

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
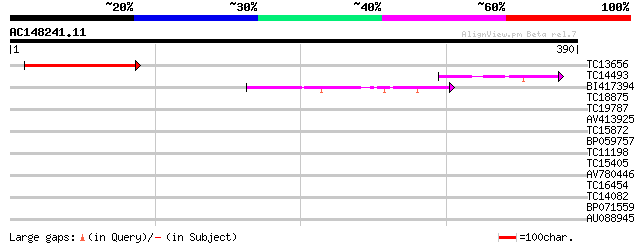
Query= AC148241.11 + phase: 0
(390 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC13656 141 2e-34
TC14493 40 7e-04
BI417394 40 9e-04
TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, pa... 36 0.013
TC19787 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, pa... 35 0.016
AV413925 30 0.69
TC15872 similar to UP|Q7MV42 (Q7MV42) DNA-binding protein HU, pa... 29 1.5
BP059757 28 2.0
TC11198 similar to UP|Q9LJT8 (Q9LJT8) Non-race specific disease ... 28 3.4
TC15405 weakly similar to GB|AAM60870.1|21536538|AY084279 root c... 27 4.5
AV780446 27 4.5
TC16454 similar to UP|O81152 (O81152) 20S proteasome beta subuni... 27 7.6
TC14082 similar to UP|Q7X9A0 (Q7X9A0) RuBisCO activase alpha for... 26 9.9
BP071559 26 9.9
AU088945 26 9.9
>TC13656
Length = 656
Score = 141 bits (356), Expect = 2e-34
Identities = 64/80 (80%), Positives = 70/80 (87%)
Frame = +3
Query: 11 KECVKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
K CV WVETYF DCLC ++DDISFSLGL SL WG+AEIPQIITIFR KS+HG+SLAFL
Sbjct: 6 KPCVGWVETYFNDCLCTMKDDISFSLGLTSLAFWGIAEIPQIITIFRTKSNHGMSLAFLC 185
Query: 71 TWVAGDICNLVGCLLEPATL 90
TWVAGD CNLVGCLLEPAT+
Sbjct: 186TWVAGDTCNLVGCLLEPATV 245
Score = 26.9 bits (58), Expect = 5.8
Identities = 29/100 (29%), Positives = 40/100 (40%), Gaps = 6/100 (6%)
Frame = +2
Query: 12 ECVKWV----ETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLA 67
+C W+ ET D CN + FS+ LM L ++F +H +S
Sbjct: 272 KCFVWLQGGPETESPDIQCNWVA-LMFSVILMMLCIISSTTFSTFFSLFVILWTHFLSFC 448
Query: 68 --FLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSF 105
F W+ + LPTQ+YTAL V NC F
Sbjct: 449 CLFFPVWIWMHVL---------MQLPTQYYTAL-VNNCKF 538
>TC14493
Length = 1214
Score = 40.0 bits (92), Expect = 7e-04
Identities = 29/88 (32%), Positives = 44/88 (49%), Gaps = 2/88 (2%)
Frame = +3
Query: 296 AIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLP--W 353
AI+ C+RIPQIW N S L + + + G +VR F +I+ N P
Sbjct: 549 AIFLCARIPQIWQNFSNRSTGEL-------SFLTSFMNFGGSMVRV--FTTIQENAPKSV 701
Query: 354 LLDATVCVALDFFIISQYIYYRYFRSSE 381
LL + VA +F I+SQ + Y+ R+ +
Sbjct: 702 LLGYAIGVATNFTILSQIVLYQKPRAEK 785
>BI417394
Length = 598
Score = 39.7 bits (91), Expect = 9e-04
Identities = 47/163 (28%), Positives = 75/163 (45%), Gaps = 20/163 (12%)
Frame = +1
Query: 164 SGIAIPNGTQKEAARGEYYYMSARSLAGSATPPSFTHLRAAKSGPSALEF--IHDSS-DD 220
S I++P Q+ + E +Y SAR L+ S TP + + L A + PS+L IH++
Sbjct: 130 SPISLPAPPQRISTGRELFYQSARYLSRSHTPTAGSIL-AQRMAPSSLNLDSIHENLLGS 306
Query: 221 DEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINL---------PLKGNSMRYGYIGFT 271
A+Q + + I ++ TFL AINL P+ N R ++ +
Sbjct: 307 AAATQSAPALKIKTTFCIVSAL-----TFLG-AINLHQSLDKRINPMFSNP-RQQFVIYV 465
Query: 272 GIKLLKVY--------VVVHSTYGQYLGWIMAAIYTCSRIPQI 306
G KL +V +++ G +LGW M +Y R+PQI
Sbjct: 466 GRKLFQVSWDRLSENGESGNNSIGTFLGWAMTFMYMGGRLPQI 594
>TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, partial (58%)
Length = 567
Score = 35.8 bits (81), Expect = 0.013
Identities = 20/53 (37%), Positives = 30/53 (55%)
Frame = +1
Query: 282 VHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
+H TY + LGW ++ S PQ+ LN +R SV GLN + L ++SY+
Sbjct: 79 LHVTY-ELLGWFAFISWSISFYPQVILNFRRKSVVGLNFDFVLLNLTKHSSYL 234
Score = 33.5 bits (75), Expect = 0.062
Identities = 14/35 (40%), Positives = 22/35 (62%)
Frame = +1
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
LG + +SW ++ PQ+I FR KS G++ F+L
Sbjct: 100 LGWFAFISWSISFYPQVILNFRRKSVVGLNFDFVL 204
>TC19787 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, partial (50%)
Length = 415
Score = 35.4 bits (80), Expect = 0.016
Identities = 19/53 (35%), Positives = 30/53 (55%)
Frame = +2
Query: 282 VHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
+H +Y + LGW ++ S PQ+ LN +R SV GLN + L +T+Y+
Sbjct: 8 LHVSY-EALGWFAFVCWSISFYPQVILNFRRKSVVGLNFDFILLNLTKHTAYL 163
Score = 32.7 bits (73), Expect = 0.11
Identities = 14/36 (38%), Positives = 22/36 (60%)
Frame = +2
Query: 35 SLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
+LG + V W ++ PQ+I FR KS G++ F+L
Sbjct: 26 ALGWFAFVCWSISFYPQVILNFRRKSVVGLNFDFIL 133
>AV413925
Length = 402
Score = 30.0 bits (66), Expect = 0.69
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +3
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSS 61
LG + +SW ++ PQ+I FR KSS
Sbjct: 93 LGWFAFISWSISFYPQVILNFRRKSS 170
Score = 26.2 bits (56), Expect = 9.9
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +3
Query: 282 VHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGS 314
+H TY + LGW ++ S PQ+ LN +R S
Sbjct: 72 LHVTY-ELLGWFAFISWSISFYPQVILNFRRKS 167
>TC15872 similar to UP|Q7MV42 (Q7MV42) DNA-binding protein HU, partial (16%)
Length = 541
Score = 28.9 bits (63), Expect = 1.5
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = -2
Query: 34 FSLGLMSLVSWGVAEIPQIITIFRNKSSHG 63
FS+ L+S++ W +PQII I R S+ G
Sbjct: 261 FSISLVSMIKWSNVCVPQIILIKR*VSAQG 172
>BP059757
Length = 373
Score = 28.5 bits (62), Expect = 2.0
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = +2
Query: 81 VGCLLEPATLPTQFYTALTVINCS 104
+GC + LPTQ ++A+T IN S
Sbjct: 263 LGCFISSFILPTQHFSAITAINIS 334
>TC11198 similar to UP|Q9LJT8 (Q9LJT8) Non-race specific disease resistance
protein-like, partial (17%)
Length = 882
Score = 27.7 bits (60), Expect = 3.4
Identities = 22/61 (36%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Frame = +3
Query: 190 AGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPR--SVDGRYG 247
A SATPPS RA + P++ + S+ E S+ S T WS+ R S R G
Sbjct: 339 APSATPPSGDSTRATRRRPASPD---PSTPPPETSRRRSTGRCTSAWSLIRR*STRSRCG 509
Query: 248 T 248
T
Sbjct: 510 T 512
>TC15405 weakly similar to GB|AAM60870.1|21536538|AY084279 root cap 1 (RCP1)
{Arabidopsis thaliana;}, partial (32%)
Length = 790
Score = 27.3 bits (59), Expect = 4.5
Identities = 12/45 (26%), Positives = 23/45 (50%), Gaps = 1/45 (2%)
Frame = +3
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNI-KRGSVEGLNPFMFVFALIAN 330
G GW ++ + Q+W NI +++GL+ F + A++ N
Sbjct: 21 GGISGWTATLLFMWMPVSQMWTNILNPENMKGLSAFSMLLAMLGN 155
>AV780446
Length = 525
Score = 27.3 bits (59), Expect = 4.5
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +3
Query: 270 FTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLN 309
F +++ +Y +VHS G+ W+ A + P +WLN
Sbjct: 39 FKCLRVFCLYNLVHSNSGE---WVCAVWFRIKDPPSVWLN 149
>TC16454 similar to UP|O81152 (O81152) 20S proteasome beta subunit PBB1 (20S
proteasome beta subunit, multicatalytic endopeptidase)
(At3g27430) , complete
Length = 1277
Score = 26.6 bits (57), Expect = 7.6
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -3
Query: 352 PWLLDATVCVALDFFIISQYIYYRYF 377
PW L +T C+A F + S Y ++F
Sbjct: 1152 PWALASTTCLASYFLVSSGLCYVKHF 1075
>TC14082 similar to UP|Q7X9A0 (Q7X9A0) RuBisCO activase alpha form
precursor, partial (90%)
Length = 1667
Score = 26.2 bits (56), Expect = 9.9
Identities = 17/50 (34%), Positives = 28/50 (56%)
Frame = -1
Query: 227 TSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLL 276
+S+IS KP+ + SV AT + LPL+ ++ +G + T +KLL
Sbjct: 356 SSDIS*AKPFHLSLSVCWLSSISAATILKLPLE--TLEFGSLELTFLKLL 213
>BP071559
Length = 392
Score = 26.2 bits (56), Expect = 9.9
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 5/56 (8%)
Frame = -2
Query: 115 CKLYTMITSSDGANIVRMLRVNCYFY--NCIILQDNEEEKRP---LNPKPSQVYSG 165
C+L +S A+ + L+V+C FY N IL +++ RP LN K Y+G
Sbjct: 208 CQLSEFPSSPSKASQIHHLKVSCIFY*ENFNILLRKKKQ*RPVVYLNKKNVYSYAG 41
>AU088945
Length = 345
Score = 26.2 bits (56), Expect = 9.9
Identities = 17/52 (32%), Positives = 27/52 (51%)
Frame = +3
Query: 231 STTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVV 282
ST+ WS +S RY + A ++G +M+YG TG+ + K +VV
Sbjct: 30 STSNRWSAKKS---RYSSSGCEASLSQMEGFNMQYGSEDCTGVVVRKXVMVV 176
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,708,522
Number of Sequences: 28460
Number of extensions: 113932
Number of successful extensions: 540
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 534
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 540
length of query: 390
length of database: 4,897,600
effective HSP length: 92
effective length of query: 298
effective length of database: 2,279,280
effective search space: 679225440
effective search space used: 679225440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148241.11


