

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
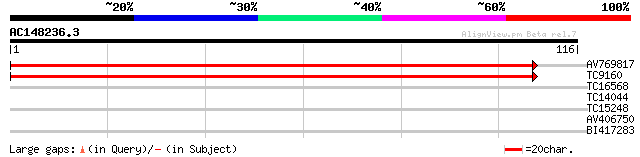
Query= AC148236.3 + phase: 0 /pseudo
(116 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV769817 196 8e-52
TC9160 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partial ... 145 1e-36
TC16568 similar to UP|AAR82759 (AAR82759) RE40656p (Fragment), p... 25 2.5
TC14044 similar to UP|Q9LN13 (Q9LN13) T6D22.2, partial (10%) 24 5.7
TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A:... 24 5.7
AV406750 23 9.6
BI417283 23 9.6
>AV769817
Length = 436
Score = 196 bits (498), Expect = 8e-52
Identities = 84/108 (77%), Positives = 101/108 (92%)
Frame = +2
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
MPWW L+FA LA +FTLP++IITATTNQTPGLN+ITEY+ G+I PGRPIANVCFKTYGY
Sbjct: 95 MPWWGLLFAGALAFVFTLPISIITATTNQTPGLNIITEYVFGLIYPGRPIANVCFKTYGY 274
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SM+QA+SFLSDFKLGHYMKIPPRSMF+VQ +GT++AGT+++GVAWWL
Sbjct: 275 ISMAQAVSFLSDFKLGHYMKIPPRSMFLVQFIGTVLAGTINIGVAWWL 418
>TC9160 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partial (53%)
Length = 1554
Score = 145 bits (367), Expect = 1e-36
Identities = 61/108 (56%), Positives = 84/108 (77%)
Frame = +2
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW ++FA LA I TLP+ +I ATTNQ PG ++I ++++G +LPG+PIAN+ FK YG
Sbjct: 320 LPWWGMLFAFALAFIVTLPIGVIQATTNQQPGYDIIAQFMIGYVLPGKPIANLLFKIYGR 499
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S A+SFLSD KLGHYMKIPPR M+ Q++GTL+A V++ VAWW+
Sbjct: 500 ISTVHALSFLSDLKLGHYMKIPPRCMYTAQLVGTLVASIVNLAVAWWM 643
>TC16568 similar to UP|AAR82759 (AAR82759) RE40656p (Fragment), partial
(13%)
Length = 421
Score = 25.4 bits (54), Expect = 2.5
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = -1
Query: 7 IFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILP 46
+ +GLA +F L A+++ +PGL V ++LP
Sbjct: 265 LVGTGLARVFLLTAALLSELETISPGLAVPEASTTSLVLP 146
>TC14044 similar to UP|Q9LN13 (Q9LN13) T6D22.2, partial (10%)
Length = 550
Score = 24.3 bits (51), Expect = 5.7
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = +2
Query: 60 YMSMSQAISFLSDFKLGHYMKIPPRS 85
Y+++S +++ L +GH +K PP S
Sbjct: 329 YLALSTSVACLVFLFIGHLLKSPPSS 406
>TC15248 weakly similar to UP|Q8GT21 (Q8GT21) Benzoyl coenzyme A: benzyl
alcohol benzoyl transferase, partial (52%)
Length = 1085
Score = 24.3 bits (51), Expect = 5.7
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -3
Query: 91 ILGTLIAGTVDVGVAWWLPRFNK 113
++G L TV V AWW+PR +
Sbjct: 234 VVG*LFVFTVVVCDAWWVPRMEE 166
>AV406750
Length = 426
Score = 23.5 bits (49), Expect = 9.6
Identities = 8/29 (27%), Positives = 18/29 (61%)
Frame = -2
Query: 81 IPPRSMFIVQILGTLIAGTVDVGVAWWLP 109
+PP + I++ + T + GT+++ + LP
Sbjct: 338 VPPHASLILEGIRTALKGTIEMAMNLMLP 252
>BI417283
Length = 557
Score = 23.5 bits (49), Expect = 9.6
Identities = 21/95 (22%), Positives = 40/95 (42%), Gaps = 7/95 (7%)
Frame = -2
Query: 19 PVAIITATTNQTPGLNVITEYIMGVILPGR-----PIANV-CFKTYGYMSMSQA-ISFLS 71
P+ ++ +T P + ++I + LP + N+ CF T GY+ ++QA I F+
Sbjct: 505 PLPLL*QSTGNVPNPVMWKQFIHSLCLPSKVRRSKQCTNLKCFSTSGYIVIAQASIVFMK 326
Query: 72 DFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
P GT++ T+++ W
Sbjct: 325 HSSSPC*QTTVPSCAIQALPQGTILRFTLEMRPVW 221
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,894,770
Number of Sequences: 28460
Number of extensions: 19067
Number of successful extensions: 121
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 121
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 121
length of query: 116
length of database: 4,897,600
effective HSP length: 78
effective length of query: 38
effective length of database: 2,677,720
effective search space: 101753360
effective search space used: 101753360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 49 (23.5 bits)
Medicago: description of AC148236.3


