

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.1 + phase: 0
(149 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
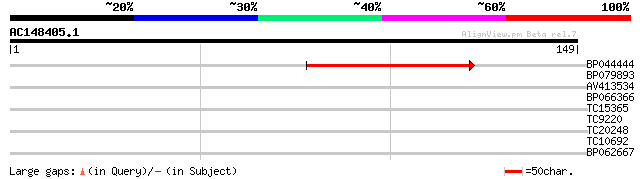
Score E
Sequences producing significant alignments: (bits) Value
BP044444 44 1e-05
BP079893 26 2.5
AV413534 25 7.3
BP066366 25 7.3
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 25 7.3
TC9220 similar to UP|Q8W1N7 (Q8W1N7) CSLC7 (Fragment), partial (... 24 9.5
TC20248 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (8%) 24 9.5
TC10692 weakly similar to UP|BAD08916 (BAD08916) Calcium-binding... 24 9.5
BP062667 24 9.5
>BP044444
Length = 482
Score = 43.9 bits (102), Expect = 1e-05
Identities = 16/44 (36%), Positives = 29/44 (65%)
Frame = -1
Query: 79 KISWPLVWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHV 122
+ISW VW + CL Y KL+ + D L+++G+++ Q+ F+ +V
Sbjct: 419 QISWRGVWAKCCLSYNNHKLLNDDDALQNFGVRNNSQVHFVPYV 288
>BP079893
Length = 425
Score = 26.2 bits (56), Expect = 2.5
Identities = 5/15 (33%), Positives = 11/15 (73%)
Frame = -1
Query: 78 EKISWPLVWGQFCLC 92
+++ W +WG +C+C
Sbjct: 203 DRLKWLYIWGHYCIC 159
>AV413534
Length = 416
Score = 24.6 bits (52), Expect = 7.3
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = -2
Query: 44 VLKLDSSSFQVEVAKTATVAV 64
V KLD +F E+A+ ATVAV
Sbjct: 271 VSKLDLKAFSCELAEEATVAV 209
>BP066366
Length = 448
Score = 24.6 bits (52), Expect = 7.3
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -2
Query: 15 SLSIQLSSMIIVDCTLPYDKLPSQNLTL 42
SL Q SSM +V +PY +P + L
Sbjct: 252 SLGEQFSSMALVSVAVPYSSVPQM*IAL 169
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 24.6 bits (52), Expect = 7.3
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 4/35 (11%)
Frame = +3
Query: 118 FIRHVSNVCNFQRKPKKRT----VNLKQHGRYEFS 148
F+ HVS +C K +KRT V++ H + F+
Sbjct: 1107 FVFHVSFLCTLSCKDEKRTHLRVVSIYSHAGFHFA 1211
>TC9220 similar to UP|Q8W1N7 (Q8W1N7) CSLC7 (Fragment), partial (39%)
Length = 885
Score = 24.3 bits (51), Expect = 9.5
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +3
Query: 106 RDYGIKDGDQLRFIRHVSNVCNFQRKPKKR 135
+D IK+G +R++ NF+RK +R
Sbjct: 75 KDQSIKEGSLCLILRNLKKKSNFRRKRHQR 164
>TC20248 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (8%)
Length = 496
Score = 24.3 bits (51), Expect = 9.5
Identities = 15/61 (24%), Positives = 28/61 (45%), Gaps = 6/61 (9%)
Frame = -1
Query: 12 PRRSLSIQLSSM------IIVDCTLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVL 65
PR S ++++ S + + C LP K+P Q+L+ + S + AT + L
Sbjct: 226 PRSSEALEVFSRKMECRTVALVCVLPITKMPEQSLSSLCTRRGGSEYSYFKHDAATTSKL 47
Query: 66 K 66
+
Sbjct: 46 R 44
>TC10692 weakly similar to UP|BAD08916 (BAD08916) Calcium-binding EF-hand
family protein-like, partial (32%)
Length = 634
Score = 24.3 bits (51), Expect = 9.5
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +3
Query: 2 SELTSVSNESPRRSLSIQLSSMIIVDCTL 30
S LTS S PRRS S + +++ ++ T+
Sbjct: 192 SRLTSASTSQPRRSSSPRSTTLCLIRLTV 278
>BP062667
Length = 528
Score = 24.3 bits (51), Expect = 9.5
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = +1
Query: 35 LPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAA 72
+P QN +LKL SSS + A VL+ E+A
Sbjct: 121 IPCQNPKTNMLKLISSSHIHRLTPAAAAPVLRPIAESA 234
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,576,023
Number of Sequences: 28460
Number of extensions: 32308
Number of successful extensions: 193
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 193
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 193
length of query: 149
length of database: 4,897,600
effective HSP length: 82
effective length of query: 67
effective length of database: 2,563,880
effective search space: 171779960
effective search space used: 171779960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148405.1


