

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
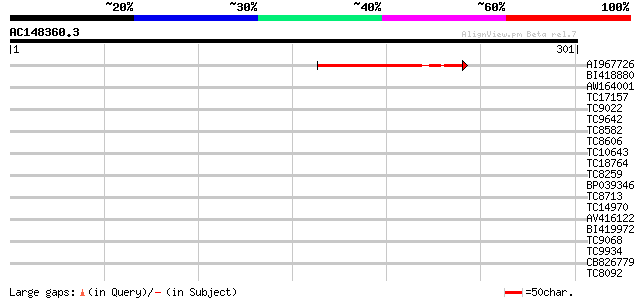
Query= AC148360.3 - phase: 0
(301 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AI967726 134 1e-32
BI418880 37 0.005
AW164001 36 0.007
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 34 0.026
TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Ara... 34 0.026
TC9642 33 0.045
TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 33 0.059
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 33 0.059
TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear phospho... 33 0.077
TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding prote... 32 0.10
TC8259 similar to UP|HMGL_SOYBN (P26585) HMG1/2-like protein (SB... 32 0.13
BP039346 32 0.17
TC8713 similar to UP|P93704 (P93704) HMG-1, partial (91%) 32 0.17
TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) prec... 27 0.28
AV416122 30 0.38
BI419972 30 0.38
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 30 0.38
TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protei... 30 0.38
CB826779 30 0.50
TC8092 similar to UP|Q8MLI6 (Q8MLI6) CG30127-PA, partial (3%) 30 0.50
>AI967726
Length = 247
Score = 134 bits (338), Expect = 1e-32
Identities = 69/81 (85%), Positives = 73/81 (89%), Gaps = 1/81 (1%)
Frame = +3
Query: 164 KPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMK-KNFGAENE 222
KPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLM KNFG
Sbjct: 15 KPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMNMKNFGG--- 185
Query: 223 DDDEDEDEEEDEADFPSKLGK 243
D+DEDED ++D+A FPSKLGK
Sbjct: 186 DEDEDED-DDDQASFPSKLGK 245
>BI418880
Length = 606
Score = 36.6 bits (83), Expect = 0.005
Identities = 25/145 (17%), Positives = 66/145 (45%), Gaps = 22/145 (15%)
Frame = +2
Query: 40 KCQVCEILAKQLYQQVQSKK------------------AEISPKKISEYQIIEIAENVCN 81
KC C+ +A++L + +K ++ ++S+ +++E+ + +C
Sbjct: 92 KCAACKAVAEELEIGLSREKPRNHLDMRHRLDSKGQRQGKLIDYRVSDLRVVELLDGLCE 271
Query: 82 LKKVEADWILRIDIVEKADRLELEEHDS----EGQCNSECKTVERACQEVMGYSDTDVAE 137
+ D+ L ID + + +++ D+ + + + K + C ++ ++ ++AE
Sbjct: 272 KMQ---DYTLMID-TDTYEWFKVDSWDNLTKNKQEARAYSKDISSYCGRLLEDTEEELAE 439
Query: 138 YLYSSKPDIDSLTNYLCKDLSKACN 162
+ + ++ LC+DLSK C+
Sbjct: 440 LIKKGSVKVGDVSKVLCQDLSKHCS 514
>AW164001
Length = 447
Score = 36.2 bits (82), Expect = 0.007
Identities = 15/40 (37%), Positives = 28/40 (69%)
Frame = +1
Query: 210 DDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKE 249
D++ K+ E+ DDD+D+D+++D+ D P+ LG + K K+
Sbjct: 52 DEVYKEEDVDEDFDDDDDDDDDDDDDDEPATLGFVDKPKD 171
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 34.3 bits (77), Expect = 0.026
Identities = 16/45 (35%), Positives = 28/45 (61%)
Frame = +2
Query: 218 GAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKG 262
G E+ED+D+D+DEEE++ D +++E+G+ K K + G
Sbjct: 791 GDEDEDEDDDDDEEEEDDDD--------DDEDDEEGEGKSKSKSG 901
>TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (75%)
Length = 1083
Score = 34.3 bits (77), Expect = 0.026
Identities = 23/101 (22%), Positives = 51/101 (49%)
Frame = +3
Query: 164 KPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENED 223
+P V ++ PG P V SS+ +E++L + E +++ + ++
Sbjct: 75 QPSHVSQNMAPG-PVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDE 251
Query: 224 DDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIV 264
D+DEDE++D+ D + G + S+ +++ ++K RK ++
Sbjct: 252 ADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAML 374
>TC9642
Length = 657
Score = 33.5 bits (75), Expect = 0.045
Identities = 18/29 (62%), Positives = 23/29 (79%), Gaps = 1/29 (3%)
Frame = -3
Query: 215 KNFGAENE-DDDEDEDEEEDEADFPSKLG 242
K+ GA N+ DDDEDEDE+EDE D S++G
Sbjct: 283 KSTGAANDTDDDEDEDEDEDE-DEASEIG 200
>TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (65%)
Length = 1341
Score = 33.1 bits (74), Expect = 0.059
Identities = 41/167 (24%), Positives = 74/167 (43%)
Frame = +1
Query: 83 KKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSS 142
KK + R DI + L L++ SE NS+ +V +V+ D D +++ S
Sbjct: 223 KKYVLGTLNRDDIPHLSMDLVLDDK-SELSHNSKGASVYFCGYKVLT-DDGDASDFSESD 396
Query: 143 KPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAP 202
+ + + + K +K + K K P AK K + +K + + +P
Sbjct: 397 EEEEELMEQENGKPETKTEDAKVAKTAKPAAAAGPS-AKQVKIVDPKKDEEEADLDVSSP 573
Query: 203 GMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKE 249
+ Y DDL+ A+ + DDE +DE +DE + P+ K+ + K+
Sbjct: 574 DVSGYE-DDLIS----ADEDSDDESDDESDDEEETPTPAKKVDQGKK 699
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 33.1 bits (74), Expect = 0.059
Identities = 22/64 (34%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Frame = -2
Query: 193 KSMEGMPGA-PGMKMYSRDDLMKKNFGAENEDDD-EDEDEEEDEADFPSKLGKILKSKEN 250
K+ EG PGA G + D+ + E++DDD ED+DEE+D + + G + E+
Sbjct: 482 KAPEGGPGAGAGPEENGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEEDNED 303
Query: 251 EKGD 254
E+ D
Sbjct: 302 EEED 291
Score = 30.0 bits (66), Expect = 0.50
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = -2
Query: 215 KNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
K AE ED ++DE +E+D+ D P G+ S E+ G
Sbjct: 635 KQHDAEGEDGNDDEGDEDDDGDGPFGEGEEDLSSEDGAG 519
Score = 29.3 bits (64), Expect = 0.85
Identities = 15/27 (55%), Positives = 19/27 (69%), Gaps = 1/27 (3%)
Frame = -2
Query: 218 GAENED-DDEDEDEEEDEADFPSKLGK 243
G E ED +DE+EDEE++EA P K K
Sbjct: 329 GVEEEDNEDEEEDEEDEEALQPPKKRK 249
>TC10643 similar to UP|BAC84847 (BAC84847) Acidic nuclear
phosphoprotein-like protein, partial (5%)
Length = 789
Score = 32.7 bits (73), Expect = 0.077
Identities = 17/53 (32%), Positives = 29/53 (54%), Gaps = 6/53 (11%)
Frame = +2
Query: 212 LMKKNFGAENEDDDEDE------DEEEDEADFPSKLGKILKSKENEKGDWKQK 258
+MK G E+ED++E+E D++EDE+ G ++ EN+ G +K
Sbjct: 17 MMKLGSGEEDEDEEEEEDSEESSDDDEDESAEGGSRGTLVDEDENDNGRSSRK 175
>TC18764 similar to UP|Q8LBJ5 (Q8LBJ5) Nucleic acid binding protein-like,
partial (35%)
Length = 848
Score = 32.3 bits (72), Expect = 0.10
Identities = 19/56 (33%), Positives = 25/56 (43%)
Frame = +1
Query: 181 KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD 236
K KE+ KS P YS+ + K E ED+DE+ EEEDE +
Sbjct: 73 KQGKESHSSNKPKSNSKRGSEPQXVKYSKGLISKDEDEGEGEDEDEEGLEEEDEEE 240
>TC8259 similar to UP|HMGL_SOYBN (P26585) HMG1/2-like protein (SB11
protein), partial (92%)
Length = 942
Score = 32.0 bits (71), Expect = 0.13
Identities = 19/61 (31%), Positives = 35/61 (57%), Gaps = 1/61 (1%)
Frame = +2
Query: 177 PFVAKSSK-EAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEA 235
P+VAK+ K +A+ EK +K+ A G ++ +K+ N+ D++D+D +ED+
Sbjct: 452 PYVAKAEKRKADYEKTMKAYNKKQ-AEGPAAADEEE-SEKSVSEVNDQDEDDDDSDEDDD 625
Query: 236 D 236
D
Sbjct: 626 D 628
>BP039346
Length = 470
Score = 31.6 bits (70), Expect = 0.17
Identities = 12/17 (70%), Positives = 15/17 (87%)
Frame = +1
Query: 217 FGAENEDDDEDEDEEED 233
FGA E++DEDEDE+ED
Sbjct: 349 FGAAEEEEDEDEDEDED 399
>TC8713 similar to UP|P93704 (P93704) HMG-1, partial (91%)
Length = 800
Score = 31.6 bits (70), Expect = 0.17
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 2/55 (3%)
Frame = +2
Query: 177 PFVAKSSK-EAEMEKLLKSME-GMPGAPGMKMYSRDDLMKKNFGAENEDDDEDED 229
P+VA++ K + E EK L++ G+ + G D K ++EDDDED++
Sbjct: 386 PYVARAEKAKEEYEKTLRAYNNGLAESKGSAEEEGSDKSKSEVNDDDEDDDEDDE 550
>TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) precursor,
partial (51%)
Length = 1566
Score = 26.9 bits (58), Expect(2) = 0.28
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +1
Query: 207 YSRDDLMKKNFGAENEDDDEDEDEEE 232
+SR + + ED+DEDEDEEE
Sbjct: 559 HSRPSGRRTRRPSREEDEDEDEDEEE 636
Score = 26.2 bits (56), Expect = 7.2
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = +1
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEK 252
E+ED+DEDE+E + P + + + +E E+
Sbjct: 604 EDEDEDEDEEERQGGRRGPGERQRHFEREEEEE 702
Score = 22.3 bits (46), Expect(2) = 0.28
Identities = 13/49 (26%), Positives = 21/49 (42%)
Frame = +1
Query: 225 DEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKH 273
+E+E+EEE P S+ +G + G+ + TLK H
Sbjct: 748 EEEEEEEESRRTIP--------SRRESRGRGGCRTSNGLEENFCTLKIH 870
>AV416122
Length = 414
Score = 30.4 bits (67), Expect = 0.38
Identities = 11/19 (57%), Positives = 16/19 (83%)
Frame = +3
Query: 218 GAENEDDDEDEDEEEDEAD 236
G E EDD+ED++EE++E D
Sbjct: 300 GEEEEDDEEDKEEEDEEED 356
Score = 28.9 bits (63), Expect = 1.1
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +3
Query: 217 FGAENEDDDEDEDEEEDEAD 236
F E E+DDE++ EEEDE +
Sbjct: 294 FEGEEEEDDEEDKEEEDEEE 353
>BI419972
Length = 498
Score = 30.4 bits (67), Expect = 0.38
Identities = 16/54 (29%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Frame = +1
Query: 207 YSRDDLMKK------NFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
+S +DL+ + N +E EDD++ +DE++D D G+ E E+ D
Sbjct: 292 FSLEDLLGRKPAYLANKDSETEDDEDGDDEDDDADDDDDDEGEDYSGDEGEEAD 453
Score = 29.6 bits (65), Expect = 0.65
Identities = 12/50 (24%), Positives = 28/50 (56%)
Frame = +1
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGK 243
S+E + G + ++D + + ++EDDD D+D++++ D+ G+
Sbjct: 295 SLEDLLGRKPAYLANKDSETEDDEDGDDEDDDADDDDDDEGEDYSGDEGE 444
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 30.4 bits (67), Expect = 0.38
Identities = 18/58 (31%), Positives = 30/58 (51%)
Frame = +2
Query: 214 KKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLK 271
K+ F + +DDD+D+D+++D+ D +++ GD KQ DT T LK
Sbjct: 59 KRLFSPKPDDDDDDDDDDDDDDDDDD-------DDDDDDGDKKQPDS----DTGTALK 199
>TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (37%)
Length = 1050
Score = 30.4 bits (67), Expect = 0.38
Identities = 13/28 (46%), Positives = 21/28 (74%)
Frame = +2
Query: 207 YSRDDLMKKNFGAENEDDDEDEDEEEDE 234
+S DDL+ + G E +D +E+E+EEE+E
Sbjct: 38 FSVDDLLDFSNGKEMDDYEEEEEEEEEE 121
>CB826779
Length = 469
Score = 30.0 bits (66), Expect = 0.50
Identities = 22/91 (24%), Positives = 40/91 (43%)
Frame = +1
Query: 173 TPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEE 232
+PG F A S ++ + P + Y +++ +ED+D+DED+E+
Sbjct: 94 SPGTVFDASSDDHQQLPSATDDKK----KPQPQEYDDAPVVEDVHDEADEDEDDDEDDED 261
Query: 233 DEADFPSKLGKILKSKENEKGDWKQKIRKGI 263
DE D + K +EK K ++ G+
Sbjct: 262 DE-DGAQGTTEGSKQSRSEKKSRKAMLKLGL 351
Score = 26.2 bits (56), Expect = 7.2
Identities = 13/45 (28%), Positives = 27/45 (59%)
Frame = +1
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIV 264
E+ D+ DEDE++DE D + G ++ +++ ++K RK ++
Sbjct: 205 EDVHDEADEDEDDDEDDEDDEDGAQGTTEGSKQSRSEKKSRKAML 339
>TC8092 similar to UP|Q8MLI6 (Q8MLI6) CG30127-PA, partial (3%)
Length = 1073
Score = 30.0 bits (66), Expect = 0.50
Identities = 14/35 (40%), Positives = 24/35 (68%), Gaps = 5/35 (14%)
Frame = -1
Query: 211 DLMKKNFGAENEDDD-----EDEDEEEDEADFPSK 240
D + + FG ENE +D E+++E+E+EAD P++
Sbjct: 404 D*VVRRFGTENEVEDAGGDAEEDEEDEEEADEPAE 300
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,421,656
Number of Sequences: 28460
Number of extensions: 73903
Number of successful extensions: 1133
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 604
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 850
length of query: 301
length of database: 4,897,600
effective HSP length: 90
effective length of query: 211
effective length of database: 2,336,200
effective search space: 492938200
effective search space used: 492938200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148360.3


