

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
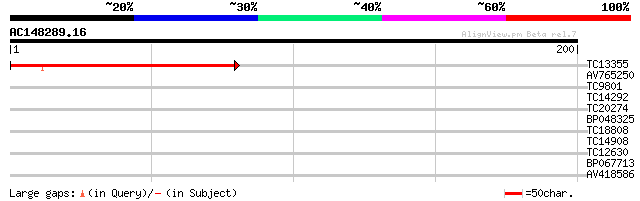
TC13355 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine pho... 136 2e-33
AV765250 38 0.001
TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like... 29 0.63
TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precur... 29 0.63
TC20274 similar to UP|OAC1_YEAST (P32332) Mitochondrial oxaloace... 26 4.1
BP048325 26 4.1
TC18808 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, part... 26 4.1
TC14908 weakly similar to PIR|A86218|A86218 protein T27G7.18 [im... 26 5.3
TC12630 similar to PIR|T39903|T39903 serine-rich protein - fissi... 25 7.0
BP067713 25 9.1
AV418586 25 9.1
>TC13355 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine phosphatase,
partial (33%)
Length = 420
Score = 136 bits (343), Expect = 2e-33
Identities = 62/82 (75%), Positives = 76/82 (92%), Gaps = 1/82 (1%)
Frame = +2
Query: 1 MIVEVENIDE-DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYP 59
++VEV++ DE DDDVL+PP NFSMVEDCI+RSS P PS+FPFL+TLNLRS+IYLCPEPYP
Sbjct: 173 ILVEVDSGDESDDDVLVPPTNFSMVEDCIFRSSFPNPSNFPFLRTLNLRSVIYLCPEPYP 352
Query: 60 EENLDFLKEQNIRLFQFGIEGK 81
+ENL+FL+ +NI+LFQFGIEGK
Sbjct: 353 QENLEFLRSENIQLFQFGIEGK 418
>AV765250
Length = 190
Score = 37.7 bits (86), Expect = 0.001
Identities = 16/21 (76%), Positives = 19/21 (90%)
Frame = -2
Query: 180 SKKRRLMYQDENIQKPRLTSF 200
SKKRRL Y ++N+QKPRLTSF
Sbjct: 186 SKKRRLSYTEDNLQKPRLTSF 124
>TC9801 homologue to UP|Q8LAC6 (Q8LAC6) Glycine rich protein-like, complete
Length = 981
Score = 28.9 bits (63), Expect = 0.63
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +2
Query: 93 IMEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
IM L +L + HP+LV K G+ G LV C
Sbjct: 95 IMLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 193
>TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precursor , partial
(95%)
Length = 1792
Score = 28.9 bits (63), Expect = 0.63
Identities = 15/36 (41%), Positives = 22/36 (60%), Gaps = 1/36 (2%)
Frame = +1
Query: 19 PNFSMVEDCIYRSSLPKPSSFPF-LQTLNLRSIIYL 53
PNFS DC++R + F F QT+N+ S++YL
Sbjct: 1603 PNFSHAFDCVHRVIIYVCPVFLF*FQTMNIFSLLYL 1710
>TC20274 similar to UP|OAC1_YEAST (P32332) Mitochondrial oxaloacetate
transport protein (Mitochondrial carrier protein PMT),
partial (7%)
Length = 509
Score = 26.2 bits (56), Expect = 4.1
Identities = 20/46 (43%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Frame = -1
Query: 43 QTLNLRSIIYLCPEP-YPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
+T NL S++Y EP Y E L+ LK Q LFQ + E SLP
Sbjct: 242 KTNNLASLVYQHFEPAYASETLEILK-QLTNLFQRHEKHNREGSLP 108
>BP048325
Length = 401
Score = 26.2 bits (56), Expect = 4.1
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 1/30 (3%)
Frame = +2
Query: 30 RSSLPK-PSSFPFLQTLNLRSIIYLCPEPY 58
RS P P++ PFL ++NL+ + L EPY
Sbjct: 290 RSKTPSFPNTNPFLSSMNLQQVECLEEEPY 379
>TC18808 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, partial (44%)
Length = 854
Score = 26.2 bits (56), Expect = 4.1
Identities = 10/21 (47%), Positives = 12/21 (56%)
Frame = -3
Query: 118 RTGCLVGCFRKLQNWCLSSAF 138
R L CF + NWC SS+F
Sbjct: 618 RDRMLAKCFCRYNNWCASSSF 556
>TC14908 weakly similar to PIR|A86218|A86218 protein T27G7.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(68%)
Length = 1219
Score = 25.8 bits (55), Expect = 5.3
Identities = 12/36 (33%), Positives = 19/36 (52%), Gaps = 4/36 (11%)
Frame = +2
Query: 113 KQGKHRTGCL----VGCFRKLQNWCLSSAFEEYQRF 144
+QG+ + CL V CF L +C S + E Y+ +
Sbjct: 98 EQGRRSSSCLLCSKVSCFLLLLIFCFSGSVEAYKNY 205
>TC12630 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe)
{Schizosaccharomyces pombe;}, partial (6%)
Length = 652
Score = 25.4 bits (54), Expect = 7.0
Identities = 13/30 (43%), Positives = 17/30 (56%), Gaps = 3/30 (10%)
Frame = +3
Query: 40 PFLQTLNLRSIIYLCPEPY---PEENLDFL 66
PF L L +++ LCPE Y P+ L FL
Sbjct: 228 PFFPLLLLLNLVLLCPEEYFLPPQTRLLFL 317
>BP067713
Length = 381
Score = 25.0 bits (53), Expect = 9.1
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = -2
Query: 20 NFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPE 56
N S E I P P S+PF L+ I LCPE
Sbjct: 236 NLSPWEMKIKTPEEP*PVSYPFSSLCVLKFFIRLCPE 126
>AV418586
Length = 416
Score = 25.0 bits (53), Expect = 9.1
Identities = 14/51 (27%), Positives = 24/51 (46%), Gaps = 1/51 (1%)
Frame = +2
Query: 9 DEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQ-TLNLRSIIYLCPEPY 58
D+D V ++D +YRS + + S P+ +L LR +Y P +
Sbjct: 233 DDDHTVTETEAAVKNLDDAVYRSEMDRLDSDPYYPLSLPLRVYVYNMPSKF 385
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,810,464
Number of Sequences: 28460
Number of extensions: 59566
Number of successful extensions: 316
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 315
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 315
length of query: 200
length of database: 4,897,600
effective HSP length: 86
effective length of query: 114
effective length of database: 2,450,040
effective search space: 279304560
effective search space used: 279304560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148289.16


