

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
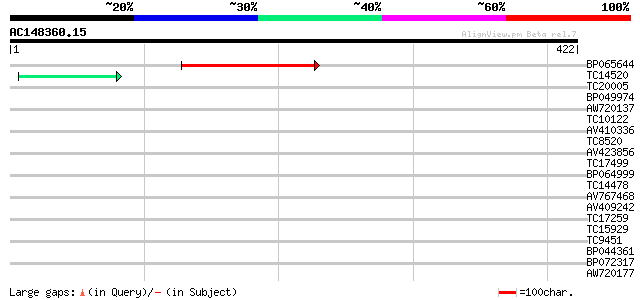
Query= AC148360.15 - phase: 0
(422 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065644 158 1e-39
TC14520 similar to UP|Q8W194 (Q8W194) Zinc finger protein LSD2, ... 42 2e-04
TC20005 similar to UP|Q7XYB8 (Q7XYB8) 18S subunit ribosomal prot... 37 0.006
BP049974 36 0.014
AW720137 31 0.34
TC10122 29 1.3
AV410336 28 2.8
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 28 2.8
AV423856 28 3.7
TC17499 weakly similar to UP|O82212 (O82212) At2g23820 protein, ... 28 3.7
BP064999 27 4.9
TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-... 27 4.9
AV767468 27 4.9
AV409242 27 4.9
TC17259 weakly similar to UP|Q94KL5 (Q94KL5) BEL1-like homeobox ... 27 6.3
TC15929 similar to UP|THM4_ARATH (Q9SEU6) Thioredoxin M-type 4, ... 27 6.3
TC9451 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase E1... 27 6.3
BP044361 27 6.3
BP072317 27 6.3
AW720177 27 6.3
>BP065644
Length = 315
Score = 158 bits (400), Expect = 1e-39
Identities = 75/102 (73%), Positives = 86/102 (83%)
Frame = +1
Query: 129 LVNEVEQEEGDGGIAGETFTDYRPSKLSIGSPHPDPIVETSSLSAVQPPEPTYDPKIKND 188
LV EVE++E GG+AGETF DYRP+K+S+G PHPD +VETSSLSAV PPEPTY+P+IK+
Sbjct: 10 LVVEVERDEDGGGVAGETFMDYRPAKVSMGPPHPDSVVETSSLSAVPPPEPTYNPRIKDH 189
Query: 189 LERSKALSCLQIETLVYACQRHLQHVPSGPRAGFFIGDGAGV 230
E SK LS LQIET VYACQRH QH+P G RAGFFIGDGAGV
Sbjct: 190 SESSKTLSSLQIETSVYACQRHTQHLPDGTRAGFFIGDGAGV 315
>TC14520 similar to UP|Q8W194 (Q8W194) Zinc finger protein LSD2, partial
(84%)
Length = 1112
Score = 42.0 bits (97), Expect = 2e-04
Identities = 18/77 (23%), Positives = 31/77 (39%)
Frame = +2
Query: 7 PPPPSTARARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNAPS 66
PP ++ C GCRT +G A + C C ++ + + C +C+ + P
Sbjct: 383 PPGMDMSQLYCGGCRTLLMYTRGAASVRCSCCHTVNLAPAANQVAHVPCGNCRTTLMYPY 562
Query: 67 NLSKFPCPQCHVRIDVH 83
C CH +V+
Sbjct: 563 GAPSVKCALCHYITNVN 613
Score = 27.3 bits (59), Expect = 4.9
Identities = 19/85 (22%), Positives = 30/85 (34%), Gaps = 21/85 (24%)
Frame = +2
Query: 17 CAGCRTYFSAAQG-----------VAELPCPNCQMPHVF--------FVDSSAVKIRCSS 57
C GCR+ +G + +P P M ++ A +RCS
Sbjct: 296 CNGCRSILLYPRGATNVCCALCNTITAVPPPGMDMSQLYCGGCRTLLMYTRGAASVRCSC 475
Query: 58 CKAVVNAP--SNLSKFPCPQCHVRI 80
C V AP + ++ PC C +
Sbjct: 476 CHTVNLAPAANQVAHVPCGNCRTTL 550
>TC20005 similar to UP|Q7XYB8 (Q7XYB8) 18S subunit ribosomal protein
(Fragment), partial (36%)
Length = 504
Score = 37.0 bits (84), Expect = 0.006
Identities = 19/74 (25%), Positives = 27/74 (35%), Gaps = 3/74 (4%)
Frame = +3
Query: 7 PPPPS---TARARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVN 63
P PP+ ++ C GCRT A G + C C + I C +C+ +
Sbjct: 222 PVPPAGMEMSQLYCGGCRTLLMYANGATSVRCSCCNTITRVPESNQVSHIHCGNCRTALM 401
Query: 64 APSNLSKFPCPQCH 77
P C CH
Sbjct: 402 YPHGALSVKCAICH 443
>BP049974
Length = 592
Score = 35.8 bits (81), Expect = 0.014
Identities = 15/54 (27%), Positives = 23/54 (41%), Gaps = 2/54 (3%)
Frame = +2
Query: 8 PPPSTARAR--CAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCK 59
PPP T A+ C GC T +G + C C ++ + + C +CK
Sbjct: 431 PPPGTEMAQLVCGGCHTLIMYIRGATSVQCSCCHTVNLALEANQVAHVHCGNCK 592
>AW720137
Length = 563
Score = 31.2 bits (69), Expect = 0.34
Identities = 15/34 (44%), Positives = 21/34 (61%)
Frame = +3
Query: 1 MNRQPQPPPPSTARARCAGCRTYFSAAQGVAELP 34
M+RQPQPPPP+ A A T+F+ + + LP
Sbjct: 201 MSRQPQPPPPNPATA--DDSLTFFNQSMELVNLP 296
>TC10122
Length = 544
Score = 29.3 bits (64), Expect = 1.3
Identities = 12/30 (40%), Positives = 14/30 (46%)
Frame = -3
Query: 217 GPRAGFFIGDGAGVGKGRTVAGLIWENWHH 246
G R G G G G G R V +W W+H
Sbjct: 368 GERVGVVAGHGVGDGGWRVVVR*VWRVWYH 279
>AV410336
Length = 358
Score = 28.1 bits (61), Expect = 2.8
Identities = 14/31 (45%), Positives = 17/31 (54%), Gaps = 5/31 (16%)
Frame = +1
Query: 159 SPHPDPIVETSSLSAV-----QPPEPTYDPK 184
SPHP P T+S S+ PP PT +PK
Sbjct: 10 SPHPSPPTPTTSTSSTAPPPHAPPPPTPNPK 102
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 28.1 bits (61), Expect = 2.8
Identities = 23/73 (31%), Positives = 27/73 (36%), Gaps = 18/73 (24%)
Frame = -1
Query: 4 QPQPPPPSTARARCAGCRTYF------------------SAAQGVAELPCPNCQMPHVFF 45
Q PPPP AR+ A +F S VA L PN +
Sbjct: 545 QKSPPPPPAARSSAAKIDEHFRLSVEFQWFLMALSVRPESKRAMVAHL-FPNRACALIMV 369
Query: 46 VDSSAVKIRCSSC 58
SS VK RCS+C
Sbjct: 368 SSSSGVKARCSTC 330
>AV423856
Length = 342
Score = 27.7 bits (60), Expect = 3.7
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +2
Query: 153 SKLSIGSPHPDPIVETSSLSAVQPPEPTYDPKI 185
S +S+ SP P SSL+ P P+ DPK+
Sbjct: 167 STISLPSPKPKTKPRASSLAISAPKPPSIDPKL 265
>TC17499 weakly similar to UP|O82212 (O82212) At2g23820 protein, partial
(40%)
Length = 502
Score = 27.7 bits (60), Expect = 3.7
Identities = 14/28 (50%), Positives = 16/28 (57%)
Frame = +3
Query: 153 SKLSIGSPHPDPIVETSSLSAVQPPEPT 180
S LSI P PI+ SS SA PP P+
Sbjct: 102 SHLSIALPLLPPIIPNSSSSATLPPSPS 185
>BP064999
Length = 547
Score = 27.3 bits (59), Expect = 4.9
Identities = 25/90 (27%), Positives = 38/90 (41%)
Frame = +2
Query: 60 AVVNAPSNLSKFPCPQCHVRIDVHADVEEVNEKIDSEPVISCFHKFSQTVSQELDMLQNP 119
A ++PS P+ H I V D+ NE + ++ + +EL +QN
Sbjct: 251 AAADSPSTSGGSAAPEFH--IGVFKDLAVANETMREAA--------AKQMVRELKEVQNA 400
Query: 120 RETVGKQSVLVNEVEQEEGDGGIAGETFTD 149
+ +G E E+EEGDGG E D
Sbjct: 401 YDGLG-------ESEKEEGDGGFKLEAEKD 469
>TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-like
protein, partial (83%)
Length = 1784
Score = 27.3 bits (59), Expect = 4.9
Identities = 13/34 (38%), Positives = 15/34 (43%)
Frame = +2
Query: 1 MNRQPQPPPPSTARARCAGCRTYFSAAQGVAELP 34
M R PPPPS AR R C + + LP
Sbjct: 482 MCRARSPPPPSAARLRSRRCSPWRCCLSRIHRLP 583
>AV767468
Length = 299
Score = 27.3 bits (59), Expect = 4.9
Identities = 13/33 (39%), Positives = 16/33 (48%)
Frame = -2
Query: 35 CPNCQMPHVFFVDSSAVKIRCSSCKAVVNAPSN 67
CP C+ P DS+ V+ SS V PSN
Sbjct: 274 CPTCRTPVRLNADSATVQSHPSSSTQFVTCPSN 176
>AV409242
Length = 315
Score = 27.3 bits (59), Expect = 4.9
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = -2
Query: 3 RQPQPPPPSTARARCAGCRT 22
R P P PS R RC GCR+
Sbjct: 227 RFPLPSQPSLLRGRCDGCRS 168
>TC17259 weakly similar to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, partial
(6%)
Length = 552
Score = 26.9 bits (58), Expect = 6.3
Identities = 13/35 (37%), Positives = 19/35 (54%), Gaps = 4/35 (11%)
Frame = +1
Query: 161 HPDPIVETSSLSAVQPPEPTY----DPKIKNDLER 191
H +V+T + SA PP PT+ DP + N + R
Sbjct: 7 HHHHLVQTPTPSATTPPPPTFQSENDPSVLNAINR 111
>TC15929 similar to UP|THM4_ARATH (Q9SEU6) Thioredoxin M-type 4, chloroplast
precursor (TRX-M4), partial (49%)
Length = 1406
Score = 26.9 bits (58), Expect = 6.3
Identities = 14/42 (33%), Positives = 19/42 (44%)
Frame = +2
Query: 1 MNRQPQPPPPSTARARCAGCRTYFSAAQGVAELPCPNCQMPH 42
++R P PPP A A C T S++ P P +PH
Sbjct: 134 LHRSPPPPPSPAAEALPGCCTTPASSSD-----PSPELVLPH 244
>TC9451 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase E1 alpha
subunit (At1g01090/T25K16_8) , partial (52%)
Length = 1082
Score = 26.9 bits (58), Expect = 6.3
Identities = 13/26 (50%), Positives = 15/26 (57%)
Frame = -2
Query: 12 TARARCAGCRTYFSAAQGVAELPCPN 37
T RARCA RTY S + LP P+
Sbjct: 778 THRARCALLRTYASNPPDLGRLPSPS 701
>BP044361
Length = 542
Score = 26.9 bits (58), Expect = 6.3
Identities = 23/93 (24%), Positives = 41/93 (43%), Gaps = 7/93 (7%)
Frame = +1
Query: 102 FHKFSQTVSQELDMLQNPRETVGKQ-SVLVNEVEQEEGDGGI------AGETFTDYRPSK 154
FHK + Q + P + +Q ++ +N + E + I +GET + + +
Sbjct: 154 FHKLCDILRQRTLLRDTPGVMIEEQLAIFLNIIGHNERNRVIQERFQHSGETISRHFNNV 333
Query: 155 LSIGSPHPDPIVETSSLSAVQPPEPTYDPKIKN 187
L V++ S +QPP+PT P+I N
Sbjct: 334 LKA--------VKSLSREVLQPPQPTTPPEILN 408
>BP072317
Length = 436
Score = 26.9 bits (58), Expect = 6.3
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = -1
Query: 147 FTDYRPSKLSIGSPHPDPIVETSSL 171
FT Y PS L+ GSP P+ +SL
Sbjct: 409 FTSYNPSPLAWGSPFEVPLTSHASL 335
>AW720177
Length = 505
Score = 26.9 bits (58), Expect = 6.3
Identities = 26/131 (19%), Positives = 53/131 (39%)
Frame = +3
Query: 94 DSEPVISCFHKFSQTVSQELDMLQNPRETVGKQSVLVNEVEQEEGDGGIAGETFTDYRPS 153
+S+P+ C H + + +E+ +QN + V + ++ D + E + +
Sbjct: 90 ESDPITDCVHLLTSLLDEEIPTVQNLKTKWSLVRVKLTNLQTHLTD--FSAEFPSSSATN 263
Query: 154 KLSIGSPHPDPIVETSSLSAVQPPEPTYDPKIKNDLERSKALSCLQIETLVYACQRHLQH 213
LS+ H + + SAV + + P L R K + +++L+ HL H
Sbjct: 264 PLSLDLFHS---LSATLHSAVSLAQQCHSP---THLPRGKLQTQSDLDSLLATLDCHLSH 425
Query: 214 VPSGPRAGFFI 224
R+G +
Sbjct: 426 CDILLRSGLLL 458
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,816,806
Number of Sequences: 28460
Number of extensions: 146225
Number of successful extensions: 1146
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1136
length of query: 422
length of database: 4,897,600
effective HSP length: 93
effective length of query: 329
effective length of database: 2,250,820
effective search space: 740519780
effective search space used: 740519780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148360.15


