

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148470.10 - phase: 0 /pseudo
(598 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
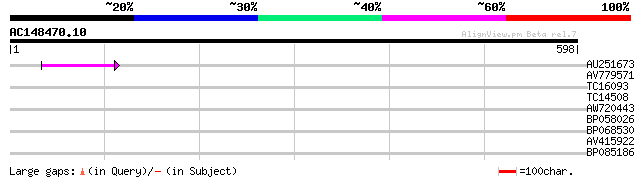
Score E
Sequences producing significant alignments: (bits) Value
AU251673 52 2e-07
AV779571 30 1.1
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 28 3.3
TC14508 similar to AAP86659 (AAP86659) 26S proteasome subunit RP... 28 3.3
AW720443 28 3.3
BP058026 27 7.2
BP068530 27 7.2
AV415922 27 7.2
BP085186 27 7.2
>AU251673
Length = 413
Score = 52.4 bits (124), Expect = 2e-07
Identities = 29/84 (34%), Positives = 44/84 (51%), Gaps = 1/84 (1%)
Frame = +2
Query: 34 CSEVQKVRFGTHMLAEEANDWWVSLLPILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEI 93
CS+ + V + L A DW+ L TWA F EF+NR++P+ VR
Sbjct: 5 CSDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVR 184
Query: 94 EFLELKQGD-MSVTEYAAEFSKSS 116
+F L+Q + M+V+EY+A F+ S
Sbjct: 185 DFERLEQAEGMTVSEYSAHFTHLS 256
>AV779571
Length = 538
Score = 30.0 bits (66), Expect = 1.1
Identities = 21/58 (36%), Positives = 38/58 (65%)
Frame = -2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLLLFV 341
+++ ++L LLL +L+LLIV ++L + I V + + LLK++LR+Q LL+L +
Sbjct: 321 LIIYHLLLSLLLLLLLLLLIVIMILSLTIE*VLML*V-----LLKMRLRLQQLLILLI 163
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 28.5 bits (62), Expect = 3.3
Identities = 22/61 (36%), Positives = 35/61 (57%), Gaps = 5/61 (8%)
Frame = -1
Query: 301 LLIVSLLLIVHIS-----WVWLYLI*MEKWLLKLQLRVQ*LLLLFV*VVHCLCLVETLKS 355
LL++ LLL++H+ W+ L L+ + WLLKL L LL+L + L L+ + KS
Sbjct: 240 LLMLFLLLLLHLLLWLLYWLLLLLLLLLNWLLKLMLV---LLVLLRMNLELLKLLSS*KS 70
Query: 356 I 356
+
Sbjct: 69 V 67
>TC14508 similar to AAP86659 (AAP86659) 26S proteasome subunit RPN5a,
partial (96%)
Length = 1957
Score = 28.5 bits (62), Expect = 3.3
Identities = 13/28 (46%), Positives = 16/28 (56%), Gaps = 1/28 (3%)
Frame = -3
Query: 454 QRGRLNFQLILFPER-SRCRWHLTVCRR 480
Q GRL F + R S CRW L+ CR+
Sbjct: 251 QSGRLGFSRLAHQSRASECRWRLSWCRK 168
>AW720443
Length = 406
Score = 28.5 bits (62), Expect = 3.3
Identities = 18/30 (60%), Positives = 21/30 (70%)
Frame = +3
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHISW 314
L S ++LL LL*+LVLLIVS LL I W
Sbjct: 204 LDSRMMLLSTLL*LLVLLIVSPLLSQSIQW 293
>BP058026
Length = 389
Score = 27.3 bits (59), Expect = 7.2
Identities = 13/31 (41%), Positives = 25/31 (79%)
Frame = -2
Query: 280 DLSEVLVSLIVLL*LLL*ILVLLIVSLLLIV 310
DL+++ + ++LL LLL +L+LL++ LLL++
Sbjct: 109 DLNKIHIKSLLLLLLLLLLLLLLLLLLLLLL 17
>BP068530
Length = 517
Score = 27.3 bits (59), Expect = 7.2
Identities = 11/30 (36%), Positives = 21/30 (69%)
Frame = -2
Query: 298 ILVLLIVSLLLIVHISWVWLYLI*MEKWLL 327
+L+L ++ LLL++ + WV L L+ + W+L
Sbjct: 462 VLLLALLFLLLLLRV*WVVLSLVAISAWVL 373
>AV415922
Length = 422
Score = 27.3 bits (59), Expect = 7.2
Identities = 13/30 (43%), Positives = 23/30 (76%)
Frame = -1
Query: 288 LIVLL*LLL*ILVLLIVSLLLIVHISWVWL 317
L++LL LLL +L+LL+ SLL+++ + W+
Sbjct: 233 LLLLLLLLLLLLLLLLSSLLVLIPLELNWV 144
>BP085186
Length = 388
Score = 27.3 bits (59), Expect = 7.2
Identities = 13/24 (54%), Positives = 21/24 (87%)
Frame = +1
Query: 288 LIVLL*LLL*ILVLLIVSLLLIVH 311
L++LL LLL +L+LL++ LLL++H
Sbjct: 190 LVLLLLLLLLLLLLLLLLLLLLLH 261
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.370 0.166 0.609
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,406,062
Number of Sequences: 28460
Number of extensions: 156261
Number of successful extensions: 2084
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1106
Number of HSP's successfully gapped in prelim test: 70
Number of HSP's that attempted gapping in prelim test: 858
Number of HSP's gapped (non-prelim): 1264
length of query: 598
length of database: 4,897,600
effective HSP length: 95
effective length of query: 503
effective length of database: 2,193,900
effective search space: 1103531700
effective search space used: 1103531700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148470.10


