

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148154.6 - phase: 0
(199 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
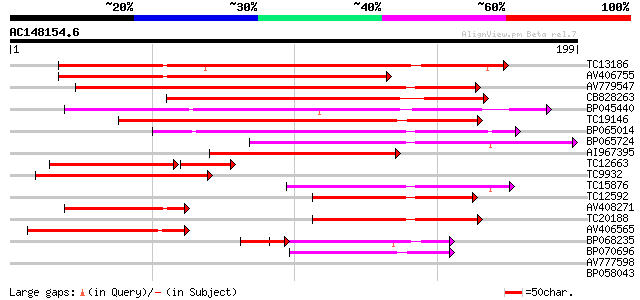
Sequences producing significant alignments: (bits) Value
TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidat... 138 6e-34
AV406755 129 3e-31
AV779547 124 1e-29
CB828263 118 8e-28
BP045440 108 8e-25
TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-contain... 103 3e-23
BP065014 98 8e-22
BP065724 75 1e-14
AI967395 71 1e-13
TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partia... 56 4e-13
TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like... 66 4e-12
TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 57 2e-09
TC12592 54 2e-08
AV408271 50 3e-07
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 49 6e-07
AV406565 47 2e-06
BP068235 40 9e-06
BP070696 40 4e-04
AV777598 36 0.004
BP058043 33 0.033
>TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (6%)
Length = 586
Score = 138 bits (348), Expect = 6e-34
Identities = 78/160 (48%), Positives = 105/160 (64%), Gaps = 2/160 (1%)
Frame = +1
Query: 18 FTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSL-IKAIEESRI 76
F Y VF++ SDTR GF+G LY AL+DKGI TFIDD L R D+I ++ KAIE SRI
Sbjct: 97 FIYDVFISLITSDTRFGFSGDLYTALSDKGILTFIDD-GLPRPDDIMSTVNSKAIESSRI 273
Query: 77 FIPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHE 136
I V S NYASS FCLDEL++I+ + KG L+ PVFY V+P+ +R+Q G +G+ HE
Sbjct: 274 AIVVISENYASSSFCLDELMYILKRREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASHE 453
Query: 137 ESFQNNKKNKERLHQWKLALTQAANLSGYH-YSPGYPTLL 175
E F + + ++ +W+ AL Q A+L +H + GY T L
Sbjct: 454 ERFGS---DSAKVRKWRNALKQVADLKAFHFFKHGYHTSL 564
>AV406755
Length = 425
Score = 129 bits (325), Expect = 3e-31
Identities = 64/117 (54%), Positives = 86/117 (72%)
Frame = +3
Query: 18 FTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIF 77
++Y VFL+FRG ++R FT HLY AL + GI F+D+ +L+RG++I+ SL+KAIE+SRI
Sbjct: 78 WSYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDN-ELQRGEDISSSLLKAIEDSRIA 254
Query: 78 IPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTK 134
I +FS NY SK+CLDEL II C +T G+ V+PVFY VDP+ IR Q G+ GE K
Sbjct: 255 IIIFSTNYTGSKWCLDELEKIIECQRTIGQEVMPVFYNVDPSDIRKQRGTVGEAFRK 425
>AV779547
Length = 556
Score = 124 bits (311), Expect = 1e-29
Identities = 69/142 (48%), Positives = 91/142 (63%)
Frame = +1
Query: 24 LNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIFIPVFSI 83
++FRG DTR+ FT HL AL KG TF DD L++G I+ LI+AIE S+I I VFS
Sbjct: 109 VSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNISTELIQAIEGSQILIVVFSK 288
Query: 84 NYASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNK 143
YASS +CL EL I K + VLPV V P+++R QSG+YGE KHEE F K
Sbjct: 289 YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQSGNYGEAFLKHEERF---K 459
Query: 144 KNKERLHQWKLALTQAANLSGY 165
++ E + +W+ AL + A+LSG+
Sbjct: 460 EDLEMVQRWRKALAEVADLSGW 525
>CB828263
Length = 570
Score = 118 bits (295), Expect = 8e-28
Identities = 55/113 (48%), Positives = 84/113 (73%)
Frame = +2
Query: 56 DLKRGDEITPSLIKAIEESRIFIPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVFYG 115
DL+RGDEI+ SL++AIEE+++ + VFS NY +SK+CLDELV I+ C +T+G++VLPVFY
Sbjct: 5 DLQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILECKRTRGQIVLPVFYD 184
Query: 116 VDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQAANLSGYHYS 168
+DP+ +R+Q+G+Y E KH + +++ +W+ AL +AANLSG+ S
Sbjct: 185 IDPSHVRNQTGTYAEAFVKHGQ--------VDKVQKWREALREAANLSGWDCS 319
>BP045440
Length = 535
Score = 108 bits (269), Expect = 8e-25
Identities = 66/173 (38%), Positives = 94/173 (54%), Gaps = 2/173 (1%)
Frame = -3
Query: 20 YQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIFIP 79
YQ+F++FRG DTRD FT LY+ L +G TF DD L+ G I L+ IEESR I
Sbjct: 527 YQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPI-EKLLDDIEESRFAII 351
Query: 80 VFSINYASSKFCLDELVHIIHCYKTKGR--LVLPVFYGVDPTQIRHQSGSYGEHLTKHEE 137
+ S NYA S++CL EL I+ C +G+ LVLP V PT++RH Y + + KHE+
Sbjct: 350 ILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKHEK 171
Query: 138 SFQNNKKNKERLHQWKLALTQAANLSGYHYSPGYPTLLKPCFSFLYFLYLIFF 190
N + + + WK AL L+G+ KP ++ ++ I+F
Sbjct: 170 ----NGIDSKTIQAWKKALFDVCTLTGFE---------KPSDWYVRSIFFIYF 51
>TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing protein
TSDC, partial (12%)
Length = 701
Score = 103 bits (256), Expect = 3e-23
Identities = 55/128 (42%), Positives = 79/128 (60%)
Frame = +1
Query: 39 LYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIFIPVFSINYASSKFCLDELVHI 98
LY AL +G TF+DD L G+EI SL+ A E+SR+ I VFS NY S +CLDELV I
Sbjct: 1 LYHALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKI 180
Query: 99 IHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQ 158
+ C +TK +LV P+FY V+ + + Q SY + HE +N K+ + + +W+ AL+
Sbjct: 181 VECMETKKQLVWPIFYKVEKSDVTSQENSYEHAMIGHE---KNYGKDSDEVKKWRSALSA 351
Query: 159 AANLSGYH 166
A L G++
Sbjct: 352 MAKLEGHN 375
>BP065014
Length = 498
Score = 98.2 bits (243), Expect = 8e-22
Identities = 58/129 (44%), Positives = 75/129 (57%)
Frame = -2
Query: 51 FIDDCDLKRGDEITPSLIKAIEESRIFIPVFSINYASSKFCLDELVHIIHCYKTKGRLVL 110
F DD L+ G I L+ AIEESR+ I V S N+A SK+CL ELV I+ C K K LVL
Sbjct: 497 FKDDLSLEDGAPIE-ELLGAIEESRVAIVVLSENFADSKWCLKELVKILDCRKKKKPLVL 321
Query: 111 PVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQAANLSGYHYSPG 170
P+FY V+PT IRH G YGE + +HE+ K+ + + +W AL+ + L G Y G
Sbjct: 320 PIFYKVEPTNIRHLKGRYGEAMAEHEKKL---GKDSQTVREWTQALSDISYLKGEPYK-G 153
Query: 171 YPTLLKPCF 179
Y L F
Sbjct: 152 YVVKLSITF 126
>BP065724
Length = 495
Score = 74.7 bits (182), Expect = 1e-14
Identities = 39/117 (33%), Positives = 68/117 (57%), Gaps = 2/117 (1%)
Frame = -1
Query: 85 YASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKK 144
YA S +CL+E+V I+ C + +LV P+FY V+P+ +RHQ SY + + K E + K
Sbjct: 495 YADSSWCLEEVVKILECKQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKQETRY---GK 325
Query: 145 NKERLHQWKLALTQAANLSGYHY--SPGYPTLLKPCFSFLYFLYLIFFPQNLILKLK 199
+ E++ +W+ +L+Q +L +HY + GY L + + + F ++LI+K K
Sbjct: 324 DSEKVKKWRSSLSQVCDLKAFHYKENSGYVLYLLLFKAMHNMINYLTF*KDLIVKNK 154
>AI967395
Length = 205
Score = 70.9 bits (172), Expect = 1e-13
Identities = 35/67 (52%), Positives = 46/67 (68%)
Frame = +3
Query: 71 IEESRIFIPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGE 130
I+ESR+ I VFS +YA S CL+ELV+I+ +LV PV+YGV+ +IRHQ G YGE
Sbjct: 3 IQESRLIIVVFSEHYAVSASCLNELVNIVDVMIMNNQLVWPVYYGVEAPEIRHQRGRYGE 182
Query: 131 HLTKHEE 137
+ K EE
Sbjct: 183 AMAKFEE 203
>TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partial (4%)
Length = 494
Score = 55.8 bits (133), Expect(2) = 4e-13
Identities = 27/45 (60%), Positives = 34/45 (75%)
Frame = +3
Query: 15 SYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKR 59
S+ T+ VFLNFRGSDT GF L K+L DKGI+TFI+D +L+R
Sbjct: 12 SHVLTFNVFLNFRGSDTCHGFISFLCKSLLDKGIYTFINDEELQR 146
Score = 33.5 bits (75), Expect(2) = 4e-13
Identities = 15/19 (78%), Positives = 17/19 (88%)
Frame = +2
Query: 61 DEITPSLIKAIEESRIFIP 79
DEITPSL+KAIE +RI IP
Sbjct: 149 DEITPSLVKAIERTRIAIP 205
>TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like, partial
(5%)
Length = 689
Score = 66.2 bits (160), Expect = 4e-12
Identities = 31/62 (50%), Positives = 43/62 (69%)
Frame = -2
Query: 10 SSSLISYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIK 69
SSS ++ + Y VF+NFRG DTR F HLY AL++ G++ F+DD +L+RG E+ P L K
Sbjct: 232 SSSSSNHQWIYDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPELQK 53
Query: 70 AI 71
I
Sbjct: 52 LI 47
>TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (13%)
Length = 558
Score = 57.4 bits (137), Expect = 2e-09
Identities = 27/82 (32%), Positives = 47/82 (56%), Gaps = 2/82 (2%)
Frame = +1
Query: 98 IIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALT 157
I+ C + +LV P+FY V+P+ +RHQ SY + + K E + K+ E++ +W+ +L+
Sbjct: 16 ILECXQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKPETRY---GKDSEKVKKWRSSLS 186
Query: 158 QAANLSGYHY--SPGYPTLLKP 177
Q +L +HY + GY P
Sbjct: 187 QVCDLKAFHYKENSGYERAFIP 252
>TC12592
Length = 534
Score = 53.9 bits (128), Expect = 2e-08
Identities = 26/58 (44%), Positives = 36/58 (61%)
Frame = +3
Query: 107 RLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQAANLSG 164
+LVLPVFYGV+PT IR+ G +GE + + E F K+ +H+WK AL + L G
Sbjct: 3 QLVLPVFYGVEPTDIRNLKGKFGEAMAEIENKF---GKDSNTIHEWKQALRDISYLKG 167
>AV408271
Length = 409
Score = 50.1 bits (118), Expect = 3e-07
Identities = 25/44 (56%), Positives = 29/44 (65%)
Frame = +1
Query: 20 YQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEI 63
Y VF++FRG D RDGF HL A K I+ F+DD LKRG EI
Sbjct: 280 YDVFVSFRGEDIRDGFLSHLADAFRQKKINAFMDD-KLKRGQEI 408
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 48.9 bits (115), Expect = 6e-07
Identities = 23/60 (38%), Positives = 37/60 (61%)
Frame = +1
Query: 107 RLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQAANLSGYH 166
+LV P+FY + + + +Q+ SYG + HE SF K+ E++ +WK AL++ A L G H
Sbjct: 4 QLVWPIFYEISXSDVSNQTKSYGHAMLAHENSF---GKDSEKVQKWKSALSEMAFLQGEH 174
>AV406565
Length = 207
Score = 47.4 bits (111), Expect = 2e-06
Identities = 27/57 (47%), Positives = 37/57 (64%)
Frame = +2
Query: 7 SYSSSSLISYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEI 63
++S++S + + VFL+FRG DTR TGHL LT + T+I D DL+RGDEI
Sbjct: 38 AWSTTSSSAPQQKHDVFLSFRGEDTRYTCTGHLQATLTRLQVKTYI-DYDLQRGDEI 205
>BP068235
Length = 511
Score = 39.7 bits (91), Expect(2) = 9e-06
Identities = 23/66 (34%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Frame = -1
Query: 92 LDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLT-KHEESFQNNKKNKERLH 150
L L + C K K +L+ P Y V+P+ IR++ SYG L ++F N + E++
Sbjct: 475 LMNLSRFLECKKMKNQLLWPCPYKVEPSDIRYRPSSYGTALA*TRTDAFGN---DSEKIQ 305
Query: 151 QWKLAL 156
WK AL
Sbjct: 304 TWKSAL 287
Score = 24.6 bits (52), Expect(2) = 9e-06
Identities = 12/17 (70%), Positives = 12/17 (70%)
Frame = -3
Query: 82 SINYASSKFCLDELVHI 98
S NYA S CLDELV I
Sbjct: 506 SENYAYSARCLDELVQI 456
>BP070696
Length = 429
Score = 39.7 bits (91), Expect = 4e-04
Identities = 18/58 (31%), Positives = 32/58 (55%)
Frame = -1
Query: 99 IHCYKTKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLAL 156
+ C K K ++ P+F ++ +I +Q SY + + HE + N K +R+ +WK AL
Sbjct: 429 LECKKKKNNIIRPIFQNMELLKIIYQINSYDDVMVAHE---RKNGKESDRVKKWKSAL 265
>AV777598
Length = 371
Score = 36.2 bits (82), Expect = 0.004
Identities = 16/23 (69%), Positives = 18/23 (77%)
Frame = +3
Query: 18 FTYQVFLNFRGSDTRDGFTGHLY 40
FTY VFL+F G DTR FTG+LY
Sbjct: 303 FTYDVFLSFCGVDTRFSFTGNLY 371
>BP058043
Length = 375
Score = 33.1 bits (74), Expect = 0.033
Identities = 16/34 (47%), Positives = 19/34 (55%)
Frame = -3
Query: 15 SYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGI 48
S+ FTY VF++ RG RD F K L KGI
Sbjct: 169 SHNFTYDVFISLRGEVLRDTFIECFLKELMQKGI 68
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,115,110
Number of Sequences: 28460
Number of extensions: 62605
Number of successful extensions: 454
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 446
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 446
length of query: 199
length of database: 4,897,600
effective HSP length: 86
effective length of query: 113
effective length of database: 2,450,040
effective search space: 276854520
effective search space used: 276854520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148154.6


