

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.4 - phase: 0
(609 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
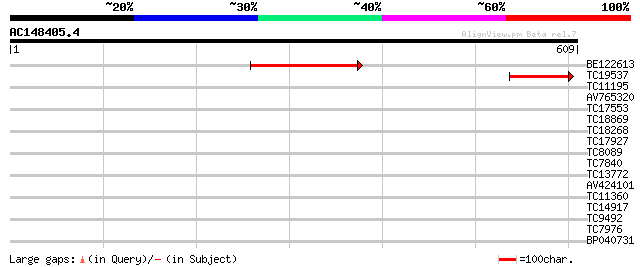
Sequences producing significant alignments: (bits) Value
BE122613 211 4e-55
TC19537 118 3e-27
TC11195 UP|Q9FJH4 (Q9FJH4) Similarity to ring finger protein, pa... 34 0.060
AV765320 34 0.078
TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrat... 32 0.30
TC18869 similar to UP|Q9SIR1 (Q9SIR1) At2g25320 protein, partial... 31 0.50
TC18268 weakly similar to UP|Q9C9S6 (Q9C9S6) Kinesin-related pro... 30 1.1
TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete 30 1.5
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 29 1.9
TC7840 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced ... 28 4.3
TC13772 27 7.3
AV424101 27 7.3
TC11360 homologue to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%) 27 7.3
TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Ar... 27 9.5
TC9492 homologue to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determi... 27 9.5
TC7976 homologue to UP|METK_LYCES (P43280) S-adenosylmethionine ... 27 9.5
BP040731 27 9.5
>BE122613
Length = 371
Score = 211 bits (536), Expect = 4e-55
Identities = 109/121 (90%), Positives = 114/121 (94%)
Frame = +1
Query: 259 GDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTD 318
G S+DLPG ASVSRETDI+ TKLKS+GDAQLVLP DQRNKLNR+LSTMQRRLVTAKTD
Sbjct: 7 GTSSLDLPGYASVSRETDIMDPTKLKSTGDAQLVLPQDQRNKLNRVLSTMQRRLVTAKTD 186
Query: 319 MEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEE 378
MEDLIVRLNQE+AAKDFL TKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEE
Sbjct: 187 MEDLIVRLNQEMAAKDFLATKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEE 366
Query: 379 L 379
L
Sbjct: 367 L 369
>TC19537
Length = 613
Score = 118 bits (295), Expect = 3e-27
Identities = 58/68 (85%), Positives = 63/68 (92%)
Frame = +1
Query: 538 SDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQT 597
+DDQIDILLAEVENLEKD GSAAS VDK+N IKDGVIC DEMRKII+DLF+DNVRLRKQT
Sbjct: 1 ADDQIDILLAEVENLEKDCGSAASYVDKSNGIKDGVICDDEMRKIIADLFIDNVRLRKQT 180
Query: 598 NRVTRHML 605
NR+TRH L
Sbjct: 181 NRITRHAL 204
>TC11195 UP|Q9FJH4 (Q9FJH4) Similarity to ring finger protein, partial (7%)
Length = 594
Score = 34.3 bits (77), Expect = 0.060
Identities = 30/111 (27%), Positives = 44/111 (39%)
Frame = +3
Query: 16 PLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKSKNSDPIVMSAFQDAS 75
PLP WSP+ + D E+PH+H + +N VM F
Sbjct: 117 PLPF---WSPSDFDFYSSDPEFPHHHHNH-----------HLSHRENQVNFVMDLFHQRV 254
Query: 76 QQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGT 126
+Q+S D + S+ S+ GL L G A D +D + A+ GT
Sbjct: 255 EQSSHHLTDDDPLPQSLTDSVFDGLGLDLG---FAPDDDTDDFFPAAGSGT 398
>AV765320
Length = 572
Score = 33.9 bits (76), Expect = 0.078
Identities = 22/68 (32%), Positives = 30/68 (43%), Gaps = 14/68 (20%)
Frame = +3
Query: 166 GLFMGQTILEQLEGL--------------PRHKVNARHGNNVTGKDRSNGNSYDSSLLGN 211
GL +GQ ++QL GL RHK+ R G + G D + +LLG
Sbjct: 207 GLAVGQLSVDQLSGLGTLVETHGVIRDGVDRHKLGLRVGGKLVGGDHIDRQDDLDTLLGR 386
Query: 212 NTMELFSQ 219
N +EL Q
Sbjct: 387 NLLELLGQ 410
>TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrate HASPP28
(Fragment), partial (61%)
Length = 754
Score = 32.0 bits (71), Expect = 0.30
Identities = 25/85 (29%), Positives = 42/85 (49%), Gaps = 7/85 (8%)
Frame = +3
Query: 451 KSLRSSQTELKKELSESIKEKYEAEKLLLHER-------EKSEQAEIAWRKQLEKCGLLL 503
KSL++ +L+K S +E+ E EK HER K+EQ+ +K LE+ L+
Sbjct: 321 KSLKARDVDLEKTTELSRREREEIEKQRAHERYMRLQEQGKTEQS----KKDLERLALIR 488
Query: 504 KQLQECSVELPYEDEDRTFLQSSSS 528
+Q E + + DE++ + S
Sbjct: 489 QQRAEAAKK---RDEEKAAKEQKKS 554
>TC18869 similar to UP|Q9SIR1 (Q9SIR1) At2g25320 protein, partial (8%)
Length = 381
Score = 31.2 bits (69), Expect = 0.50
Identities = 23/117 (19%), Positives = 50/117 (42%)
Frame = +1
Query: 414 LLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYE 473
L + + QLE L + + + K A+ KVL + + ++LK + +K+ +
Sbjct: 22 LTEQVQEIESQLEWLRSERDDEKVKLSAEKKVLQDRLHDADTQLSQLKSRKRDELKKVVK 201
Query: 474 AEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTD 530
+ L + +E A + ++L++ + + + EDE R Q+ T+
Sbjct: 202 EKNALAERLKNAEAARKRFDEELKR--FATENVTREEIRQSLEDEVRRLTQTVGQTE 366
>TC18268 weakly similar to UP|Q9C9S6 (Q9C9S6) Kinesin-related protein,
partial (5%)
Length = 617
Score = 30.0 bits (66), Expect = 1.1
Identities = 27/108 (25%), Positives = 51/108 (47%), Gaps = 5/108 (4%)
Frame = +2
Query: 371 QMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSK 430
Q+++ M+ L + E + KL+ + + +D + I + L QDLD A+ E
Sbjct: 104 QLKFTMQ-LEQTKFEEKKKLEEQDFSRLKKDKVRNEI--EISALKQDLDMAKRTYEEHVS 274
Query: 431 QYGELEAKSKADIKVLVKEVKSLRSSQTELKKEL-----SESIKEKYE 473
Q A+SKA+ + + E+K + + K+L S S+ K++
Sbjct: 275 QLELQAAESKAEYEKRIVELKHQLADARKQVKDLEAFSESRSLNRKHK 418
>TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete
Length = 1793
Score = 29.6 bits (65), Expect = 1.5
Identities = 52/238 (21%), Positives = 91/238 (37%), Gaps = 1/238 (0%)
Frame = +1
Query: 117 TPYDASEVGTPRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQ 176
T Y++ + + G DD+ + DED + +E VKYG N + Q
Sbjct: 709 TRYESHKESSQMEGDDDDDD--------DEDDVDDLENEVKYGQGNRMKARL-------Q 843
Query: 177 LEGLPRHKVNARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQP-GHAKVLGHIRKLSNE 235
E ++ H + + +NG + G N S P G +K + + + +
Sbjct: 844 WEEDADLSSSSGHDSQMQNPHLTNG----QLMSGENLCATTSGPMGQSKKVQSLPYVDPK 1011
Query: 236 SVGSDGSSIRGSDISNFGIPNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPL 295
G + SD +P G+ + SAS G +++ +G+ Q
Sbjct: 1012QPGLE------SDEEIRRVPEFGGESAAT---SASRPDTGSNPGPERVQGAGEGQKKRGR 1164
Query: 296 DQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQK 353
+K ++ L + R V+A+ E ++ A L TKVKDLE K++
Sbjct: 1165SSADKESKRLKRLLRNRVSAQQARE-------RKKAYLTDLETKVKDLETNNSELKER 1317
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/39 (35%), Positives = 17/39 (42%)
Frame = +3
Query: 4 RSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHR 42
+ P HRH +P L + P P RD HNHR
Sbjct: 222 KRPQHHRHSRQAPRVLHLQPLPHPHPPRRRDQPTDHNHR 338
>TC7840 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced response
protein, complete
Length = 1585
Score = 28.1 bits (61), Expect = 4.3
Identities = 60/268 (22%), Positives = 118/268 (43%), Gaps = 26/268 (9%)
Frame = +1
Query: 336 LTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESG 395
L+ +VK L+V ET K K+N+ ++ + L K+++ KL S++
Sbjct: 382 LSLRVKQLDVRCET---KTKDNVFVTVVASIQ--------YRALADKAVDAYYKL-SDTK 525
Query: 396 GNVGQ---DLTKESIVQQNDVLLQDLDAAREQLEILSKQYGE-----LEAKSKADIKVLV 447
+ D+ + S+ + +LD+A EQ ++K E + A ++ L+
Sbjct: 526 AQIQAYVFDVIRASVPKM------ELDSAFEQKNEIAKAVEEELEKAMSAYGYEIVQTLI 687
Query: 448 KEVKS---LRSSQTELKK--ELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLL 502
+++ ++ + E+ L + KEK EAEK+L +R + + A K L GL
Sbjct: 688 VDIEPDERVKKAMNEINAAARLRVATKEKAEAEKILQIKRAEGD----AESKYL--AGLG 849
Query: 503 LKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAE--VENLEKDAGSAA 560
+ + ++ V+ D F ++ T +S D +D++L + +++ SA
Sbjct: 850 IARQRQAIVD-GLRDSVLAFSENVPGT-------SSKDIMDMVLVTQYFDTMKEIGASAK 1005
Query: 561 SNV-----------DKTNDIKDGVICGD 577
SN D T+ I+DG++ G+
Sbjct: 1006SNAVFIPHGPGAVKDITSQIRDGLLQGN 1089
>TC13772
Length = 409
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/31 (38%), Positives = 20/31 (63%)
Frame = -1
Query: 338 TKVKDLEVELETSKQKNKENLQQAILIERER 368
+K KD+ +EL K++ + N Q+A +RER
Sbjct: 370 SKYKDIAIELAKKKKQTERNKQKATGAKRER 278
>AV424101
Length = 235
Score = 27.3 bits (59), Expect = 7.3
Identities = 19/77 (24%), Positives = 40/77 (51%)
Frame = +3
Query: 381 RKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSK 440
R S + + S+S + + + +ND+L Q++ + R Q++ L+ + LEA+
Sbjct: 3 RISCQSTSPVSSKSSPRQSYEDIHDDLKHRNDMLNQEVLSLRAQVDNLTSKSRSLEAELD 182
Query: 441 ADIKVLVKEVKSLRSSQ 457
+K L KEV ++ + +
Sbjct: 183 RTLKQL-KEVTAVAAEE 230
>TC11360 homologue to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%)
Length = 651
Score = 27.3 bits (59), Expect = 7.3
Identities = 7/16 (43%), Positives = 12/16 (74%)
Frame = +3
Query: 36 EWPHNHRTGWSYCVII 51
+W H ++ GWSYC ++
Sbjct: 18 QWYHTNKIGWSYCSLL 65
>TC14917 similar to GB|AAP37839.1|30725634|BT008480 At5g64750 {Arabidopsis
thaliana;}, partial (17%)
Length = 1051
Score = 26.9 bits (58), Expect = 9.5
Identities = 20/63 (31%), Positives = 29/63 (45%), Gaps = 1/63 (1%)
Frame = +3
Query: 60 SKNSDPIVMSAFQDASQQNSVKDPDSNNRDYSVQSSL-HSGLSLTAGSSSVASDYGSDTP 118
S +S+PIV + Q + Q S SN DYS S + +S T S + S S +
Sbjct: 441 SNSSEPIVHAEAQHHTFQGSSGSNSSNFYDYSPMSLYDQANVSSTMASHHLQSSSSSSSS 620
Query: 119 YDA 121
Y +
Sbjct: 621 YSS 629
>TC9492 homologue to UP|Q9MBA2 (Q9MBA2) MinD (Septum site-determining MinD)
(At5g24020), partial (33%)
Length = 633
Score = 26.9 bits (58), Expect = 9.5
Identities = 23/94 (24%), Positives = 42/94 (44%)
Frame = -2
Query: 422 REQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHE 481
++ I S + + K+K I++L E + +T L L+ +LLHE
Sbjct: 404 KQHKTITSLAHTKTNIKTKERIRILASE----K*EKTTLGLFLNHD-----SLHSILLHE 252
Query: 482 REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPY 515
+S ++ Q +CG L++ +E S+ PY
Sbjct: 251 PPRS-----LFKSQTSQCGRLVQNQREPSIGTPY 165
>TC7976 homologue to UP|METK_LYCES (P43280) S-adenosylmethionine synthetase
1 (Methionine adenosyltransferase 1) (AdoMet synthetase
1) , partial (67%)
Length = 862
Score = 26.9 bits (58), Expect = 9.5
Identities = 22/83 (26%), Positives = 35/83 (41%)
Frame = +2
Query: 503 LKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASN 562
L+ S LP++ TFL +S S + + K D D +L + D+ A
Sbjct: 17 LRHCSNFSAPLPFKMAAETFLFTSESVNEGHPDKLCDQISDAVLDACLEQDPDSKVACET 196
Query: 563 VDKTNDIKDGVICGDEMRKIISD 585
KTN + ++ G+ K I D
Sbjct: 197 CTKTNLV---MVFGEITTKAIVD 256
>BP040731
Length = 492
Score = 26.9 bits (58), Expect = 9.5
Identities = 15/34 (44%), Positives = 17/34 (49%)
Frame = -1
Query: 9 HRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHR 42
H TS LPLG SP+PR T H+HR
Sbjct: 462 HERTTTSTLPLGRSSSPSPRD----ATAGHHHHR 373
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.129 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,636,002
Number of Sequences: 28460
Number of extensions: 116329
Number of successful extensions: 505
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 503
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 505
length of query: 609
length of database: 4,897,600
effective HSP length: 96
effective length of query: 513
effective length of database: 2,165,440
effective search space: 1110870720
effective search space used: 1110870720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148405.4


