

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
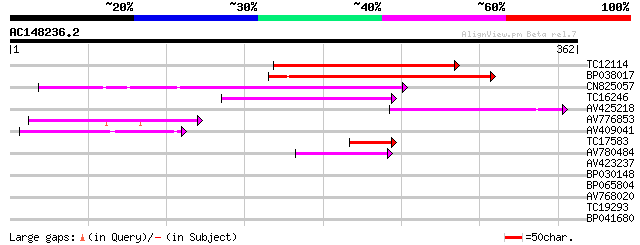
Sequences producing significant alignments: (bits) Value
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 207 3e-54
BP038017 128 2e-30
CN825057 124 3e-29
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 77 3e-15
AV425218 73 9e-14
AV776853 59 1e-09
AV409041 53 9e-08
TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired re... 52 2e-07
AV780484 43 7e-05
AV423237 34 0.044
BP030148 32 0.13
BP065804 28 1.8
AV768020 28 3.1
TC19293 28 3.1
BP041680 26 9.1
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 207 bits (526), Expect = 3e-54
Identities = 96/119 (80%), Positives = 107/119 (89%)
Frame = +1
Query: 169 KKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGD 228
K+R+RSL+ GEAGY+LQYFQRK VEN FYHAYQLD EDQITNVFW DARML+DY YFGD
Sbjct: 13 KRRQRSLLHGEAGYILQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFGD 192
Query: 229 VVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQK 287
+VSLDSTYCT+SS+RPLA+ SGFNHHR AVIFGAALLYDET ESY WLFE+FLEAHK +
Sbjct: 193 MVSLDSTYCTHSSNRPLAVFSGFNHHRKAVIFGAALLYDETTESY*WLFESFLEAHKHE 369
>BP038017
Length = 570
Score = 128 bits (321), Expect = 2e-30
Identities = 64/145 (44%), Positives = 93/145 (64%)
Frame = +2
Query: 166 YLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSY 225
Y++ ++R+L + +A LL YF++ +NP FY+A QLD E+++ NVFWADAR Y+Y
Sbjct: 131 YVKSSQKRTLGR-DAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAYNY 307
Query: 226 FGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHK 285
FGD V D+ Y N P A +G NHH V+FG ALL DE+ S+ WLF T+L A
Sbjct: 308 FGDAVIFDTMYRPNQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTWLFRTWLSAMN 487
Query: 286 QKMPQTVFTDQAKSMAKALAEVMPE 310
+ P ++ TDQ +++ A+A+V PE
Sbjct: 488 DRPPVSITTDQDRAIQAAVAQVFPE 562
>CN825057
Length = 721
Score = 124 bits (310), Expect = 3e-29
Identities = 69/238 (28%), Positives = 126/238 (51%), Gaps = 2/238 (0%)
Frame = +1
Query: 19 AWEFWNTYXGKXGFGVR-KDYTHRKKDGSVSSCRFVCCKEGMRKPDKRDYKTKNPRLETR 77
A+ F+ Y GF K+ KK +F C + G+ P+ + ++ +
Sbjct: 1 AYSFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVT-PESDGGSNRRSSVK-K 174
Query: 78 TNCEARLGLKNV-DGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVG 136
T+C+A + +K DGK ++H+F+++HNHELL P + R V +++ +V
Sbjct: 175 TDCKACMHVKRKPDGKWIIHEFIKEHNHELL-PALAYHFRIHRNVKLAEKNNMDILHAVS 351
Query: 137 VQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPT 196
+ +K + +S++ GG N+ + D + +K + ++ +G+A +L+YF+ ENP
Sbjct: 352 ERTRKMYVEMSRQSGGCLNIESLVGDLNDQFKKGQYLAMDEGDAQVMLEYFKHIQKENPN 531
Query: 197 FYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHH 254
F+++ L+ E ++ N+FW DA+ + DY F DVVS D++Y ++ P A G NHH
Sbjct: 532 FFYSIDLNEEQRLRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVGVNHH 705
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 77.4 bits (189), Expect = 3e-15
Identities = 35/112 (31%), Positives = 60/112 (53%)
Frame = +2
Query: 136 GVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENP 195
GV+ L+ GG +LGF + D N++ K + + G+A L Y + K +P
Sbjct: 20 GVRACHIMALMLGPKGGHESLGFTKTDLSNHIAKPKRERIQNGDAAAALSYLEGKADNDP 199
Query: 196 TFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAI 247
F++ + ++ + N+FW D +DY+ FGDV++ DSTY N ++PL +
Sbjct: 200 MFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYNKPLVV 355
>AV425218
Length = 419
Score = 72.8 bits (177), Expect = 9e-14
Identities = 39/114 (34%), Positives = 62/114 (54%)
Frame = +2
Query: 243 RPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAK 302
RPLA + G NHH +V+FG ALL E +ES+ WLF++ L PQ + TD +++M K
Sbjct: 2 RPLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGVPPQGIITDHSEAMKK 181
Query: 303 ALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
A+ V+P H C ++M+ + L + ES + +Y +++FE
Sbjct: 182 AIETVLPSTRHRWCLSYIMKKLPQKLLGYAQYES-IRHHLQNVVYDAVVIDEFE 340
>AV776853
Length = 625
Score = 58.9 bits (141), Expect = 1e-09
Identities = 40/115 (34%), Positives = 59/115 (50%), Gaps = 4/115 (3%)
Frame = +1
Query: 13 FCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMR--KPDKRDYKTK 70
F S + A++F+ Y GF VRKD R G+V+ +FVC ++G+R K R + +
Sbjct: 244 FGSEEEAYQFYQAYAKYQGFIVRKDDIGRDYHGNVNMRQFVCNRQGLRSKKHYNRTDRKR 423
Query: 71 NPRLETRTNCEA--RLGLKNVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSE 123
+ + T TNC A R+ L GK V F E HNHEL H + R +++
Sbjct: 424 DHKPVTHTNCLAKLRVHLDYKIGKWKVVSFEECHNHELTPARFVHFIPPYRVMND 588
>AV409041
Length = 349
Score = 52.8 bits (125), Expect = 9e-08
Identities = 35/109 (32%), Positives = 52/109 (47%), Gaps = 2/109 (1%)
Frame = +3
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGS-VSSCRFVCCKEGMRKPDKR 65
P D+ F S + A+ F+ Y GFG K + R + +F C + G ++
Sbjct: 33 PHYDIEFESHEAAYAFYKEYAKSAGFGTAKLSSRRSRASKEFIDAKFSCIRYGNKQQSD- 209
Query: 66 DYKTKNPRLETRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTH 113
NPR + C+A + +K DGK V+ FV++HNHE LLP H
Sbjct: 210 --DAINPRPSPKIGCKASMHVKRRQDGKWYVYSFVKEHNHE-LLPAQAH 347
>TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired response
protein, mutator-like transposase-like protein,
phytochrome A signaling protein-like, partial (3%)
Length = 666
Score = 52.0 bits (123), Expect = 2e-07
Identities = 21/30 (70%), Positives = 28/30 (93%)
Frame = +3
Query: 218 RMLVDYSYFGDVVSLDSTYCTNSSHRPLAI 247
RM++DY YFGDVVSLD+TY TN+++RPLA+
Sbjct: 3 RMIMDYEYFGDVVSLDTTYSTNNAYRPLAV 92
>AV780484
Length = 529
Score = 43.1 bits (100), Expect = 7e-05
Identities = 20/62 (32%), Positives = 32/62 (51%)
Frame = -1
Query: 183 LLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSH 242
+ +YF ENP F++A LD+ +T+VFW D + +DY +V + + Y N
Sbjct: 199 MTEYFVSTQGENPNFFYAIDLDLNRHLTSVFWVDIKGRLDYETSMMLVLIHTHYLKNKYK 20
Query: 243 RP 244
P
Sbjct: 19 IP 14
>AV423237
Length = 310
Score = 33.9 bits (76), Expect = 0.044
Identities = 21/62 (33%), Positives = 33/62 (52%)
Frame = +2
Query: 215 ADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYE 274
ADA ++YSYFGD V D+T ++ + G + VI+ AL+ +E+ S+
Sbjct: 14 ADATSRMNYSYFGDAVIFDTT--IDNHIESICSSWG*SSWATCVIWLVALIANESESSFV 187
Query: 275 WL 276
WL
Sbjct: 188 WL 193
>BP030148
Length = 398
Score = 32.3 bits (72), Expect = 0.13
Identities = 15/48 (31%), Positives = 24/48 (49%), Gaps = 1/48 (2%)
Frame = +3
Query: 286 QKMPQ-TVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLM 332
+ MP+ T+ +D+ K + + P A+HG C HL + K N M
Sbjct: 6 ENMPRLTILSDRQKGIVDGVEANFPTAFHGFCMRHLSDSFRKEFNNTM 149
>BP065804
Length = 492
Score = 28.5 bits (62), Expect = 1.8
Identities = 20/65 (30%), Positives = 28/65 (42%)
Frame = -1
Query: 6 NPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKR 65
+P + C + W W + GF R+ R VS+CR C +G+R D
Sbjct: 231 SPASRILSCLFCHKWYSWGLLNSE-GFRWRRSSKGRI---IVSACRPSCNSDGLR*LDPX 64
Query: 66 DYKTK 70
DY TK
Sbjct: 63 DYATK 49
>AV768020
Length = 268
Score = 27.7 bits (60), Expect = 3.1
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = -2
Query: 98 FVEDHNHELLLPETTHMLSSQRKVSEIHC 126
F+ H+H L+ TTH SSQ + +C
Sbjct: 240 FLHPHHHHPLISPTTHRCSSQSTTTRWYC 154
>TC19293
Length = 472
Score = 27.7 bits (60), Expect = 3.1
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = +2
Query: 240 SSHRPLAIISGFNHHRGAVIFGAALLYDETAESY 273
SS P + GF+ A+ FG+ L+Y AESY
Sbjct: 119 SSRVPPGSVKGFSIQGCAMYFGSFLVYSVAAESY 220
>BP041680
Length = 525
Score = 26.2 bits (56), Expect = 9.1
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +1
Query: 20 WEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRF 52
WE + + G + +R+ +TH K +SS RF
Sbjct: 109 WEAFGSVLGARDYALREKFTHPIKSIHLSSMRF 207
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,649,056
Number of Sequences: 28460
Number of extensions: 89461
Number of successful extensions: 404
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 401
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 403
length of query: 362
length of database: 4,897,600
effective HSP length: 91
effective length of query: 271
effective length of database: 2,307,740
effective search space: 625397540
effective search space used: 625397540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148236.2


