

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
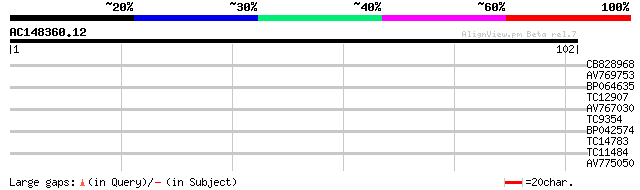
Query= AC148360.12 + phase: 0 /pseudo
(102 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB828968 25 1.6
AV769753 25 2.1
BP064635 25 2.1
TC12907 UP|Q40201 (Q40201) RAB1A, complete 25 2.1
AV767030 24 3.6
TC9354 similar to UP|Q9FN03 (Q9FN03) UVB-resistance protein UVR8... 23 6.1
BP042574 23 6.1
TC14783 UP|O04360 (O04360) Late nodulin Nlj16, complete 23 8.0
TC11484 similar to PIR|JC5965|JC5965 cytochrome P450 CYP86A1 - A... 23 8.0
AV775050 23 8.0
>CB828968
Length = 572
Score = 25.4 bits (54), Expect = 1.6
Identities = 8/20 (40%), Positives = 11/20 (55%)
Frame = -3
Query: 3 TKVDPPFDVIWVYSAIKFGC 22
+K PPF + WV+ F C
Sbjct: 354 SK*SPPFSIFWVFQDFSFQC 295
>AV769753
Length = 496
Score = 25.0 bits (53), Expect = 2.1
Identities = 12/37 (32%), Positives = 20/37 (53%)
Frame = +1
Query: 44 ISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMS 80
+S CS S SKS + + + +K+ S+V G +S
Sbjct: 343 LSCCSNSAENSKSSSSASARCLKDFTSIVSFASGGLS 453
>BP064635
Length = 504
Score = 25.0 bits (53), Expect = 2.1
Identities = 11/26 (42%), Positives = 17/26 (65%), Gaps = 3/26 (11%)
Frame = -3
Query: 64 AMKEVKSLVDMI---LGFMSICCSKI 86
A V LVD++ LGF+++CCS +
Sbjct: 373 AYGRVLGLVDLVVSMLGFLALCCSTV 296
>TC12907 UP|Q40201 (Q40201) RAB1A, complete
Length = 869
Score = 25.0 bits (53), Expect = 2.1
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +2
Query: 10 DVIWVYSAIKFGCRKSLKGDILEQI 34
DV W++ K C KSL +L I
Sbjct: 791 DVTWMFETAKISCSKSLYNIVLNLI 865
>AV767030
Length = 460
Score = 24.3 bits (51), Expect = 3.6
Identities = 18/52 (34%), Positives = 25/52 (47%)
Frame = +3
Query: 48 SASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICCSKISEEKDLDLVLSFS 99
SA + GSK M V L D++LG + I S S ++ LVL+ S
Sbjct: 270 SA*LAGSKQRVKSTSLVMDLVTDLADLVLGALEIAASVFSVKR---LVLTSS 416
>TC9354 similar to UP|Q9FN03 (Q9FN03) UVB-resistance protein UVR8
(AT5g63860/MGI19_6), partial (8%)
Length = 585
Score = 23.5 bits (49), Expect = 6.1
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 11 VIWVYSAIKFGCRKSLKGDILEQIS 35
V W+Y KFG R SL+ + +I+
Sbjct: 461 VCWLYDL*KFGLRYSLQATYVVEIN 535
>BP042574
Length = 505
Score = 23.5 bits (49), Expect = 6.1
Identities = 16/44 (36%), Positives = 21/44 (47%)
Frame = -1
Query: 48 SASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICCSKISEEKD 91
S+S S ALL+ E +S + + SI S ISE KD
Sbjct: 379 SSSRASWASAALLSTSNNSETRSSISCSVRGSSITLSSISEGKD 248
>TC14783 UP|O04360 (O04360) Late nodulin Nlj16, complete
Length = 540
Score = 23.1 bits (48), Expect = 8.0
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +3
Query: 77 GFMSICCSKISEEKDLDLVLS 97
G S+C K +EE+D+ L+LS
Sbjct: 432 GAFSLCXEKETEEEDVFLLLS 494
>TC11484 similar to PIR|JC5965|JC5965 cytochrome P450 CYP86A1 - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (43%)
Length = 752
Score = 23.1 bits (48), Expect = 8.0
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = -2
Query: 81 ICCSKISEEKDLDLVLSFSSL 101
ICC +SEE+ L+ L S L
Sbjct: 604 ICCQMLSEEEGLEDPLELSPL 542
>AV775050
Length = 382
Score = 23.1 bits (48), Expect = 8.0
Identities = 6/17 (35%), Positives = 13/17 (76%)
Frame = -2
Query: 9 FDVIWVYSAIKFGCRKS 25
F ++W +++++FGC S
Sbjct: 96 FIILWFHNSVEFGCSVS 46
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,656,513
Number of Sequences: 28460
Number of extensions: 16291
Number of successful extensions: 79
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 79
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 79
length of query: 102
length of database: 4,897,600
effective HSP length: 78
effective length of query: 24
effective length of database: 2,677,720
effective search space: 64265280
effective search space used: 64265280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 47 (22.7 bits)
Medicago: description of AC148360.12


