

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
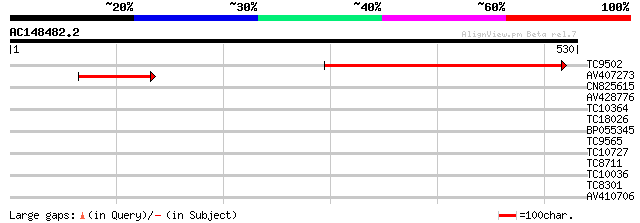
Sequences producing significant alignments: (bits) Value
TC9502 similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate cotran... 387 e-108
AV407273 144 4e-35
CN825615 30 0.74
AV428776 30 0.97
TC10364 similar to UP|Q9ARE3 (Q9ARE3) ZF-HD homeobox protein, pa... 29 2.2
TC18026 similar to UP|AAR87415 (AAR87415) Surface antigen (Fragm... 29 2.2
BP055345 28 4.8
TC9565 similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (19%) 28 4.8
TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein... 27 6.3
TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor,... 27 8.2
TC10036 weakly similar to UP|Q9LFB5 (Q9LFB5) Anthranilate N-benz... 27 8.2
TC8301 27 8.2
AV410706 27 8.2
>TC9502 similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate
cotransporter-like, partial (42%)
Length = 1253
Score = 387 bits (993), Expect = e-108
Identities = 190/226 (84%), Positives = 210/226 (92%)
Frame = +2
Query: 295 LEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAVLLFI 354
LEALGPMSFAEKMVL VFGLLIVLWMTRRI+DD+PGWG FHGLVGDGS+SV+AAVLLFI
Sbjct: 5 LEALGPMSFAEKMVLSVFGLLIVLWMTRRITDDIPGWGSFFHGLVGDGSVSVMAAVLLFI 184
Query: 355 IPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYL 414
IPNMK+ GEKLMDWNECKKLPW+LILLLGAGFA+ADGVQSSGLADV+S+AL+FL D PYL
Sbjct: 185 IPNMKKEGEKLMDWNECKKLPWSLILLLGAGFAIADGVQSSGLADVLSRALDFLDDAPYL 364
Query: 415 AIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLP 474
AI PAVSL+ SIITEFITSNDATATL++PLLYHIA M VHPLLLMIPG IATEFAF LP
Sbjct: 365 AIAPAVSLMSSIITEFITSNDATATLIIPLLYHIARTMRVHPLLLMIPGAIATEFAFLLP 544
Query: 475 TSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLL 520
TSTPSNVVGFATGHIEI+D LKVGVPLKVAG+++L+ LMPTLG ++
Sbjct: 545 TSTPSNVVGFATGHIEIQDTLKVGVPLKVAGIALLSFLMPTLGAVV 682
>AV407273
Length = 224
Score = 144 bits (363), Expect = 4e-35
Identities = 68/72 (94%), Positives = 71/72 (98%)
Frame = +1
Query: 65 MLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILA 124
ML VIAWVFTWW+T+AVPLPVTSMCPLFLFPLFGI+SADTVAHSYMDDVITLVLGSFILA
Sbjct: 7 MLAVIAWVFTWWVTSAVPLPVTSMCPLFLFPLFGISSADTVAHSYMDDVITLVLGSFILA 186
Query: 125 LAVERYNVHRRL 136
LAVERYNVHRRL
Sbjct: 187 LAVERYNVHRRL 222
>CN825615
Length = 572
Score = 30.4 bits (67), Expect = 0.74
Identities = 20/63 (31%), Positives = 34/63 (53%)
Frame = +2
Query: 428 TEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATG 487
T+ ITSND T L L+ + ++++ + +L M GG+ ++ A P S +V F +G
Sbjct: 350 TQTITSNDGTYVTLDAPLHVVNVHLYQNTILPMQQGGLISKDARMTPFSL---IVTFPSG 520
Query: 488 HIE 490
E
Sbjct: 521 AAE 529
>AV428776
Length = 331
Score = 30.0 bits (66), Expect = 0.97
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = -1
Query: 335 FHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDW 368
+HGL+ D V+AA L+ I+P+ +G +L W
Sbjct: 181 YHGLLYDPPFLVVAAFLMLILPSSWDDGLRLRAW 80
>TC10364 similar to UP|Q9ARE3 (Q9ARE3) ZF-HD homeobox protein, partial (14%)
Length = 1117
Score = 28.9 bits (63), Expect = 2.2
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -3
Query: 250 WFFFGFPVAVLLLLCFWCILCL 271
W FF FP++ LL +WCI L
Sbjct: 659 WIFFSFPISEEDLLWWWCIQTL 594
>TC18026 similar to UP|AAR87415 (AAR87415) Surface antigen (Fragment),
partial (13%)
Length = 522
Score = 28.9 bits (63), Expect = 2.2
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = -2
Query: 304 AEKMVLCVFGLLIVLWMTRRISDDLPGW 331
+ K++LC + L+I LW++ + PGW
Sbjct: 311 SNKLMLCTWALVIFLWISFSVQLRKPGW 228
>BP055345
Length = 504
Score = 27.7 bits (60), Expect = 4.8
Identities = 19/59 (32%), Positives = 28/59 (47%)
Frame = +2
Query: 150 NPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVIL 208
+P + +L TT S W+H A M +PVA + R P E +L + RA+ L
Sbjct: 260 HPELSILWPAHTTQVDSYWIHIFQLADMQIPVA--LSGRTPCRESSCQLHQELYRALYL 430
>TC9565 similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (19%)
Length = 872
Score = 27.7 bits (60), Expect = 4.8
Identities = 16/44 (36%), Positives = 25/44 (56%)
Frame = +3
Query: 13 PLQEPIQETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKL 56
PL P T ++ ++I S + Y +LGP LS ++ LLV+L
Sbjct: 387 PLPPP---TSSSGYLSISSAASSLERYCLLGPCLSFLVLLLVEL 509
>TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein
(RNA-binding protein S1, serine-rich domain), partial
(15%)
Length = 932
Score = 27.3 bits (59), Expect = 6.3
Identities = 15/55 (27%), Positives = 27/55 (48%)
Frame = -2
Query: 19 QETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVF 73
Q+TPN+NS+ + +L+ Q I++ +S + P + +LG W F
Sbjct: 790 QDTPNSNSMLMIFLLHTQLHSILIQGGISSLNVKNNNHQIPNSKFNILGHCCWCF 626
>TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor, partial
(74%)
Length = 1591
Score = 26.9 bits (58), Expect = 8.2
Identities = 16/43 (37%), Positives = 20/43 (46%), Gaps = 3/43 (6%)
Frame = -1
Query: 56 LDAPTTSTKMLGVIAWVFTWWITNAVPLPVTS---MCPLFLFP 95
L P ST + ++ + T W T VPL S C LF FP
Sbjct: 385 LALPVWSTGSVTLLFFTLTVWSTRCVPLLTLSPPLSCSLFSFP 257
>TC10036 weakly similar to UP|Q9LFB5 (Q9LFB5) Anthranilate
N-benzoyltransferase-like protein (AT5g01210/F7J8_190),
partial (32%)
Length = 502
Score = 26.9 bits (58), Expect = 8.2
Identities = 18/70 (25%), Positives = 32/70 (45%), Gaps = 1/70 (1%)
Frame = +2
Query: 430 FITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHI 489
+IT NDA + H+ +N V P L+ + FA+ +P S + + A +
Sbjct: 293 YITCNDAGVDFIHAKAKHLTLNAVVSPGLVDVHPCFKEFFAYDMPVSYSGHHIPLAAVQV 472
Query: 490 -EIKDMLKVG 498
E+ D + +G
Sbjct: 473 TELADAVFIG 502
>TC8301
Length = 1243
Score = 26.9 bits (58), Expect = 8.2
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -1
Query: 246 SFNSWFFFGFPVAVLLLLCFWCILCL 271
+F+S FF + LLC+WCIL L
Sbjct: 82 NFSSLDFFSQTAIHVELLCWWCILSL 5
>AV410706
Length = 427
Score = 26.9 bits (58), Expect = 8.2
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 3/47 (6%)
Frame = +1
Query: 144 FCGDPLNPSMLLLGLCATTFFVSMWLHNVA---TAVMMMPVATGILH 187
FC + SMLLLG +++ + LH +A ++M P TG LH
Sbjct: 247 FCRTIASSSMLLLGKS*SSWLMIEMLHMLA*FWLLILMSPCFTGGLH 387
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,350,826
Number of Sequences: 28460
Number of extensions: 162260
Number of successful extensions: 1369
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1352
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1368
length of query: 530
length of database: 4,897,600
effective HSP length: 95
effective length of query: 435
effective length of database: 2,193,900
effective search space: 954346500
effective search space used: 954346500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148482.2


