

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
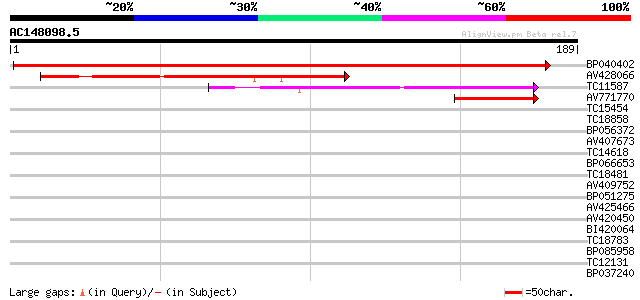
Sequences producing significant alignments: (bits) Value
BP040402 306 1e-84
AV428066 120 2e-28
TC11587 homologue to UP|Q9FN36 (Q9FN36) Gb|AAF34833.1, partial (... 96 4e-21
AV771770 39 4e-04
TC15454 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabi... 31 0.12
TC18858 29 0.45
BP056372 28 0.76
AV407673 28 1.3
TC14618 homologue to UP|AAQ76040 (AAQ76040) Ubiquitin extension ... 27 1.7
BP066653 27 2.2
TC18481 similar to UP|O23166 (O23166) Thiol-disulfide interchang... 27 2.9
AV409752 27 2.9
BP051275 26 3.8
AV425466 26 3.8
AV420450 26 3.8
BI420064 26 3.8
TC18783 similar to UP|Q9FNN8 (Q9FNN8) Cleft lip and palate assoc... 26 3.8
BP085958 26 4.9
TC12131 similar to UP|Q9LN41 (Q9LN41) F18O14.29, partial (17%) 26 4.9
BP037240 26 4.9
>BP040402
Length = 596
Score = 306 bits (784), Expect = 1e-84
Identities = 135/179 (75%), Positives = 159/179 (88%)
Frame = +3
Query: 2 ESGLPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKG 61
E+GLPLL+ LLQ TLR +C+ P STSSKWVYAVFWR++PR FPPPRWEFGGT+LDRSKG
Sbjct: 9 ENGLPLLNCLLQQTLRTICTSPNFSTSSKWVYAVFWRILPRNFPPPRWEFGGTALDRSKG 188
Query: 62 NKRNWIIVWEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGH 121
NKRNWI+VWEDGFCDF+ECEQR+SG LN RFGADVFFKMSHEVYSYGEGL+GKVAAD H
Sbjct: 189 NKRNWILVWEDGFCDFDECEQRRSGYLNSRFGADVFFKMSHEVYSYGEGLLGKVAADTSH 368
Query: 122 KWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
KWVY+D + CE +Y+ WNAS++ QP+AWEFQ NSGIQ+I +IAV+EG+VQLGSF+K+
Sbjct: 369 KWVYNDQHSECEPSYIASWNASVEPQPKAWEFQFNSGIQSIVLIAVREGVVQLGSFNKI 545
>AV428066
Length = 429
Score = 120 bits (300), Expect = 2e-28
Identities = 59/118 (50%), Positives = 77/118 (65%), Gaps = 15/118 (12%)
Frame = +1
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC +S+WVYAVFWR++PR +PPP+WE G + DRS+GN+RNWI+VW
Sbjct: 91 LLQHTLRSLCIHE----NSQWVYAVFWRILPRNYPPPKWE-GQGAYDRSRGNRRNWILVW 255
Query: 71 EDGFCDFNEC---EQRKSGCLN------------ERFGADVFFKMSHEVYSYGEGLVG 113
EDGFC+F E C + ++FFKMSHE+Y+YGEGL+G
Sbjct: 256 EDGFCNFAASAAPEVNSGDCPTSSVYGNCEFQPYQGLQPELFFKMSHEIYNYGEGLIG 429
>TC11587 homologue to UP|Q9FN36 (Q9FN36) Gb|AAF34833.1, partial (36%)
Length = 697
Score = 95.9 bits (237), Expect = 4e-21
Identities = 53/114 (46%), Positives = 68/114 (59%), Gaps = 4/114 (3%)
Frame = +3
Query: 67 IIVWEDGFCDFNECEQRKSGCLNERFGAD----VFFKMSHEVYSYGEGLVGKVAADNGHK 122
+++WEDGFC + L++ G D VF KMS ++Y+YGEGL+GKVA+D HK
Sbjct: 18 MLMWEDGFC--------RGRGLDDIDGEDPVRKVFSKMSIQLYNYGEGLMGKVASDKCHK 173
Query: 123 WVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGS 176
WV+ + CE N W +S D P W Q SGIQTIAVI GL+QLGS
Sbjct: 174 WVFKEPTE-CEPNISNYWQSSFDALPPEWTDQFESGIQTIAVIQAGHGLLQLGS 332
>AV771770
Length = 288
Score = 39.3 bits (90), Expect = 4e-04
Identities = 18/28 (64%), Positives = 23/28 (81%)
Frame = +3
Query: 149 RAWEFQLNSGIQTIAVIAVKEGLVQLGS 176
R WE Q SGI+TIA+IAV+EG+VQ G+
Sbjct: 3 RTWEAQFLSGIKTIALIAVREGVVQSGA 86
>TC15454 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;}, partial (5%)
Length = 544
Score = 31.2 bits (69), Expect = 0.12
Identities = 11/16 (68%), Positives = 15/16 (93%)
Frame = +1
Query: 95 DVFFKMSHEVYSYGEG 110
++FFKMSHE+Y+Y EG
Sbjct: 13 ELFFKMSHEIYNYEEG 60
>TC18858
Length = 409
Score = 29.3 bits (64), Expect = 0.45
Identities = 13/19 (68%), Positives = 16/19 (83%)
Frame = +1
Query: 6 PLLHSLLQHTLRNLCSFPT 24
PL H+L HTL++LCSFPT
Sbjct: 127 PLSHNL--HTLKSLCSFPT 177
>BP056372
Length = 632
Score = 28.5 bits (62), Expect = 0.76
Identities = 9/28 (32%), Positives = 16/28 (57%)
Frame = +2
Query: 47 PRWEFGGTSLDRSKGNKRNWIIVWEDGF 74
P W+F GT + +G K ++++W F
Sbjct: 518 PTWDFDGTKSSK*RGLKATFVMLWSPAF 601
>AV407673
Length = 427
Score = 27.7 bits (60), Expect = 1.3
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = +3
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPR 42
L LR +C +S W Y+VFW + PR
Sbjct: 318 LHEALRTVC------LNSDWTYSVFWTIRPR 392
>TC14618 homologue to UP|AAQ76040 (AAQ76040) Ubiquitin extension protein,
partial (97%)
Length = 644
Score = 27.3 bits (59), Expect = 1.7
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -1
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVF 36
+P HS L H+ +L +FP SST W A F
Sbjct: 419 VPAPHSALGHSFLSL*TFPESSTL*NWRTASF 324
>BP066653
Length = 513
Score = 26.9 bits (58), Expect = 2.2
Identities = 10/27 (37%), Positives = 15/27 (55%)
Frame = +3
Query: 24 TSSTSSKWVYAVFWRVVPRIFPPPRWE 50
T+S W+Y + + P IFP +WE
Sbjct: 378 TNSFVFLWIYYISPNIFPTIFPRSKWE 458
>TC18481 similar to UP|O23166 (O23166) Thiol-disulfide interchange like
protein (Thioredoxin-like protein)
(AT4g37200/C7A10_160), partial (42%)
Length = 602
Score = 26.6 bits (57), Expect = 2.9
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -3
Query: 99 KMSHEVYSYGEGLVGKVAADNGHKWVYSD 127
K S +Y YG ++ + A+NGH + SD
Sbjct: 573 KNSRHLYLYGNFMIIDIGANNGHGYRVSD 487
>AV409752
Length = 421
Score = 26.6 bits (57), Expect = 2.9
Identities = 18/53 (33%), Positives = 23/53 (42%), Gaps = 3/53 (5%)
Frame = -2
Query: 45 PPPRWEFGGTSLDRSKGNKRNWIIVWEDGFCDFNE---CEQRKSGCLNERFGA 94
PPP W GG+ + + EDGF D E +R+ CL FGA
Sbjct: 150 PPPVWSLGGSDTGSMRFSDE-----LEDGFRDIMEKTRRRRRRGFCLLFLFGA 7
>BP051275
Length = 295
Score = 26.2 bits (56), Expect = 3.8
Identities = 9/21 (42%), Positives = 14/21 (65%), Gaps = 1/21 (4%)
Frame = -3
Query: 30 KWVYAVFWRVVPRI-FPPPRW 49
KW++ + ++P FPPPRW
Sbjct: 248 KWIFVLSLALLPAAYFPPPRW 186
>AV425466
Length = 345
Score = 26.2 bits (56), Expect = 3.8
Identities = 7/18 (38%), Positives = 13/18 (71%)
Frame = +1
Query: 59 SKGNKRNWIIVWEDGFCD 76
SK ++ W+++W+DG D
Sbjct: 10 SKAEQQQWVLLWQDGTLD 63
>AV420450
Length = 413
Score = 26.2 bits (56), Expect = 3.8
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +2
Query: 36 FWRVVPRIFPPPRWEFGGTSLDR 58
FW+ +PR+FPP F +L+R
Sbjct: 92 FWKPLPRMFPPDSPFFAPGNLER 160
>BI420064
Length = 410
Score = 26.2 bits (56), Expect = 3.8
Identities = 15/32 (46%), Positives = 18/32 (55%)
Frame = +1
Query: 1 MESGLPLLHSLLQHTLRNLCSFPTSSTSSKWV 32
+ S P HS L + +LC FP SST S WV
Sbjct: 13 LSSCYPHPHSNLHSS--SLCVFPISSTFSTWV 102
>TC18783 similar to UP|Q9FNN8 (Q9FNN8) Cleft lip and palate associated
transmembrane protein-like, partial (20%)
Length = 406
Score = 26.2 bits (56), Expect = 3.8
Identities = 14/47 (29%), Positives = 24/47 (50%)
Frame = +3
Query: 3 SGLPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRW 49
S +P S+L+ + LC+FP + + V +++ RI PRW
Sbjct: 180 SRIPASASILEGHILCLCTFPRAKRIKRRVCWQVQQILLRIKRQPRW 320
>BP085958
Length = 507
Score = 25.8 bits (55), Expect = 4.9
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = +1
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVP 41
LPL+ SLL T+ L S SS SS +Y R P
Sbjct: 142 LPLIFSLLLSTILILLSIFVSSDSSSLLYLYRSRATP 252
>TC12131 similar to UP|Q9LN41 (Q9LN41) F18O14.29, partial (17%)
Length = 480
Score = 25.8 bits (55), Expect = 4.9
Identities = 7/15 (46%), Positives = 11/15 (72%)
Frame = +3
Query: 59 SKGNKRNWIIVWEDG 73
S N + W+++WEDG
Sbjct: 9 SASNNQQWVLLWEDG 53
>BP037240
Length = 484
Score = 25.8 bits (55), Expect = 4.9
Identities = 14/21 (66%), Positives = 15/21 (70%)
Frame = +3
Query: 9 HSLLQHTLRNLCSFPTSSTSS 29
HSLLQ L NL S P +STSS
Sbjct: 198 HSLLQTPLPNLPSPPPTSTSS 260
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,573,187
Number of Sequences: 28460
Number of extensions: 79560
Number of successful extensions: 346
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 341
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 341
length of query: 189
length of database: 4,897,600
effective HSP length: 85
effective length of query: 104
effective length of database: 2,478,500
effective search space: 257764000
effective search space used: 257764000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148098.5


