQMKNARWSAYYLNYKCMLNCRALIRSAKFYIN 134
>TC9716
Length = 601
Score = 25.0 bits (53), Expect = 3.0
Identities = 8/19 (42%), Positives = 13/19 (68%)
Frame = +3
Query: 44 LCIDCSAVHRSLGVHISFV 62
+C CS +HR VH+S++
Sbjct: 405 ICFMCSEIHRI*PVHVSYI 461
>AV418159
Length = 282
Score = 24.6 bits (52), Expect = 3.9
Identities = 10/25 (40%), Positives = 16/25 (64%), Gaps = 2/25 (8%)
Frame = -2
Query: 35 WASVTYGIFLCIDCSA--VHRSLGV 57
W S GI+ CI+ +A + RS+G+
Sbjct: 140 WTSTITGIYCCINLNAQQIRRSMGI 66
>TC17509
Length = 497
Score = 23.5 bits (49), Expect = 8.6
Identities = 8/20 (40%), Positives = 14/20 (70%)
Frame = +3
Query: 17 KTKSENKSCFDCNAKNPTWA 36
K+KSE+ +CF C + +W+
Sbjct: 69 KSKSESSTCFCCRSGP*SWS 128
>TC10034 similar to UP|Q84PB7 (Q84PB7) Inositol phosphatase-like protein,
partial (18%)
Length = 588
Score = 23.5 bits (49), Expect = 8.6
Identities = 7/20 (35%), Positives = 12/20 (60%)
Frame = +1
Query: 43 FLCIDCSAVHRSLGVHISFV 62
F+C+ C+ HR I+F+
Sbjct: 481 FMCVHCTDTHRDSPFEINFI 540
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.132 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,318,012
Number of Sequences: 28460
Number of extensions: 16374
Number of successful extensions: 133
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 133
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 133
length of query: 63
length of database: 4,897,600
effective HSP length: 39
effective length of query: 24
effective length of database: 3,787,660
effective search space: 90903840
effective search space used: 90903840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC148483.13


