

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
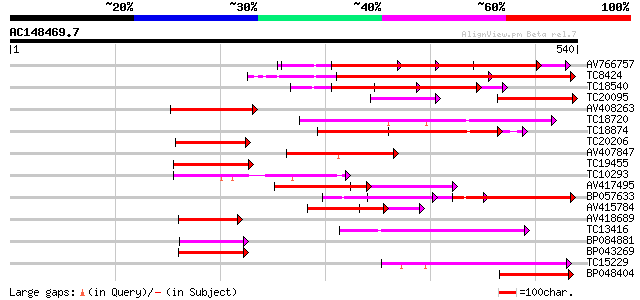
Query= AC148469.7 - phase: 0
(540 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV766757 238 2e-63
TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm rep... 186 6e-48
TC18540 similar to UP|Q944I3 (Q944I3) AT3g01400/T13O15_4, partia... 138 3e-33
TC20095 137 4e-33
AV408263 119 9e-28
TC18720 weakly similar to UP|SCWA_YEAST (Q04951) Probable family... 108 2e-24
TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5,... 106 1e-23
TC20206 97 7e-21
AV407847 95 3e-20
TC19455 similar to UP|Q8S8Z6 (Q8S8Z6) Syringolide-induced protei... 88 4e-18
TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%) 87 9e-18
AV417495 83 1e-16
BP057633 82 2e-16
AV415784 74 5e-14
AV418689 74 6e-14
TC13416 weakly similar to GB|AAP21292.1|30102748|BT006484 At1g67... 73 1e-13
BP084881 72 3e-13
BP043269 69 3e-12
TC15229 weakly similar to UP|O60585 (O60585) Ser/Arg-related nuc... 64 8e-11
BP048404 59 2e-09
>AV766757
Length = 601
Score = 238 bits (607), Expect = 2e-63
Identities = 120/200 (60%), Positives = 154/200 (77%)
Frame = +2
Query: 307 VSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATL 366
V LL+ D TQE+AVT++LNLSIYENNKG I+ +GA+P IV VLR G+MEARENAAATL
Sbjct: 2 VGLLSVPDSRTQEHAVTALLNLSIYENNKGCIVSSGAVPGIVHVLRKGSMEARENAAATL 181
Query: 367 FSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGII 426
FSLS+ DENK+ IG+SGAI LV LL G+ RGKKDAATALFNLCIYQGNKG+A+RAG+I
Sbjct: 182 FSLSVVDENKVTIGSSGAIPPLVTLLSEGTQRGKKDAATALFNLCIYQGNKGKAVRAGVI 361
Query: 427 TALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAA 486
L+ +LT+ S MVDEAL I+++L+SH + K +I A +P+L++ + G PRNKEN+A
Sbjct: 362 PTLMKLLTEPSGGMVDEALAILAILSSHPDGKAAIGAADAVPILVEFIGNGSPRNKENSA 541
Query: 487 AILLALCKRDTDNLSCISRL 506
A+L+ L D L+ +L
Sbjct: 542 AVLVHLSSGDQQYLAQAHKL 601
Score = 89.7 bits (221), Expect = 1e-18
Identities = 55/151 (36%), Positives = 88/151 (57%)
Frame = +2
Query: 260 AIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQE 319
A+ +V L S+E A A + SLS +N++ I +GAIP LV+LL+ ++
Sbjct: 110 AVPGIVHVLRKGSMEARENAAATLFSLSVVD-ENKVTIGSSGAIPPLVTLLSEGTQRGKK 286
Query: 320 NAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIII 379
+A T++ NL IY+ NKG + AG IP+++++L + + A A L LS + K I
Sbjct: 287 DAATALFNLCIYQGNKGKAVRAGVIPTLMKLLTEPSGGMVDEALAILAILSSHPDGKAAI 466
Query: 380 GASGAISALVDLLQNGSPRGKKDAATALFNL 410
GA+ A+ LV+ + NGSPR K+++A L +L
Sbjct: 467 GAADAVPILVEFIGNGSPRNKENSAAVLVHL 559
Score = 51.2 bits (121), Expect = 4e-07
Identities = 32/93 (34%), Positives = 51/93 (54%)
Frame = +2
Query: 442 DEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS 501
+ A+T + L+ ++ K IV + +P ++ +LR G +ENAAA L +L D +N
Sbjct: 38 EHAVTALLNLSIYENNKGCIVSSGAVPGIVHVLRKGSMEARENAAATLFSLSVVD-ENKV 214
Query: 502 CISRLGAVIPLSELARTGTERAKRKATSLLEHL 534
I GA+ PL L GT+R K+ A + L +L
Sbjct: 215 TIGSSGAIPPLVTLLSEGTQRGKKDAATALFNL 313
Score = 46.2 bits (108), Expect = 1e-05
Identities = 36/119 (30%), Positives = 55/119 (45%)
Frame = +2
Query: 256 GDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDV 315
G AI LV LS + + A + +L N+ AG IP L+ LLT
Sbjct: 221 GSSGAIPPLVTLLSEGTQRGKKDAATALFNLCIYQ-GNKGKAVRAGVIPTLMKLLTEPSG 397
Query: 316 MTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADE 374
+ A+ + LS + + K I A A+P +V+ + G+ +EN+AA L LS D+
Sbjct: 398 GMVDEALAILAILSSHPDGKAAIGAADAVPILVEFIGNGSPRNKENSAAVLVHLSSGDQ 574
>TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm repeat
containing protein homolog-like {Oryza sativa (japonica
cultivar-group);} , partial (49%)
Length = 1586
Score = 186 bits (473), Expect = 6e-48
Identities = 99/228 (43%), Positives = 148/228 (64%)
Frame = +3
Query: 312 SEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSL 371
S D TQE+AVT++LNLS+ + NK LI AGA+ S++ VL+ GT +++NAA L SL+L
Sbjct: 390 SSDPWTQEHAVTALLNLSLNDENKKLITNAGAVKSLIYVLKTGTETSKQNAACALLSLAL 569
Query: 372 ADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLN 431
+EN+ IGASGAI LV LL NGS RGKKDA T L+ LC + NK RA+ AG++ L++
Sbjct: 570 VEENRSSIGASGAIPPLVSLLINGSSRGKKDALTTLYKLCSVKLNKERAVNAGVVKPLVD 749
Query: 432 MLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLA 491
++ + + ++A+ +++ LA+ Q+ K +IV+ I L++ + G + KE A LL
Sbjct: 750 LVAEQGTGLAEKAMVVLNSLAAVQDGKDAIVEEGGIAALVEAIEDGSVKGKEFAVLTLLQ 929
Query: 492 LCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
LC N + R G + PL L+++GT RAK KA +LL +LR+ +Q
Sbjct: 930 LCVDSVRNRGLLVREGGIPPLVSLSQSGTPRAKHKAETLLRYLRESRQ 1073
Score = 80.1 bits (196), Expect = 8e-16
Identities = 68/236 (28%), Positives = 117/236 (48%), Gaps = 2/236 (0%)
Frame = +3
Query: 227 WCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSL 286
W EH + T L N + +D + + +T + A+++L+ L + + A + SL
Sbjct: 402 WTQEHAV---TALLN--LSLNDENKKLIT-NAGAVKSLIYVLKTGTETSKQNAACALLSL 563
Query: 287 SKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPS 346
+ +NR I +GAIP LVSLL + +++A+T++ L + NK + AG +
Sbjct: 564 ALVE-ENRSSIGASGAIPPLVSLLINGSSRGKKDALTTLYKLCSVKLNKERAVNAGVVKP 740
Query: 347 IVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATA 406
+V ++ E A L SL+ + K I G I+ALV+ +++GS +GK+ A
Sbjct: 741 LVDLVAEQGTGLAEKAMVVLNSLAAVQDGKDAIVEEGGIAALVEAIEDGSVKGKEFAVLT 920
Query: 407 LFNLCIYQ-GNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVL-ASHQEAKVS 460
L LC+ N+G +R G I L+++ + +A T++ L S QEA S
Sbjct: 921 LLQLCVDSVRNRGLLVREGGIPPLVSLSQSGTPRAKHKAETLLRYLRESRQEASSS 1088
>TC18540 similar to UP|Q944I3 (Q944I3) AT3g01400/T13O15_4, partial (40%)
Length = 433
Score = 138 bits (347), Expect = 3e-33
Identities = 72/143 (50%), Positives = 102/143 (70%)
Frame = +3
Query: 307 VSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATL 366
+SL++S+D+ QE VT+ILNLS+ + NK LI +GAI +V+ L+AGT A+ENAA L
Sbjct: 3 ISLISSQDLQLQEYGVTAILNLSLCDENKELIASSGAIKPLVRALKAGTPTAKENAACAL 182
Query: 367 FSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGII 426
LS A+E+K IG SGAI LV LLQ G RGKKDA+TAL++LC + NK RA++AGI+
Sbjct: 183 LRLSQAEESKAAIGRSGAIPLLVSLLQTGGIRGKKDASTALYSLCSVRENKIRAVKAGIM 362
Query: 427 TALLNMLTDSSKSMVDEALTIMS 449
L+ ++ D +MVD++ ++S
Sbjct: 363 KVLVELMADFESNMVDKSAYVVS 431
Score = 71.6 bits (174), Expect = 3e-13
Identities = 42/127 (33%), Positives = 71/127 (55%)
Frame = +3
Query: 348 VQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATAL 407
+ ++ + ++ +E + +LSL DENK +I +SGAI LV L+ G+P K++AA AL
Sbjct: 3 ISLISSQDLQLQEYGVTAILNLSLCDENKELIASSGAIKPLVRALKAGTPTAKENAACAL 182
Query: 408 FNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTI 467
L + +K R+G I L+++L +A T + L S +E K+ VKA +
Sbjct: 183 LRLSQAEESKAAIGRSGAIPLLVSLLQTGGIRGKKDASTALYSLCSVRENKIRAVKAGIM 362
Query: 468 PVLIDLL 474
VL++L+
Sbjct: 363 KVLVELM 383
Score = 67.0 bits (162), Expect = 7e-12
Identities = 43/125 (34%), Positives = 70/125 (55%)
Frame = +3
Query: 268 LSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILN 327
+S + ++ V I +LS +N+ LIA +GAI LV L + +ENA ++L
Sbjct: 12 ISSQDLQLQEYGVTAILNLSL-CDENKELIASSGAIKPLVRALKAGTPTAKENAACALLR 188
Query: 328 LSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISA 387
LS E +K I +GAIP +V +L+ G + +++A+ L+SL ENKI +G +
Sbjct: 189 LSQAEESKAAIGRSGAIPLLVSLLQTGGIRGKKDASTALYSLCSVRENKIRAVKAGIMKV 368
Query: 388 LVDLL 392
LV+L+
Sbjct: 369 LVELM 383
>TC20095
Length = 475
Score = 137 bits (345), Expect = 4e-33
Identities = 69/76 (90%), Positives = 74/76 (96%)
Frame = +2
Query: 465 STIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAK 524
STIPVLIDLLRTGLPRNKENAAAILLALCK+D +NL+CISRLG+VIPL+EL RTGTERAK
Sbjct: 2 STIPVLIDLLRTGLPRNKENAAAILLALCKKDAENLACISRLGSVIPLTELTRTGTERAK 181
Query: 525 RKATSLLEHLRKLQQL 540
RKATSLLEHLRKLQQL
Sbjct: 182 RKATSLLEHLRKLQQL 229
Score = 43.1 bits (100), Expect = 1e-04
Identities = 25/68 (36%), Positives = 38/68 (55%), Gaps = 1/68 (1%)
Frame = +2
Query: 344 IPSIVQVLRAGTMEARENAAATLFSLSLAD-ENKIIIGASGAISALVDLLQNGSPRGKKD 402
IP ++ +LR G +ENAAA L +L D EN I G++ L +L + G+ R K+
Sbjct: 8 IPVLIDLLRTGLPRNKENAAAILLALCKKDAENLACISRLGSVIPLTELTRTGTERAKRK 187
Query: 403 AATALFNL 410
A + L +L
Sbjct: 188 ATSLLEHL 211
Score = 30.8 bits (68), Expect = 0.58
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 1/65 (1%)
Frame = +2
Query: 303 IPVLVSLLTSEDVMTQENAVTSILNLSIYE-NNKGLIMLAGAIPSIVQVLRAGTMEAREN 361
IPVL+ LL + +ENA +L L + N I G++ + ++ R GT A+
Sbjct: 8 IPVLIDLLRTGLPRNKENAAAILLALCKKDAENLACISRLGSVIPLTELTRTGTERAKRK 187
Query: 362 AAATL 366
A + L
Sbjct: 188 ATSLL 202
>AV408263
Length = 331
Score = 119 bits (299), Expect = 9e-28
Identities = 55/83 (66%), Positives = 64/83 (76%)
Frame = +2
Query: 154 KKPDAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLT 213
K A+VIP DF CPISLELM+DPVIV+TGQTYERS I++W+ G+ TCPKTQQ L
Sbjct: 77 KNHPALVIPTDFKCPISLELMQDPVIVSTGQTYERSCIEKWLQAGHGTCPKTQQSLTSTV 256
Query: 214 LTPNYVLRSLVSQWCIEHNIEQP 236
LTPNYVLRSL+ QWC + IE P
Sbjct: 257 LTPNYVLRSLIEQWCEANGIEPP 325
>TC18720 weakly similar to UP|SCWA_YEAST (Q04951) Probable family 17
glucosidase SCW10 precursor (Soluble cell wall protein
10) , partial (7%)
Length = 871
Score = 108 bits (271), Expect = 2e-24
Identities = 80/254 (31%), Positives = 138/254 (53%), Gaps = 10/254 (3%)
Frame = +3
Query: 277 RAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSI-YENNK 335
R A A +R L+K + R +A GAIP L ++L SED+ +Q+ ++ ++LNL I + NK
Sbjct: 114 REAAATVRMLAKEDSHVRGTLAMLGAIPPLGAMLDSEDLDSQKASLYALLNLGIGNDANK 293
Query: 336 GLIMLAGAIPSIVQVLRAGT-MEAR--ENAAATLFSLSLADENKIIIGASGAISALVDLL 392
I+ G++ +++++ + +++R E A LS D NK IIG+S A+ LV L
Sbjct: 294 AAIVKVGSVHKMLKLIESPEGLDSRVSEAIVANFLGLSALDSNKPIIGSSAAVPFLVRTL 473
Query: 393 QNG----SPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIM 448
Q+ S + K+DA AL+NL I+ GN + ++ L+N + D + + +L+I+
Sbjct: 474 QSSDVKTSSQAKQDALRALYNLSIFPGNISFILETHLVPFLINSIGD--MEVTERSLSIL 647
Query: 449 SVLASHQEAKVSI-VKASTIPVLIDLLR-TGLPRNKENAAAILLALCKRDTDNLSCISRL 506
S L S +E + +I T P+L+D+L T P +E + +L+ + + + +
Sbjct: 648 SNLVSTREGRKAISAVPDTFPILVDVLNWTDSPECQEKTSYVLMVMAHKSFGDRQGMIEA 827
Query: 507 GAVIPLSELARTGT 520
G V L EL+ GT
Sbjct: 828 GIVSSLLELSLLGT 869
>TC18874 similar to UP|Q9FJJ0 (Q9FJJ0) Genomic DNA, chromosome 5, TAC
clone:K19B1, partial (33%)
Length = 543
Score = 106 bits (264), Expect = 1e-23
Identities = 55/176 (31%), Positives = 107/176 (60%)
Frame = +2
Query: 294 RILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRA 353
R+ + + L L+ S V+ Q NAV S++NLS+ + NK I+ +G +P ++ VL+
Sbjct: 20 RVSLCTPRLLSALRLLVESRYVVVQVNAVASLVNLSLEKTNKVRIVRSGFVPFLIDVLKG 199
Query: 354 GTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIY 413
G E++E+AA LFSL+L D+NK+ IG GA+ L+ L++ S R + D+A AL++L +
Sbjct: 200 GFSESQEHAAGALFSLALDDDNKMAIGVLGALQPLMHALRSESERTRHDSALALYHLTLV 379
Query: 414 QGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPV 469
Q N+ + ++ G++ LL+M+ + ++ L ++ +A+ E + +++ + + +
Sbjct: 380 QSNRVKLVKLGVVPTLLSMVANG--NLASRVLLVLCNVAACVEGRTAMLDGNAVEI 541
Score = 63.5 bits (153), Expect = 8e-11
Identities = 44/134 (32%), Positives = 71/134 (52%), Gaps = 1/134 (0%)
Frame = +2
Query: 361 NAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRA 420
NA A+L +LSL NK+ I SG + L+D+L+ G ++ AA ALF+L + NK
Sbjct: 98 NAVASLVNLSLEKTNKVRIVRSGFVPFLIDVLKGGFSESQEHAAGALFSLALDDDNKMAI 277
Query: 421 IRAGIITALLNML-TDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLP 479
G + L++ L ++S ++ D AL + L Q +V +VK +P L+ ++ G
Sbjct: 278 GVLGALQPLMHALRSESERTRHDSALALYH-LTLVQSNRVKLVKLGVVPTLLSMVANG-- 448
Query: 480 RNKENAAAILLALC 493
A+ +LL LC
Sbjct: 449 ---NLASRVLLVLC 481
>TC20206
Length = 508
Score = 97.1 bits (240), Expect = 7e-21
Identities = 42/71 (59%), Positives = 52/71 (73%)
Frame = +1
Query: 159 IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNY 218
I IP + CPISLELMRDPV V TGQTY+RS I+ W++ GNTTCP T+ L TL PN+
Sbjct: 127 IQIPYHYRCPISLELMRDPVTVCTGQTYDRSSIESWVNTGNTTCPVTRAPLTEFTLIPNH 306
Query: 219 VLRSLVSQWCI 229
LR L+ +WC+
Sbjct: 307 TLRRLIQEWCV 339
>AV407847
Length = 423
Score = 95.1 bits (235), Expect = 3e-20
Identities = 61/111 (54%), Positives = 76/111 (67%), Gaps = 4/111 (3%)
Frame = +1
Query: 264 LVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLT--SEDVMTQENA 321
LV KL+ ++ES A V E+R L+K +D+R IAEAGAIPVLV LT S Q NA
Sbjct: 37 LVNKLNVSDLDESNAVVYELRVLAKTDSDSRACIAEAGAIPVLVRYLTTGSHHPSLQVNA 216
Query: 322 VTSILNLSIYENNKGLIM-LAGAIPSIVQVLRAG-TMEARENAAATLFSLS 370
VT+ILNLSI E NK IM GA+ + +VLR+G T EA+ NAAAT+FSL+
Sbjct: 217 VTTILNLSILEANKTRIMETDGALNGVAEVLRSGATWEAKANAAATVFSLT 369
>TC19455 similar to UP|Q8S8Z6 (Q8S8Z6) Syringolide-induced protein 13-1-1,
partial (25%)
Length = 485
Score = 87.8 bits (216), Expect = 4e-18
Identities = 36/76 (47%), Positives = 53/76 (69%)
Frame = +1
Query: 157 DAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTP 216
+ IVIP F CP+SL+LM+DPV + TG TY+R+ I++WI+ GN CP T Q L L P
Sbjct: 148 EEIVIPNHFRCPVSLDLMKDPVTLPTGITYDRTSIEKWIEAGNKKCPVTNQILTTFDLIP 327
Query: 217 NYVLRSLVSQWCIEHN 232
N+ LR ++ WC++++
Sbjct: 328 NHSLRRMIQDWCVKNS 375
>TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%)
Length = 757
Score = 86.7 bits (213), Expect = 9e-18
Identities = 61/179 (34%), Positives = 89/179 (49%), Gaps = 11/179 (6%)
Frame = +3
Query: 157 DAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNT----TCPKTQQKLQ-- 210
D + +P FLCPISLE+M+DPV V TG TY+R I++W+ N TCP T+Q L
Sbjct: 96 DEVDVPSYFLCPISLEIMKDPVTVTTGITYDRESIEKWLFSKNNTKTITCPVTKQPLSDC 275
Query: 211 --HLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKL 268
LTLTPN+ LR L+ WC + S G R T +T + KL
Sbjct: 276 SADLTLTPNHTLRRLIQAWC--------------TMNASHGVERIPTPKPPVNKTQISKL 413
Query: 269 ---SCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTS 324
+ RS + ++S++ S N+ I AG + L S++ + + ++AVTS
Sbjct: 414 LKDASRSPLMQIKCLQRLKSIASGSETNKRCIEAAGGVEFLASIVMN----SIDSAVTS 578
>AV417495
Length = 316
Score = 82.8 bits (203), Expect = 1e-16
Identities = 47/92 (51%), Positives = 58/92 (62%)
Frame = +3
Query: 253 DVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTS 312
D++G A + LV L ++ R A EIR L+K + DNRI IA GAI +LV LL S
Sbjct: 36 DLSGIEAQVRGLVEGLRSSDLDTQREATVEIRFLAKHNMDNRIAIANCGAISLLVELLKS 215
Query: 313 EDVMTQENAVTSILNLSIYENNKGLIMLAGAI 344
D QENAVT++LNLSI +NNK I AGAI
Sbjct: 216 TDTRIQENAVTALLNLSINDNNKAAIANAGAI 311
Score = 50.8 bits (120), Expect = 5e-07
Identities = 33/103 (32%), Positives = 54/103 (52%), Gaps = 1/103 (0%)
Frame = +3
Query: 325 ILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLAD-ENKIIIGASG 383
I++ S E L + + +V+ LR+ ++ + A + L+ + +N+I I G
Sbjct: 3 IISSSAIETRADLSGIEAQVRGLVEGLRSSDLDTQREATVEIRFLAKHNMDNRIAIANCG 182
Query: 384 AISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGII 426
AIS LV+LL++ R +++A TAL NL I NK AG I
Sbjct: 183 AISLLVELLKSTDTRIQENAVTALLNLSINDNNKAAIANAGAI 311
>BP057633
Length = 570
Score = 82.0 bits (201), Expect = 2e-16
Identities = 43/118 (36%), Positives = 75/118 (63%)
Frame = -2
Query: 422 RAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRN 481
+AG + L+ ++ +++ MVD+A+ ++S L++ E + I + IP+L++++ +G R
Sbjct: 569 QAGAVKFLVQLM-NTADGMVDKAVALLSNLSTITEGRSEIAREGGIPLLVEIIESGSQRG 393
Query: 482 KENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
KENAA+ILL LC + + + + GAV PL L+++GT RAK KA LL H R ++
Sbjct: 392 KENAASILLQLCLHSSKFCTLVLQEGAVPPLVALSQSGTPRAKEKAQQLLSHFRNQRE 219
Score = 62.4 bits (150), Expect = 2e-10
Identities = 38/110 (34%), Positives = 61/110 (54%), Gaps = 1/110 (0%)
Frame = -2
Query: 299 EAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEA 358
+AGA+ LV L+ + D M + AV + NLS + I G IP +V+++ +G+
Sbjct: 569 QAGAVKFLVQLMNTADGMV-DKAVALLSNLSTITEGRSEIAREGGIPLLVEIIESGSQRG 393
Query: 359 RENAAATLFSLSL-ADENKIIIGASGAISALVDLLQNGSPRGKKDAATAL 407
+ENAA+ L L L + + ++ GA+ LV L Q+G+PR K+ A L
Sbjct: 392 KENAASILLQLCLHSSKFCTLVLQEGAVPPLVALSQSGTPRAKEKAQQLL 243
Score = 48.9 bits (115), Expect = 2e-06
Identities = 31/117 (26%), Positives = 60/117 (50%), Gaps = 1/117 (0%)
Frame = -2
Query: 341 AGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGK 400
AGA+ +VQ++ + A A L +LS E + I G I LV+++++GS RGK
Sbjct: 566 AGAVKFLVQLMNTADGMV-DKAVALLSNLSTITEGRSEIAREGGIPLLVEIIESGSQRGK 390
Query: 401 KDAATALFNLCIYQGN-KGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQE 456
++AA+ L LC++ ++ G + L+ + + ++A ++S + +E
Sbjct: 389 ENAASILLQLCLHSSKFCTLVLQEGAVPPLVALSQSGTPRAKEKAQQLLSHFRNQRE 219
>AV415784
Length = 242
Score = 74.3 bits (181), Expect = 5e-14
Identities = 44/79 (55%), Positives = 56/79 (70%), Gaps = 2/79 (2%)
Frame = +1
Query: 284 RSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIM-LAG 342
R L+K +NR IAEAGAIP L +LL+S + + QEN+VT++LNLSI++ NK IM G
Sbjct: 4 RFLAKTGRENRAFIAEAGAIPYLRNLLSSHNAVAQENSVTALLNLSIFDKNKSRIMDEEG 183
Query: 343 AIPSIVQVLRAG-TMEARE 360
+ SIV VLR G T EARE
Sbjct: 184 CLGSIVDVLRFGHTTEARE 240
Score = 42.0 bits (97), Expect = 3e-04
Identities = 23/63 (36%), Positives = 38/63 (59%), Gaps = 1/63 (1%)
Frame = +1
Query: 334 NKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKI-IIGASGAISALVDLL 392
N+ I AGAIP + +L + A+EN+ L +LS+ D+NK I+ G + ++VD+L
Sbjct: 31 NRAFIAEAGAIPYLRNLLSSHNAVAQENSVTALLNLSIFDKNKSRIMDEEGCLGSIVDVL 210
Query: 393 QNG 395
+ G
Sbjct: 211 RFG 219
Score = 36.2 bits (82), Expect = 0.014
Identities = 20/61 (32%), Positives = 37/61 (59%), Gaps = 1/61 (1%)
Frame = +1
Query: 374 ENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAI-RAGIITALLNM 432
EN+ I +GAI L +LL + + ++++ TAL NL I+ NK R + G + +++++
Sbjct: 28 ENRAFIAEAGAIPYLRNLLSSHNAVAQENSVTALLNLSIFDKNKSRIMDEEGCLGSIVDV 207
Query: 433 L 433
L
Sbjct: 208 L 210
>AV418689
Length = 371
Score = 73.9 bits (180), Expect = 6e-14
Identities = 31/61 (50%), Positives = 44/61 (71%)
Frame = +3
Query: 161 IPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVL 220
IP F CPISL+LM+DPV ++TG TY+R +++W D GN TCP T Q +++ + PN+ L
Sbjct: 189 IPNHFRCPISLDLMKDPVTLSTGITYDRDSVEKWFDDGNYTCPVTNQIVKNFDMIPNHSL 368
Query: 221 R 221
R
Sbjct: 369 R 371
>TC13416 weakly similar to GB|AAP21292.1|30102748|BT006484 At1g67530
{Arabidopsis thaliana;}, partial (23%)
Length = 557
Score = 72.8 bits (177), Expect = 1e-13
Identities = 56/185 (30%), Positives = 96/185 (51%), Gaps = 4/185 (2%)
Frame = +2
Query: 315 VMTQENAVTSILNLSIYEN-NKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLAD 373
+M QE ++ NL++ N NK +++ AG +P + +++ + +A A +LS +
Sbjct: 2 LMAQEIGAMALFNLAVNNNGNKEMMLSAGVLPVLEEMI--SNTSSYGSATALYLNLSCLE 175
Query: 374 ENKIIIGASGAISALVDLLQNGSP-RGKKDAATALFNLCIYQGNKGRAIRAGIITALLNM 432
E K +IG S A+ L LLQ+ S + K+D+ AL+NL N + +GII L ++
Sbjct: 176 EAKPMIGTSQAVQFLTQLLQSDSDIQCKQDSLHALYNLSTVSTNIPYLL*SGIINGLQSL 355
Query: 433 LTDSSKSM-VDEALTIMSVLASHQEAKVSIVKA-STIPVLIDLLRTGLPRNKENAAAILL 490
L D + ++ + ++ LA+ Q + +V A I L +L G +E AA+ LL
Sbjct: 356 LVDQGDCVWTEKCIAVLINLATSQAGREEMVSAPGLISALASILDIGELIVQEQAASCLL 535
Query: 491 ALCKR 495
LC R
Sbjct: 536 ILCNR 550
>BP084881
Length = 488
Score = 71.6 bits (174), Expect = 3e-13
Identities = 32/66 (48%), Positives = 39/66 (58%)
Frame = -3
Query: 162 PEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLR 221
P F+CPI E+MRDP + A G TYE I+ W+D G+ T P T KL H L PN LR
Sbjct: 426 PSYFICPIFQEVMRDPHVAADGFTYEAEAIRGWLDSGHDTSPMTNSKLAHQNLVPNRALR 247
Query: 222 SLVSQW 227
S + W
Sbjct: 246 SAIQDW 229
>BP043269
Length = 413
Score = 68.6 bits (166), Expect = 3e-12
Identities = 27/67 (40%), Positives = 43/67 (63%)
Frame = -3
Query: 161 IPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVL 220
+P F+CPI ++M++P + A G +YE I++W+ GN P T +++H LTPN+ L
Sbjct: 366 VPSVFICPILQDVMKNPHVAADGFSYELEAIEQWLQSGNERSPMTNLRMKHTFLTPNHTL 187
Query: 221 RSLVSQW 227
RSL+ W
Sbjct: 186 RSLIDDW 166
>TC15229 weakly similar to UP|O60585 (O60585) Ser/Arg-related nuclear matrix
protein, partial (4%)
Length = 965
Score = 63.5 bits (153), Expect = 8e-11
Identities = 55/189 (29%), Positives = 87/189 (45%), Gaps = 8/189 (4%)
Frame = +3
Query: 355 TMEARENAAATLFSLSL----ADENKIIIGASGAISALVDLLQN--GSPRGKKDAATALF 408
+++ R N+AA + + AD + G +V++L++ PR K ALF
Sbjct: 66 SLDVRVNSAALIEMVVAGTHSADLRSQVSSVEGIHDGVVEILRSPISYPRALKVGVKALF 245
Query: 409 NLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIV-KASTI 467
LC+ + ++ RA+ AG L++ L D K + AL + +L A T+
Sbjct: 246 ALCLVKQSRQRAVAAGAPAVLIDRLADFEKCDAERALATVELLCRVPAGCAEFAGHALTV 425
Query: 468 PVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTG-TERAKRK 526
P+L+ ++ R E AA LL+LC G + L L ++ TERAKRK
Sbjct: 426 PMLVKIILKISDRATEYAAGALLSLCSESERLQMEAVAAGVLTQLLLLVQSDCTERAKRK 605
Query: 527 ATSLLEHLR 535
A LL+ LR
Sbjct: 606 AQLLLKLLR 632
>BP048404
Length = 457
Score = 58.9 bits (141), Expect = 2e-09
Identities = 34/73 (46%), Positives = 46/73 (62%), Gaps = 2/73 (2%)
Frame = -3
Query: 467 IPVLIDLLRTGL-PRNKENAAAILLALCKRDTDNLSCI-SRLGAVIPLSELARTGTERAK 524
+P L+ ++R GL R+KEN AIL A+C D L + + +SELARTGT RAK
Sbjct: 440 VPSLLSIMREGLCERSKENCVAILQAICLYDRSKLKEVRDEENSHRTISELARTGTSRAK 261
Query: 525 RKATSLLEHLRKL 537
RKAT +L+ L K+
Sbjct: 260 RKATGILDRLNKI 222
Score = 35.4 bits (80), Expect = 0.024
Identities = 21/72 (29%), Positives = 39/72 (54%), Gaps = 3/72 (4%)
Frame = -3
Query: 342 GAIPSIVQVLRAGTME-ARENAAATLFSLSLADENKI--IIGASGAISALVDLLQNGSPR 398
GA+PS++ ++R G E ++EN A L ++ L D +K+ + + + +L + G+ R
Sbjct: 446 GAVPSLLSIMREGLCERSKENCVAILQAICLYDRSKLKEVRDEENSHRTISELARTGTSR 267
Query: 399 GKKDAATALFNL 410
K+ A L L
Sbjct: 266 AKRKATGILDRL 231
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.131 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,486,592
Number of Sequences: 28460
Number of extensions: 88051
Number of successful extensions: 512
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 461
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 493
length of query: 540
length of database: 4,897,600
effective HSP length: 95
effective length of query: 445
effective length of database: 2,193,900
effective search space: 976285500
effective search space used: 976285500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148469.7


