

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148404.10 + phase: 0
(400 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
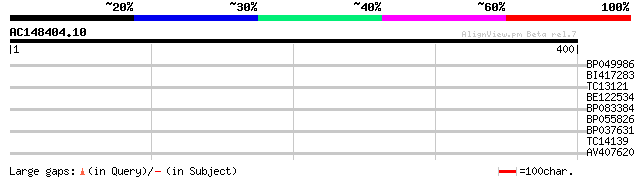
Score E
Sequences producing significant alignments: (bits) Value
BP049986 28 2.7
BI417283 28 2.7
TC13121 similar to UP|SYFB_ARATH (Q9SGE9) Probable phenylalanyl-... 27 4.6
BE122534 27 4.6
BP083384 27 6.0
BP055826 27 7.9
BP037631 27 7.9
TC14139 27 7.9
AV407620 27 7.9
>BP049986
Length = 608
Score = 28.1 bits (61), Expect = 2.7
Identities = 12/31 (38%), Positives = 21/31 (67%)
Frame = +3
Query: 6 SLRLHQLPFFIIFFLQLFTAFSNASSSSSST 36
SLR+H + + F + T+ S++SS+SSS+
Sbjct: 387 SLRIHTIAALVFFIIAKLTSSSSSSSTSSSS 479
>BI417283
Length = 557
Score = 28.1 bits (61), Expect = 2.7
Identities = 13/24 (54%), Positives = 18/24 (74%)
Frame = +3
Query: 16 IIFFLQLFTAFSNASSSSSSTHSN 39
++ FL L A S++SSSSSS+H N
Sbjct: 69 LLLFLLLLVAPSSSSSSSSSSHEN 140
>TC13121 similar to UP|SYFB_ARATH (Q9SGE9) Probable phenylalanyl-tRNA
synthetase beta chain (Phenylalanine--tRNA ligase beta
chain) (PheRS) , partial (37%)
Length = 721
Score = 27.3 bits (59), Expect = 4.6
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +3
Query: 214 PLIPIRIIGSASLIAYVARNPYVQIG 239
P +P R S S +V R+PY+QIG
Sbjct: 276 PALP*RTCSSPSSFLWVPRDPYLQIG 353
>BE122534
Length = 358
Score = 27.3 bits (59), Expect = 4.6
Identities = 16/29 (55%), Positives = 18/29 (61%), Gaps = 1/29 (3%)
Frame = -2
Query: 12 LPFFIIFFLQLFTAFSNASS-SSSSTHSN 39
LPFF I LQLF F ++SS S S H N
Sbjct: 351 LPFFEIASLQLFNIFMHSSSFQSCSIHFN 265
>BP083384
Length = 362
Score = 26.9 bits (58), Expect = 6.0
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = -1
Query: 198 SNQKNEFWPFLQSMCVPLIPIRII 221
S Q FWPFL S+ + +IP+ II
Sbjct: 80 SLQLKGFWPFLLSL*IKIIPLIII 9
>BP055826
Length = 544
Score = 26.6 bits (57), Expect = 7.9
Identities = 15/37 (40%), Positives = 19/37 (50%), Gaps = 1/37 (2%)
Frame = +3
Query: 86 SSKNNFT-VKFSDEISSWNHNKFTTTPKPDLASLVDQ 121
S N FT + +D I SWN K+ P P AS + Q
Sbjct: 45 SLHNYFTKI*LNDLIYSWNQYKYN*NPAPTSASAIQQ 155
>BP037631
Length = 529
Score = 26.6 bits (57), Expect = 7.9
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 8/51 (15%)
Frame = -2
Query: 354 KPLSIKKVKPFIESDSVSWANLMSNLSYTKL--------RPVLLPPEALTL 396
K + I V+PF + ++W L S LSY+ R L+ PE +T+
Sbjct: 459 KSIQIVPVQPFATNS*INWTCLNSCLSYSNCSL*CSMVSRSTLVTPEHITV 307
>TC14139
Length = 482
Score = 26.6 bits (57), Expect = 7.9
Identities = 10/38 (26%), Positives = 21/38 (54%)
Frame = -1
Query: 3 TMISLRLHQLPFFIIFFLQLFTAFSNASSSSSSTHSNI 40
+M+ +HQ PF + F +F + AS++ + ++ I
Sbjct: 146 SMVHYNIHQFPFIVSFTSSIFIKKNTASANKQTIYTPI 33
>AV407620
Length = 414
Score = 26.6 bits (57), Expect = 7.9
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +3
Query: 9 LHQLPFFIIFFLQLFTAFSNASSSSSSTHSNITHI 43
LH + I LFTAF +SS SS+H+ I I
Sbjct: 15 LHGIHLLIFIISTLFTAFIISSSPPSSSHT*IQAI 119
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,243,620
Number of Sequences: 28460
Number of extensions: 102643
Number of successful extensions: 700
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 675
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 693
length of query: 400
length of database: 4,897,600
effective HSP length: 92
effective length of query: 308
effective length of database: 2,279,280
effective search space: 702018240
effective search space used: 702018240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148404.10


