

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.2 + phase: 0
(209 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
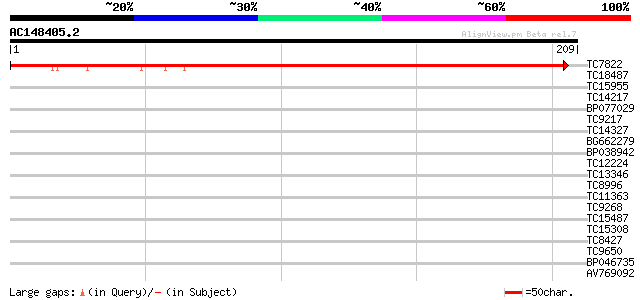
Score E
Sequences producing significant alignments: (bits) Value
TC7822 similar to UP|Q42139 (Q42139) H+-transporting ATP synthas... 272 3e-74
TC18487 similar to UP|O80953 (O80953) At2g39280 protein, partial... 36 0.006
TC15955 33 0.028
TC14217 28 0.89
BP077029 28 0.89
TC9217 similar to UP|Q9XQC7 (Q9XQC7) GrpE protein homolog, parti... 28 1.2
TC14327 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2G... 28 1.2
BG662279 28 1.2
BP038942 27 2.0
TC12224 similar to PIR|D96602|D96602 nucleolar protein [imported... 27 3.4
TC13346 26 4.4
TC8996 homologue to UP|BAD10669 (BAD10669) Lipoxygenase-like pro... 26 5.8
TC11363 similar to UP|Q9SWV4 (Q9SWV4) ER66 protein (Fragment), p... 26 5.8
TC9268 similar to UP|Q9LHE6 (Q9LHE6) Genomic DNA, chromosome 3, ... 26 5.8
TC15487 similar to UP|O23144 (O23144) Proton pump interactor (AT... 25 7.5
TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%) 25 7.5
TC8427 homologue to UP|Q84L15 (Q84L15) Cytosolic glutamine synth... 25 7.5
TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogen... 25 7.5
BP046735 25 7.5
AV769092 25 9.8
>TC7822 similar to UP|Q42139 (Q42139) H+-transporting ATP synthase CHAIN9 -
like protein (AT4G32260/F10M6_100) , partial (76%)
Length = 907
Score = 272 bits (696), Expect = 3e-74
Identities = 159/215 (73%), Positives = 180/215 (82%), Gaps = 9/215 (4%)
Frame = +3
Query: 1 MASMIMASSKPIIR-VP-TNSLPSTPKLP-SLQTITPNLKLQLSKLKSMT---LAATSLS 54
MA+M+MASSKP+I +P +NSLP TP+L SL + LK Q+ LKS LAA+SLS
Sbjct: 102 MANMVMASSKPLIPPLPISNSLPPTPRLQISLPKLPLKLKHQIPNLKSSLSSLLAASSLS 281
Query: 55 FV-FAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRG 111
+ FAPPSLA EKAALFDFNLTLPIIV EFLFLM ALDK+YF+PLG FMD+RDA IR
Sbjct: 282 LLAFAPPSLAEEIEKAALFDFNLTLPIIVAEFLFLMVALDKIYFSPLGKFMDDRDAAIRE 461
Query: 112 KLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAEL 171
KLNSV +TS EVK+LEEQANAVL+AARAEI+ ALN+MKKETQ EVE+KIAEGR KVDAEL
Sbjct: 462 KLNSVKDTSSEVKQLEEQANAVLKAARAEISAALNKMKKETQVEVEQKIAEGRLKVDAEL 641
Query: 172 QEALANLEKQKEETVKALDSQIAALSQDIVNKVLP 206
QEALANLEKQKEET+K+LDSQIAALSQDIV KVLP
Sbjct: 642 QEALANLEKQKEETIKSLDSQIAALSQDIVKKVLP 746
>TC18487 similar to UP|O80953 (O80953) At2g39280 protein, partial (13%)
Length = 592
Score = 35.8 bits (81), Expect = 0.006
Identities = 22/87 (25%), Positives = 53/87 (60%), Gaps = 8/87 (9%)
Frame = +1
Query: 111 GKLNSVTNTSEEV--------KELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAE 162
G ++SV + E+V + LEE+ +A+LRA E A+ + +K++ + ++ K+ +
Sbjct: 259 GDIDSVPDLHEQVVGLKVELCRLLEEKRSAILRAEELETAL-MEMVKQDNRRQLSAKVEQ 435
Query: 163 GRKKVDAELQEALANLEKQKEETVKAL 189
+++ A+L++AL++ ++Q+ ++ L
Sbjct: 436 LEEEI-AQLRQALSDKQEQETAMIQVL 513
>TC15955
Length = 1042
Score = 33.5 bits (75), Expect = 0.028
Identities = 30/104 (28%), Positives = 51/104 (48%), Gaps = 8/104 (7%)
Frame = +1
Query: 98 LGTFMDNRDAEIRGKLNSVTN----TSEEVKELEEQAN--AVLRAARAEIAVALNQMKKE 151
L +F D GKL + ++E LE + N A +++ E+ +N MKKE
Sbjct: 265 LNSFADEIKKNSMGKLEESISLRQKSNENAARLEGKRNQLAAMQSYIDEMEAQMNIMKKE 444
Query: 152 TQAEVEEKIAEGRKKV-DAELQE-ALANLEKQKEETVKALDSQI 193
TQ AE +K + D +L + L N+E++ E +KA + ++
Sbjct: 445 TQDYTGRCSAEAQKMLEDVQLADHDLDNMEREVAEVLKASELKL 576
>TC14217
Length = 985
Score = 28.5 bits (62), Expect = 0.89
Identities = 22/77 (28%), Positives = 36/77 (46%), Gaps = 7/77 (9%)
Frame = +1
Query: 122 EVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVD-------AELQEA 174
E K++EEQ A E +V + +K +TQA ++K KVD ++E
Sbjct: 127 ETKKVEEQNEAEAATTETE-SVKVENVKNDTQAVDDKKEENADTKVDEISTAISEPVRET 303
Query: 175 LANLEKQKEETVKALDS 191
LA+ K +E +D+
Sbjct: 304 LASKFKDEESIETGVDN 354
>BP077029
Length = 486
Score = 28.5 bits (62), Expect = 0.89
Identities = 23/74 (31%), Positives = 40/74 (53%), Gaps = 3/74 (4%)
Frame = -3
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDA---ELQEALAN 177
++ KELEEQ NA+ +A ++V Q K T + + ++ E K + A EL+E + +
Sbjct: 484 QKKKELEEQINAI----KANLSVF--QTAKVTAIKRKREVFEEGKALKAQRDELREQVPH 323
Query: 178 LEKQKEETVKALDS 191
L ++E K D+
Sbjct: 322 LRDEREVAKKIQDN 281
>TC9217 similar to UP|Q9XQC7 (Q9XQC7) GrpE protein homolog, partial (48%)
Length = 993
Score = 28.1 bits (61), Expect = 1.2
Identities = 26/90 (28%), Positives = 39/90 (42%), Gaps = 1/90 (1%)
Frame = +1
Query: 98 LGTFMDN-RDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEV 156
+G DN R R +L+ V+N EV E A+A+I V + E
Sbjct: 58 IGADFDNFRKRTERDRLSLVSNAQGEVVESLLPVLDNFERAKAQIKV---------ETEA 210
Query: 157 EEKIAEGRKKVDAELQEALANLEKQKEETV 186
EEKI+ + + + E L +L + ETV
Sbjct: 211 EEKISNSYQSIYKQFIEILTSLGVEPVETV 300
>TC14327 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2G14_40
{Arabidopsis thaliana;}, partial (22%)
Length = 1038
Score = 28.1 bits (61), Expect = 1.2
Identities = 14/42 (33%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = +3
Query: 11 PIIRVPTNSLPSTPKLPSLQTITP--NLKLQLSKLKSMTLAA 50
P+++ P +PSTP + S++ P N + QL K+M A
Sbjct: 129 PVVKPPVAPIPSTPIVKSMKDCIPLCNYRCQLHSRKNMCTRA 254
>BG662279
Length = 337
Score = 28.1 bits (61), Expect = 1.2
Identities = 18/48 (37%), Positives = 22/48 (45%)
Frame = +1
Query: 16 PTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPSLA 63
PT+ PS+P LP T P S + TSLS + P SLA
Sbjct: 184 PTSKSPSSPSLPKPPTPKPRSSDHPSATSLIYQPRTSLSSLEPPSSLA 327
>BP038942
Length = 617
Score = 27.3 bits (59), Expect = 2.0
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +1
Query: 13 IRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPSL 62
+ P + P T P LQ TP+ +LS S + TSLS + P+L
Sbjct: 382 LATPLSPSPPTSLFPFLQPPTPSRLSRLSSFTSNPSSVTSLSTKPSSPTL 531
>TC12224 similar to PIR|D96602|D96602 nucleolar protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(16%)
Length = 315
Score = 26.6 bits (57), Expect = 3.4
Identities = 20/55 (36%), Positives = 27/55 (48%), Gaps = 4/55 (7%)
Frame = -1
Query: 111 GKLNSVTNTSEEVKELEEQANAVLRAARAEI----AVALNQMKKETQAEVEEKIA 161
GKL S T + V E A LRA+RAE+ +L + K Q+E E + A
Sbjct: 288 GKLVSSTVLNSSVNNPSETALHCLRASRAEVKGLKERSLTTLPKRFQSETELRTA 124
>TC13346
Length = 458
Score = 26.2 bits (56), Expect = 4.4
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -1
Query: 83 FLFLMFALDKLYFTPLGTFMDN 104
FL+L F DKLYF LG D+
Sbjct: 191 FLYLTFRHDKLYFISLGKL*DH 126
>TC8996 homologue to UP|BAD10669 (BAD10669) Lipoxygenase-like protein,
partial (5%)
Length = 562
Score = 25.8 bits (55), Expect = 5.8
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = -3
Query: 20 LPSTPKLPSLQTITPNLKLQLSKLKSMTLAAT 51
+ S+PKL L+ +T LK+++ K++ AAT
Sbjct: 371 ISSSPKLALLKPLTQKLKIEMRKVEGNGGAAT 276
>TC11363 similar to UP|Q9SWV4 (Q9SWV4) ER66 protein (Fragment), partial
(18%)
Length = 557
Score = 25.8 bits (55), Expect = 5.8
Identities = 16/47 (34%), Positives = 27/47 (57%), Gaps = 1/47 (2%)
Frame = +1
Query: 134 LRAARAEIAVALNQMKKETQAEVE-EKIAEGRKKVDAELQEALANLE 179
LR ++E N ++ E+ E E + + EGRK+ + LQ ALA ++
Sbjct: 115 LRGFKSEGVSEGNMVQGESSTEDEYDLLKEGRKQTEQRLQIALARVK 255
>TC9268 similar to UP|Q9LHE6 (Q9LHE6) Genomic DNA, chromosome 3, P1 clone:
MZE19, partial (30%)
Length = 805
Score = 25.8 bits (55), Expect = 5.8
Identities = 17/50 (34%), Positives = 25/50 (50%)
Frame = +3
Query: 8 SSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVF 57
S KP+ P+ + STP P+ + +P+ LQL S +AT S F
Sbjct: 369 SGKPLAPSPS-AASSTPLTPTPTSTSPSSTLQLPATSSAATSATKPSAWF 515
>TC15487 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (16%)
Length = 1153
Score = 25.4 bits (54), Expect = 7.5
Identities = 17/48 (35%), Positives = 26/48 (53%)
Frame = +2
Query: 145 LNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQ 192
L +MK+E + ++ E +KK+ AE A A L+ QKE K D +
Sbjct: 440 LKEMKREEEIAKAKQAWERKKKL-AEKAAAKAALKAQKEAEKKLKDRE 580
>TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%)
Length = 1103
Score = 25.4 bits (54), Expect = 7.5
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +1
Query: 5 IMASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKS 45
+MA I+ V N LP KLP LQT+ + ++ SK ++
Sbjct: 301 VMAYMAQILLVLLNILPPR-KLPPLQTLKMSFSMRTSKTRA 420
>TC8427 homologue to UP|Q84L15 (Q84L15) Cytosolic glutamine synthetase,
partial (41%)
Length = 683
Score = 25.4 bits (54), Expect = 7.5
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 2/42 (4%)
Frame = +1
Query: 20 LPSTPKLPSLQTITPNLKLQ--LSKLKSMTLAATSLSFVFAP 59
L STP L QT+ P L+L+ K+ +TL L + P
Sbjct: 268 LTSTPSLGVWQTVEPQLELEEKQRKMGKVTLRTEGLPLTWIP 393
>TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogenase-like
protein , partial (47%)
Length = 1041
Score = 25.4 bits (54), Expect = 7.5
Identities = 11/33 (33%), Positives = 21/33 (63%)
Frame = -2
Query: 6 MASSKPIIRVPTNSLPSTPKLPSLQTITPNLKL 38
+A+++ + +PT +LP+T LP+ + T L L
Sbjct: 440 VAAARSTLSIPTPALPTTLSLPAEDSNTSRLTL 342
>BP046735
Length = 535
Score = 25.4 bits (54), Expect = 7.5
Identities = 15/59 (25%), Positives = 28/59 (47%)
Frame = +2
Query: 23 TPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPSLAFEKAALFDFNLTLPIIVV 81
TP + P L+ L + + A + SF+ + PSL F L +++T +I++
Sbjct: 119 TPSKAFYHLVNPGGSGSLTLLSTPSRATSKASFIPSNPSLLFA*TNLGRYSITFSLILL 295
>AV769092
Length = 427
Score = 25.0 bits (53), Expect = 9.8
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +2
Query: 15 VPTNSLPSTPKLPSLQTITPNLKLQL 40
+P SLP P LPS ++ P L L +
Sbjct: 272 IPN*SLPQFPLLPS*HSLIPTLPLHI 349
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.128 0.329
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,247,175
Number of Sequences: 28460
Number of extensions: 24137
Number of successful extensions: 204
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 201
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 202
length of query: 209
length of database: 4,897,600
effective HSP length: 86
effective length of query: 123
effective length of database: 2,450,040
effective search space: 301354920
effective search space used: 301354920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148405.2


