

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.2 + phase: 0 /pseudo
(813 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
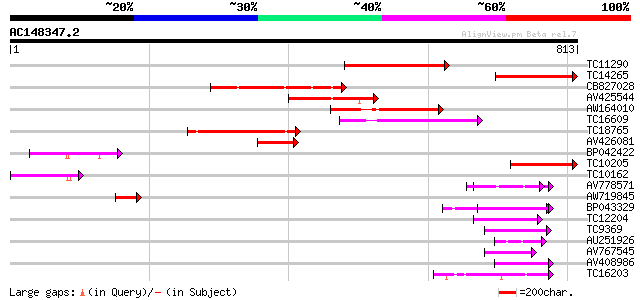
Sequences producing significant alignments: (bits) Value
TC11290 weakly similar to UP|AAN62760 (AAN62760) Disease resista... 186 2e-47
TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resista... 171 4e-43
CB827028 152 2e-37
AV425544 140 8e-34
AW164010 124 8e-29
TC16609 weakly similar to PIR|T48468|T48468 disease resistance-l... 110 7e-25
TC18765 weakly similar to UP|Q9LVT1 (Q9LVT1) Disease resistance ... 100 7e-22
AV426081 99 3e-21
BP042422 94 1e-19
TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%) 86 2e-17
TC10162 79 4e-15
AV778571 62 4e-10
AW719845 58 5e-09
BP043329 51 6e-07
TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protei... 47 9e-06
TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resista... 45 5e-05
AU251926 45 6e-05
AV767545 44 1e-04
AV408986 44 1e-04
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 43 2e-04
>TC11290 weakly similar to UP|AAN62760 (AAN62760) Disease resistance
protein-like protein MsR1, partial (18%)
Length = 459
Score = 186 bits (471), Expect = 2e-47
Identities = 97/152 (63%), Positives = 119/152 (77%), Gaps = 2/152 (1%)
Frame = +3
Query: 481 GIYQSAKEPFEQRKRLIIDINKNKSG--LAEKQQGLMTCILSKFMRLCVKRNPQQLTARI 538
GIYQS +EP +RKRL ID N++K L EKQQG+M+ LSK RLC+K+ PQ + AR
Sbjct: 3 GIYQSIQEPVGKRKRLNIDTNEDKLEWLLQEKQQGIMSRTLSKIRRLCLKQKPQLVLART 182
Query: 539 LSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKL 598
LS+S DETC D S ++PA+ EVLILNL TK+YSLPE + +M+KL+ LI+TNY FHPS+L
Sbjct: 183 LSISTDETCTPDLSLIQPAEAEVLILNLRTKKYSLPEILEEMNKLKALIVTNYGFHPSEL 362
Query: 599 NNIELLGSLQNLERIRLERIYVPSFGTLKNLK 630
NN ELL SL NL+RIRLE+I VPSFGTLKNLK
Sbjct: 363 NNFELLDSLSNLKRIRLEQISVPSFGTLKNLK 458
>TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resistance
protein-like protein MsR1, partial (5%)
Length = 766
Score = 171 bits (433), Expect = 4e-43
Identities = 84/117 (71%), Positives = 99/117 (83%)
Frame = +1
Query: 697 QDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNL 756
Q+IGKLENLELL LSSCTDL+ +P SIG+L L+ LDISNCISL SLPEE GNLCNLK+L
Sbjct: 1 QEIGKLENLELLRLSSCTDLKGLPDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSL 180
Query: 757 DMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
M SCA ELP S+VNLQNL T+ CDEETAA+WE F++++P++KIEV VDVNLNWL
Sbjct: 181 YMTSCAGCELPSSIVNLQNL-TVVCDEETAASWEAFEYVIPHLKIEVPQVDVNLNWL 348
Score = 32.0 bits (71), Expect = 0.40
Identities = 20/55 (36%), Positives = 27/55 (48%), Gaps = 1/55 (1%)
Frame = -3
Query: 716 LEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGN-LCNLKNLDMASCASIELPFS 769
L +P S GKL L D+S +S SLP E GN +++ L + S P S
Sbjct: 167 LHRLPISSGKLGRLMQXDMSRSLSFESLPIESGNPFKSVQELSLRSSKFSNFPIS 3
Score = 31.2 bits (69), Expect = 0.69
Identities = 25/76 (32%), Positives = 42/76 (54%)
Frame = +1
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERI 618
+E+L L+ T LP+ I ++SKLR+L I+N PS I L +L++L
Sbjct: 25 LELLRLSSCTDLKGLPDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSLYMTSCAGC 204
Query: 619 YVPSFGTLKNLKKLSL 634
+PS ++ NL+ L++
Sbjct: 205 ELPS--SIVNLQNLTV 246
>CB827028
Length = 572
Score = 152 bits (385), Expect = 2e-37
Identities = 91/197 (46%), Positives = 127/197 (64%), Gaps = 2/197 (1%)
Frame = +2
Query: 288 SRVAFPRFSTTC--ILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLT 345
SR A R+ + +L+ L+ + A +LF H A + NS++I +LV+K+V+ C+ P+
Sbjct: 2 SRSAIARYDSGSPYVLERLSLDYATSLFLHSASLSPNSTEI-PVSLVKKIVKGCKRSPIA 178
Query: 346 IKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDI 405
+ V TSLKN+ +WR K LS+G SIL+SN D+LT LQK DVL+ P +ECF D+
Sbjct: 179 LTVTGTSLKNKNLAVWRSRAKALSKGQSILESNHDVLTCLQKSLDVLD--PIAIECFKDL 352
Query: 406 ALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNN 465
LFPED RIP AALVDMWAEL DD AME I +L +N+A++ + RK A+D +
Sbjct: 353 GLFPEDQRIPAAALVDMWAELRCGDDE--TAMEKIYELVNLNMADIFVTRKVANDI--VD 520
Query: 466 YNNHFIILHDILRELGI 482
YN H++ H +LREL I
Sbjct: 521 YNYHYVAQHGLLRELAI 571
>AV425544
Length = 425
Score = 140 bits (353), Expect = 8e-34
Identities = 74/133 (55%), Positives = 93/133 (69%), Gaps = 3/133 (2%)
Frame = +1
Query: 400 ECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNAS 459
ECFMD+ LFPED RIPV AL+DMWAELY LD++G AM I+ L NL N I+ RK AS
Sbjct: 19 ECFMDLGLFPEDQRIPVTALIDMWAELYNLDEDGRNAMTIVLDLTSRNLINFIVTRKVAS 198
Query: 460 DTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDI---NKNKSGLAEKQQGLMT 516
D YNNHF++LHD+LREL I+QS EPFEQRKRLIID+ N+ + + + QQG +
Sbjct: 199 DA-GVCYNNHFVMLHDLLRELAIHQSKGEPFEQRKRLIIDLNGDNRPEWWVGQNQQGFIG 375
Query: 517 CILSKFMRLCVKR 529
+ S R+ VK+
Sbjct: 376 RLFSFLPRMLVKQ 414
>AW164010
Length = 494
Score = 124 bits (310), Expect = 8e-29
Identities = 65/161 (40%), Positives = 102/161 (62%)
Frame = +2
Query: 461 TDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILS 520
+D +YN H++ H +LR+L I Q+++E ++R RLII+IN+N L
Sbjct: 26 SDIVDYNYHYVTQHGLLRDLAILQTSEETEDKRDRLIIEINRNN--------------LP 163
Query: 521 KFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKM 580
+ L N + ARILS+S DE + + ++P +VE L+LNL K+Y+LP ++ KM
Sbjct: 164 AWWSL---ENEYPIAARILSISTDEAFSSELCNLQPTKVEALVLNLRGKKYTLPMFMEKM 334
Query: 581 SKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVP 621
+KL+VL ITNYD +P++L N E+L L NL+R+RLE++ +P
Sbjct: 335 NKLKVLTITNYDLYPAELENFEMLDHLSNLKRLRLEKVRIP 457
>TC16609 weakly similar to PIR|T48468|T48468 disease resistance-like protein
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(12%)
Length = 621
Score = 110 bits (276), Expect = 7e-25
Identities = 66/207 (31%), Positives = 104/207 (49%), Gaps = 2/207 (0%)
Frame = +1
Query: 474 HDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQ 533
HDILR+L + S + +R RL++ + L ++ ++ Q
Sbjct: 43 HDILRDLALNLSNRGSINERLRLVMPKREGNGQLPKEW---------------LRYRGQP 177
Query: 534 LTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDF 593
L ARI+S+ E DW ++E + EVLILN + +Y LP +IA+M LR LI+ NY
Sbjct: 178 LEARIVSIHTGEMTEGDWCELEFPKAEVLILNFTSSEYFLPPFIARMPSLRALIVINYST 357
Query: 594 HPSKLNNIELLGSLQNLERIRLERIYVPSFG--TLKNLKKLSLYMCNTILAFEKGSILIS 651
+ L+N+ + +L NL + LE++ P L+ L KL + +C + + ++
Sbjct: 358 TYAHLHNVSVFKNLSNLRSLWLEKVSTPQLSGIVLEKLGKLFIVLCQVNNSLKGKDANLA 537
Query: 652 DAFPNLEELNIDYCKDLVVLPIGICDI 678
FPNL E +D+C DL LP IC I
Sbjct: 538 HIFPNLSEFTLDHCDDLTELPSSICGI 618
>TC18765 weakly similar to UP|Q9LVT1 (Q9LVT1) Disease resistance
protein-like, partial (18%)
Length = 688
Score = 100 bits (250), Expect = 7e-22
Identities = 61/164 (37%), Positives = 99/164 (60%), Gaps = 2/164 (1%)
Frame = +2
Query: 255 SPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPR-FSTTCILKPLAHEEAVTLF 313
S +L+VLDDVW + P++E++ ++ K LV SR F R F+ T ++ L+ +A++LF
Sbjct: 206 SQILVVLDDVWSL--PVLEQLVLRVPGCKYLVVSRFKFQRIFNDTYDVELLSEGDALSLF 379
Query: 314 HHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHS 373
H+A K+ N+NL+++VV C LPL +KVI SL+++ W + LSQG S
Sbjct: 380 CHHAFGHKSIPFGANQNLIKQVVAECGRLPLALKVIGASLRDQNEMFWLSVKTRLSQGLS 559
Query: 374 ILDS-NTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPV 416
I +S +L+ R+ + L + + ECF+D+ FPED +IP+
Sbjct: 560 IGESYEVNLIDRMAISTNYLPEK--VKECFLDLCAFPEDKKIPL 685
>AV426081
Length = 179
Score = 98.6 bits (244), Expect = 3e-21
Identities = 45/59 (76%), Positives = 52/59 (87%)
Frame = +1
Query: 356 RPRDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRI 414
RP ++W+KI KELSQGHS+LDS+T+LLTRLQKI DV EDNP I ECFMD+ALFPED RI
Sbjct: 1 RPYELWQKIVKELSQGHSVLDSHTELLTRLQKILDVAEDNPVIKECFMDLALFPEDQRI 177
>BP042422
Length = 526
Score = 93.6 bits (231), Expect = 1e-19
Identities = 59/154 (38%), Positives = 84/154 (54%), Gaps = 20/154 (12%)
Frame = +3
Query: 29 KAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLDHPREEINSLIEENDAGES-------- 80
K ++ + R+++R +++ V Q + HLD PREEI +LI E DAGE
Sbjct: 66 KEEEQHTRRKNIRGCWLNMADVAQ-LAFLTLHLDPPREEIKTLIREKDAGEGDKCLCWGN 242
Query: 81 ------ACT---CSSENDSYGYVVEENQSLTVNDVEETLYKTREILELLNHEFNED---G 128
C C + D + V E Q+ NDV++TLYK R LEL++ E E G
Sbjct: 243 FISLLQLCVNKLCHNNKDDFLNVDGEKQAQMANDVKDTLYKVRGFLELISKEDFEQKFSG 422
Query: 129 QLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGG 162
KRP+ VP N +FTVG+D+P +KLK+ELL+ G
Sbjct: 423 APIKRPYGVPANPEFTVGVDVPLNKLKVELLKDG 524
Score = 28.5 bits (62), Expect = 4.4
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = +2
Query: 196 IHCWVGYTI*QVENGASEGW 215
IHCW G + VE GAS+ W
Sbjct: 464 IHCWGGCAVE*VEGGASQRW 523
>TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%)
Length = 586
Score = 85.9 bits (211), Expect = 2e-17
Identities = 42/96 (43%), Positives = 65/96 (66%), Gaps = 1/96 (1%)
Frame = +3
Query: 719 IPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI-ELPFSVVNLQNLK 777
+P SI L ++ LDIS+CISLS LPE+ G L L+ L++ C+ + ELP S+++L+NL+
Sbjct: 9 LPKSITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRLTELPTSIMDLENLE 188
Query: 778 TITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
+ CDEE A WE Q +L ++++ V D NL++L
Sbjct: 189 EVECDEEIAELWEPVQQILSDLQLHVAPTDFNLDFL 296
>TC10162
Length = 577
Score = 78.6 bits (192), Expect = 4e-15
Identities = 46/122 (37%), Positives = 70/122 (56%), Gaps = 16/122 (13%)
Frame = +1
Query: 1 MSSATHIITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDH 60
M+ AT ++ L +F ++L KISQ+++K++K+ ++R LR L + + VV +IK+YN+H
Sbjct: 199 MADATQLVALAPLARFQEILKKISQMVEKSRKSGIAQRILRSTLIEFNHVVLEIKRYNEH 378
Query: 61 LDHPREEINSLIEENDAGESA-----CTCS-----------SENDSYGYVVEENQSLTVN 104
L PREEI +LI E DAGE C S ++DS +E Q+LT
Sbjct: 379 LAQPREEIKTLIREKDAGEEGEEGDKCVNSCLQVVTDRVGHGKDDSDSVDADEKQALTAK 558
Query: 105 DV 106
DV
Sbjct: 559 DV 564
>AV778571
Length = 452
Score = 62.0 bits (149), Expect = 4e-10
Identities = 35/113 (30%), Positives = 62/113 (53%)
Frame = +3
Query: 655 PNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCT 714
PNLE+L+ Y ++L+ + + + L+ L + C K+ + P KL +LE L LS C+
Sbjct: 84 PNLEKLSFVYSENLIKIHESVGFLNKLRILDIEGCCKIRTFPPM--KLASLEELYLSHCS 257
Query: 715 DLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIELP 767
LE+ P + K+ N+ + + + + LP GNL L+ L++ C ++LP
Sbjct: 258 SLESFPEILAKMENITLIGVYHT-PIKELPSSIGNLTRLRRLELHGCGMLQLP 413
Score = 48.9 bits (115), Expect = 3e-06
Identities = 31/115 (26%), Positives = 54/115 (46%)
Frame = +3
Query: 665 CKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIG 724
C+ + +P + + L+KL L + + +G L L +L + C + P
Sbjct: 45 CESITQIP-DLSGLPNLEKLSFVYSENLIKIHESVGFLNKLRILDIEGCCKIRTFPPM-- 215
Query: 725 KLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTI 779
KL +L+ L +S+C SL S PE + N+ + + ELP S+ NL L+ +
Sbjct: 216 KLASLEELYLSHCSSLESFPEILAKMENITLIGVYHTPIKELPSSIGNLTRLRRL 380
>AW719845
Length = 209
Score = 58.2 bits (139), Expect = 5e-09
Identities = 25/38 (65%), Positives = 32/38 (83%)
Frame = +3
Query: 152 SKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEV 189
SKLKMELL+ G S + +TGLGGLGKTTLA +CWD+++
Sbjct: 6 SKLKMELLKDGVSVVAVTGLGGLGKTTLAAMVCWDKQI 119
Score = 32.7 bits (73), Expect = 0.24
Identities = 19/38 (50%), Positives = 24/38 (63%)
Frame = +2
Query: 206 QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS* 243
Q E+GAS+G L D FR +GEN+ G LLG* +*
Sbjct: 8 QAEDGASQGRCLGRRCDRFRWVGENNFGCNGLLG*TN* 121
>BP043329
Length = 484
Score = 51.2 bits (121), Expect = 6e-07
Identities = 35/111 (31%), Positives = 58/111 (51%), Gaps = 2/111 (1%)
Frame = +3
Query: 671 LPIGICDIFLLKKLRVT-NCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNL 729
+P + + LK L ++ N +P +IG L NLE++ L+ C + IP SIG L L
Sbjct: 6 IPPSLGTLTTLKMLNLSYNPFYPGRIPPEIGNLTNLEVVWLTQCNLVGVIPDSIGNLKKL 185
Query: 730 KHLDISNCISLSSLPEEFGNLCNLKNLDM-ASCASIELPFSVVNLQNLKTI 779
K LD++ S+P L +L+ +++ + S ELP + NL L+ +
Sbjct: 186 KDLDLALNDLYGSIPSSLTGLTSLRQIELYNNSLSGELPRGMGNLTELRLL 338
Score = 42.0 bits (97), Expect = 4e-04
Identities = 46/158 (29%), Positives = 73/158 (46%), Gaps = 2/158 (1%)
Frame = +3
Query: 621 PSFGTLKNLKKLSLYMCNTILAFEKGSILISDA-FPNLEELNIDYCKDLVVLPIGICDIF 679
PS GTL LK L+L + F G I NLE + + C + V+P I ++
Sbjct: 12 PSLGTLTTLKMLNL----SYNPFYPGRIPPEIGNLTNLEVVWLTQCNLVGVIPDSIGNLK 179
Query: 680 LLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCIS 739
LK L + S+P + L +L + L + + +P +G L L+ LD S
Sbjct: 180 KLKDLDLALNDLYGSIPSSLTGLTSLRQIELYNNSLSGELPRGMGNLTELRLLDASMNHL 359
Query: 740 LSSLPEEFGNLCNLKNLDM-ASCASIELPFSVVNLQNL 776
+PEE +L L++L++ + ELP S+ + NL
Sbjct: 360 TGRIPEELCSL-PLESLNLYENRFEGELPASIADSPNL 470
>TC12204 weakly similar to UP|Q9ZSN3 (Q9ZSN3) NBS-LRR-like protein cD7
(Fragment), partial (3%)
Length = 328
Score = 47.4 bits (111), Expect = 9e-06
Identities = 32/100 (32%), Positives = 50/100 (50%), Gaps = 1/100 (1%)
Frame = +1
Query: 666 KDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGK 725
+ LVV + + + L LR+ NC S P+ + N+ L L C L++ P + K
Sbjct: 10 ESLVVTGVQLQYLQSLNSLRICNCPNFESFPEGGLRAPNMTNLHLEKCKKLKSFPQQMNK 189
Query: 726 -LLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI 764
LL+L L+I C L S+PE G +L L++ CA +
Sbjct: 190 MLLSLMTLNIKECPELESIPEG-GFPDSLNLLEIFHCAKL 306
Score = 38.5 bits (88), Expect = 0.004
Identities = 30/110 (27%), Positives = 53/110 (47%), Gaps = 1/110 (0%)
Frame = +1
Query: 608 QNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKD 667
QNLE + + + + L++L L + C +F +G + PN+ L+++ CK
Sbjct: 1 QNLESLVVTGVQLQY---LQSLNSLRICNCPNFESFPEGGLRA----PNMTNLHLEKCKK 159
Query: 668 LVVLPIGICDIFL-LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
L P + + L L L + C +L S+P+ G ++L LL + C L
Sbjct: 160 LKSFPQQMNKMLLSLMTLNIKECPELESIPEG-GFPDSLNLLEIFHCAKL 306
>TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resistance
protein-like protein, partial (70%)
Length = 859
Score = 45.1 bits (105), Expect = 5e-05
Identities = 32/98 (32%), Positives = 52/98 (52%), Gaps = 1/98 (1%)
Frame = +1
Query: 681 LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISL 740
L +L + N +P IG+L+ L++L+L +AIP IG+L +L HL +S
Sbjct: 13 LTRLDLHNNKLTGPIPPQIGRLKRLKILNLRWNKLQDAIPPEIGELKSLTHLYLSFNSFK 192
Query: 741 SSLPEEFGNLCNLKNLDMASCASI-ELPFSVVNLQNLK 777
+P+E NL +L+ L + I +P + LQNL+
Sbjct: 193 GEIPKELANLPDLRYLYLHENRLIGRIPPELGTLQNLR 306
>AU251926
Length = 335
Score = 44.7 bits (104), Expect = 6e-05
Identities = 29/75 (38%), Positives = 41/75 (54%)
Frame = +2
Query: 695 LPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLK 754
LP +GKL +L L LS + A+P++IG L +L LD+ + + LP+ GNL NL
Sbjct: 107 LPDSLGKLSSLVTLDLSE-NRIVALPSTIGGLSSLTRLDL-HTNRIQELPDSIGNLLNLV 280
Query: 755 NLDMASCASIELPFS 769
LD+ LP S
Sbjct: 281 YLDLRGNQLPSLPAS 325
Score = 39.3 bits (90), Expect = 0.003
Identities = 24/59 (40%), Positives = 35/59 (58%)
Frame = +2
Query: 690 HKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFG 748
+++ +LP IG L +L L L + ++ +P SIG LLNL +LD+ L SLP FG
Sbjct: 161 NRIVALPSTIGGLSSLTRLDLHT-NRIQELPDSIGNLLNLVYLDLRG-NQLPSLPASFG 331
>AV767545
Length = 562
Score = 43.9 bits (102), Expect = 1e-04
Identities = 27/75 (36%), Positives = 41/75 (54%)
Frame = -1
Query: 681 LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISL 740
LK L + + + +P I L +LE L +S L AIP+S+G+LL L+ LD+S
Sbjct: 535 LKVLNLGSNNISGPIPVSISNLIDLERLDISRNHILGAIPSSLGQLLELQWLDVSINSLA 356
Query: 741 SSLPEEFGNLCNLKN 755
S+P + NLK+
Sbjct: 355 GSIPSSLSQITNLKH 311
>AV408986
Length = 434
Score = 43.9 bits (102), Expect = 1e-04
Identities = 27/86 (31%), Positives = 44/86 (50%), Gaps = 1/86 (1%)
Frame = +3
Query: 695 LPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLK 754
+P +IG + +LELL+L + AIP +GKL LK L + ++P E GN N
Sbjct: 15 IPPEIGNISSLELLALHQNSFSGAIPKELGKLSGLKRLYVYTNQLNGTIPTELGNCTNAI 194
Query: 755 NLDMASCASIE-LPFSVVNLQNLKTI 779
+D++ I +P + + NL +
Sbjct: 195 EIDLSENRLIGIIPKELGQISNLSLL 272
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/88 (27%), Positives = 44/88 (49%)
Frame = +3
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
+P I +I L+ L + ++P+++GKL L+ L + + IPT +G N
Sbjct: 15 IPPEIGNISSLELLALHQNSFSGAIPKELGKLSGLKRLYVYTNQLNGTIPTELGNCTNAI 194
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDM 758
+D+S + +P+E G + NL L +
Sbjct: 195 EIDLSENRLIGIIPKELGQISNLSLLHL 278
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation
aberrant root formation protein), complete
Length = 3308
Score = 43.1 bits (100), Expect = 2e-04
Identities = 60/207 (28%), Positives = 91/207 (42%), Gaps = 35/207 (16%)
Frame = +2
Query: 608 QNLERIRLERIYVPSFG-------TLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEEL 660
QNL + L VP FG L+ L+ L++ M N L + S L S +L+ L
Sbjct: 323 QNLRVVALNVTLVPLFGHLPPEIGLLEKLENLTISMNN--LTDQLPSDLAS--LTSLKVL 490
Query: 661 NIDYCKDLVVLPIGIC-DIFLLKKLRVTNCHKLSSLPQDIGKLE---------------- 703
NI + P I + L+ L + LP++I KLE
Sbjct: 491 NISHNLFSGQFPGNITVGMTELEALDAYDNSFSGPLPEEIVKLEKLKYLHLAGNYFSGTI 670
Query: 704 --------NLELLSLSSCTDLEAIPTSIGKLLNLK--HLDISNCISLSSLPEEFGNLCNL 753
+LE L L++ + +P S+ KL LK HL SN +P FG++ NL
Sbjct: 671 PESYSEFQSLEFLGLNANSLTGRVPESLAKLKTLKELHLGYSNAYE-GGIPPAFGSMENL 847
Query: 754 KNLDMASC-ASIELPFSVVNLQNLKTI 779
+ L+MA+C + E+P S+ NL L ++
Sbjct: 848 RLLEMANCNLTGEIPPSLGNLTKLHSL 928
Score = 31.2 bits (69), Expect = 0.69
Identities = 52/225 (23%), Positives = 89/225 (39%), Gaps = 49/225 (21%)
Frame = +2
Query: 560 EVLILNLHTKQYS--LPEWIAKMSKLRVLIITNYDFHP------SKLNNIELLG------ 605
E+ L+ + +S LPE I K+ KL+ L + F S+ ++E LG
Sbjct: 551 ELEALDAYDNSFSGPLPEEIVKLEKLKYLHLAGNYFSGTIPESYSEFQSLEFLGLNANSL 730
Query: 606 ---------SLQNLERIRL------ERIYVPSFGTLKNLKKLSLYMCNTI--LAFEKGSI 648
L+ L+ + L E P+FG+++NL+ L + CN + G++
Sbjct: 731 TGRVPESLAKLKTLKELHLGYSNAYEGGIPPAFGSMENLRLLEMANCNLTGEIPPSLGNL 910
Query: 649 L-ISDAFPNLEELN------IDYCKDLVVLPIGICDIF--------LLKKLRVTNCHK-- 691
+ F + L + L+ L + I D+ LK L + N +
Sbjct: 911 TKLHSLFVQMNNLTGTIPPELSSMMSLMSLDLSINDLTGEIPESFSKLKNLTLMNFFQNK 1090
Query: 692 -LSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDIS 735
SLP IG L NLE L + +P ++G + D++
Sbjct: 1091FRGSLPSFIGDLPNLETLQVWENNFSFVLPHNLGGNGRFLYFDVT 1225
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,152,255
Number of Sequences: 28460
Number of extensions: 199152
Number of successful extensions: 994
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 932
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 971
length of query: 813
length of database: 4,897,600
effective HSP length: 98
effective length of query: 715
effective length of database: 2,108,520
effective search space: 1507591800
effective search space used: 1507591800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148347.2


