

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147960.5 - phase: 0
(617 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
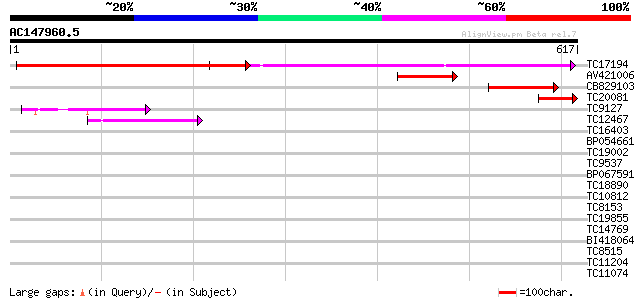
Score E
Sequences producing significant alignments: (bits) Value
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 247 3e-66
AV421006 64 9e-11
CB829103 64 9e-11
TC20081 similar to UP|AAQ73157 (AAQ73157) LysM domain-containing... 50 1e-06
TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-... 42 2e-04
TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete 40 8e-04
TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein... 40 0.001
BP054661 39 0.002
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 35 0.036
TC9537 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 32 0.23
BP067591 32 0.39
TC18890 similar to UP|Q7YS88 (Q7YS88) Receptor activity modifyin... 30 0.87
TC10812 similar to UP|O65440 (O65440) CLV1 receptor kinase like ... 30 0.87
TC8153 weakly similar to UP|Q8VAJ3 (Q8VAJ3) Wsv416 (WSSV475), pa... 30 1.1
TC19855 similar to UP|Q84Y10 (Q84Y10) Carbonic anhydrase 2 , pa... 30 1.1
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 29 1.9
BI418064 29 2.5
TC8515 28 3.3
TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Ar... 28 4.3
TC11074 28 4.3
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 247 bits (631), Expect = 3e-66
Identities = 187/404 (46%), Positives = 230/404 (56%), Gaps = 6/404 (1%)
Frame = +3
Query: 218 VPFHTRSIRWSYNWNICWSSSRTNSIIILHLCYVLPKEEDKEARISFGRIQRDLRSS*E* 277
VP + RS W W I + R + IL++C + P+E +E+ I+ +
Sbjct: 777 VPQNRRSS*WCSCWYIYCRNLRASVTSILYVC*I-PEEGRRES*IANRYFYGPFNT--RW 947
Query: 278 *SLR*CYIWNF--RFCESC*-YDWN*--GGKIRRVFI*RTSQCHK*LQHG*QNRSRWFR* 332
* L *C I NF ++ C* Y * GG+I V I TS+ +K*L G* N SRW
Sbjct: 948 *CL**CRI*NFWIQWARDC*CYRSY*HYGGEINGVLISGTSEGYK*L*LG**NWSRWIWS 1127
Query: 333 SLLRRAEWRESCNKKDGHESNKRISC*VEGVDACSSREPGTLDRILC*GIPFPCL*IH** 392
LL R E +E+ N +DG S RIS *VEG++ CS E G LD IL *GI PCL* +*
Sbjct: 1128 CLLCRIERQENSN*EDGCTSINRISL*VEGLNTCSPLESGALDWILR*GISIPCL*TY*Q 1307
Query: 393 WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLC 452
WKLRP FR RT MV N S S+R *IHS + +Y SR ++ K+ R +L
Sbjct: 1308 WKLRPIFAWFR*RTIAMV*PSTNSSRCSKRP*IHS*AHCACVYPSRCEICKHIDR*ELAW 1487
Query: 453 KGCRLRINQAD*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFWSSSL*TNFC* 511
KGCR ++QA * W +++ G +IW+HA RIC IW* F KNRCIC WS S *T FC
Sbjct: 1488 KGCRFWLDQAY*SWELHTTNS-SGGNIWIHAPRICSIW*YFSKNRCICIWSCSF*TYFCK 1664
Query: 512 SSCNNG*RFWC*FKGPCSFV*RSF*SATSYRRS*KTGGS*AWR*LSN*SCIQDGTTCESM 571
C+ C* KGPCSFV*RS * S S +TGGS*AWR LSN* C QD TT ES+
Sbjct: 1665 ECCSEDR*ISC*IKGPCSFV*RST**E*SL*CSSQTGGS*AWRKLSN*FCSQDCTTRESL 1844
Query: 572 YK**SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNV 615
YK *S A +KYE CSY Y*GL +++KS S K V
Sbjct: 1845 YKR*STAKTKYEIFSCCSYDPFITY*GL***IFLRKSNSHKFTV 1976
Score = 236 bits (602), Expect = 8e-63
Identities = 121/258 (46%), Positives = 172/258 (65%), Gaps = 3/258 (1%)
Frame = +1
Query: 8 LSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTN-LTYVSNIMQSNLVTKPEDIVSY 66
L F +LL F ES C K C++ALASYY+ L ++ MQS +V+ + I SY
Sbjct: 136 LLFFILLLGHVCFHVESNCLKGCDLALASYYILPGVFILQNITTFMQSEIVSSNDAITSY 315
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N D I N +QSF R+N+PFPCDCI EFLGH+F+Y + DTY ++A+ Y+NLTT +
Sbjct: 316 NKDKILNDINIQSFQRLNIPFPCDCIGGEFLGHVFEYSASKGDTYETIANLYYANLTTVD 495
Query: 127 WLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEID 186
L+ FNSY +IP +NVTVNCSCGNS VSKDYGLFITYP+RP D+L+ I+N++ +D
Sbjct: 496 LLKRFNSYDPKNIPVNAKVNVTVNCSCGNSQVSKDYGLFITYPIRPGDTLQDIANQSSLD 675
Query: 187 AELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNWNICWSSSRTNSIIIL 246
A L+Q +NP VNFS+ SG+ +IPG+ +N YVP + R+ + + S + T +++L
Sbjct: 676 AGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLLLL 855
Query: 247 HLC-YV-LPKEEDKEARI 262
C YV K+E+++A++
Sbjct: 856 AFCMYVRYQKKEEEKAKL 909
>AV421006
Length = 203
Score = 63.5 bits (153), Expect = 9e-11
Identities = 33/65 (50%), Positives = 40/65 (60%)
Frame = +1
Query: 423 S*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNFISSHC*YGWHIWLH 482
S*I+S Y IYSS HKV KY RQKL +GC I+Q W F++SH W IW+H
Sbjct: 1 S*IYS*AYSACIYSS*HKVSKYINRQKLPGEGC*FWIDQTHRSWKFLASHWSSCWDIWIH 180
Query: 483 ATRIC 487
AT +C
Sbjct: 181 ATGVC 195
>CB829103
Length = 547
Score = 63.5 bits (153), Expect = 9e-11
Identities = 41/77 (53%), Positives = 48/77 (62%), Gaps = 1/77 (1%)
Frame = +3
Query: 522 C*FKGPCSFV*RSF*S-ATSYRRS*KTGGS*AWR*LSN*SCIQDGTTCESMYK**SAATS 580
C* PCSFV* S+* S+ RS*KTGGS AWR L N* DG+TC+S+++ S TS
Sbjct: 69 C*V*EPCSFV**SY*P*GESH*RS*KTGGSEAWRELLN*FHS*DGSTCQSLHRPRSKTTS 248
Query: 581 KYEFCCSCSYHTNFNY* 597
E CC CS F Y*
Sbjct: 249 TNEICCGCSNGFKFCY* 299
>TC20081 similar to UP|AAQ73157 (AAQ73157) LysM domain-containing
receptor-like kinase 6, partial (9%)
Length = 506
Score = 49.7 bits (117), Expect = 1e-06
Identities = 23/42 (54%), Positives = 27/42 (63%)
Frame = +3
Query: 576 SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNVWK 617
S SKYE C C Y T FN * LG + ++KSKSC+S VWK
Sbjct: 24 STTASKYEIHCGCPYDTFFNP*RLGCWLLLRKSKSCESYVWK 149
>TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-anchored
protein 2 precursor, partial (46%)
Length = 1016
Score = 42.4 bits (98), Expect = 2e-04
Identities = 40/151 (26%), Positives = 65/151 (42%), Gaps = 11/151 (7%)
Frame = +1
Query: 14 LLASKSFIAESKC---SKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYNTDT 70
LLA A+ KC + C+ ALA Y + T L + + V Y D
Sbjct: 379 LLAEAQIEAKFKCISENAPCH-ALADYSHPNGTTLRRIQTLFT----------VKYLPDI 525
Query: 71 ITNKDFVQSFTRV------NVPFPCDCIHDEFLGH-IFQYQVATKDTYLSVASNNYSNLT 123
+ + + TRV VPFPC C + L + + +Y++ DT +A+ ++ L
Sbjct: 526 LGANNLPANTTRVAPDQVIKVPFPCRCSNGTGLSNKVPRYKIKKGDTLYDIATTVFAGLV 705
Query: 124 TSEWLQNFNSYP-SNDIPDTGTLNVTVNCSC 153
+Q N P +N+I T+ + + CSC
Sbjct: 706 KYPQIQVANEIPDANNITAGDTIWIPLPCSC 798
>TC12467 UP|CAE02597 (CAE02597) Nod-factor receptor 5, complete
Length = 2292
Score = 40.4 bits (93), Expect = 8e-04
Identities = 35/129 (27%), Positives = 59/129 (45%), Gaps = 4/129 (3%)
Frame = +3
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFN-SYPSNDIPDTG 143
VP C C + + YQ+ D+Y VA+ Y NLT +Q N +P+
Sbjct: 429 VPVTCGCAGNHSSANT-SYQIQLGDSYDFVATTLYENLTNWNIVQASNPGVNPYLLPERV 605
Query: 144 TLNVTVNCSC-GNSDVSKDYGLFITYPLRPEDSLELISNKTEID-AELLQKYNPGVNFSQ 201
+ + C C + ++K ITY +P D++ L+S K A++L + G +F+
Sbjct: 606 KVVFPLFCRCPSKNQLNKGIQYLITYVWKPNDNVSLVSAKFGASPADILTENRYGQDFTA 785
Query: 202 GSGL-VYIP 209
+ L + IP
Sbjct: 786 ATNLPILIP 812
>TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, partial
(55%)
Length = 876
Score = 40.0 bits (92), Expect = 0.001
Identities = 32/105 (30%), Positives = 48/105 (45%), Gaps = 1/105 (0%)
Frame = +2
Query: 404 WRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD 463
W + +M + *NC SQR+*I R Y S H + ++ + CK C ++
Sbjct: 26 WFSSLMDSES*NCCWSSQRT*ISP*EGRDTYYPSLH*I**HTAFR*RRCKDC*F*FVKSS 205
Query: 464 *CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFWSSSL*T 507
+ SS W++ L +RIC W FK C+ FWS + T
Sbjct: 206 PRCSSTSSFYPCSWNLRLSCSRICNDWATHFKK*CLQFWSCTFGT 340
>BP054661
Length = 449
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/47 (48%), Positives = 27/47 (56%)
Frame = -2
Query: 571 MYK**SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNVWK 617
+++ *S TSKYE C CS T F LG + KS SCKSNV K
Sbjct: 448 VHRA*SNGTSKYEICYGCSNDT*FYNSKLGPCILL*KSSSCKSNVGK 308
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 35.0 bits (79), Expect = 0.036
Identities = 56/199 (28%), Positives = 80/199 (40%), Gaps = 14/199 (7%)
Frame = +3
Query: 303 KIRRVFI*RTSQCHK*LQHG*QNRSRWFR*SLLRRA-EWRESCNKKD---GHESNKRISC 358
++ VF R + C+K* Q *Q RW R +L A W +C++K +S I C
Sbjct: 15 RMEGVFFERIALCYK*FQL**QAW*RWIRQCILGPALGWITNCSEKIESLEQQSGHGICC 194
Query: 359 *VEGVDACSSREPGTLDRILC*-GIPFPCL*IH**WKLRPTSTQFRWRTFVMV------- 410
* + +++E +LC* +*++ +L PT + W + +
Sbjct: 195 *S*DIGKSTTQESS*SSWLLC*RSGTVDRI*LY--AELEPTLSS-SWTALIRMPS*LEST 365
Query: 411 --YQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNF 468
Y C G S + TY P Y S+ V I +L GC Q D W
Sbjct: 366 DEYCNRVCRGNCISSPPSNTTYHPQRY*SKQCV----IGFRLPGSGC*FWFRQVDP*WCN 533
Query: 469 ISSHC*YGWHIWLHATRIC 487
H G H WL TRIC
Sbjct: 534 SCDHKSQG-HPWLPCTRIC 587
>TC9537 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 480
Score = 32.3 bits (72), Expect = 0.23
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 4/62 (6%)
Frame = +3
Query: 430 YRPYIYSSRHKVRKYSIRQKLLCKGCRL----RINQAD*CWNFISSHC*YGWHIWLHATR 485
+ PY SSR+K +++ + CK C +N+ + +S + W IWLH TR
Sbjct: 96 FTPYC-SSRYKDKQHPFGYWIQCKSC*FWSG*DVNEVGAI*HHVSCY----WLIWLHGTR 260
Query: 486 IC 487
IC
Sbjct: 261 IC 266
>BP067591
Length = 358
Score = 31.6 bits (70), Expect = 0.39
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +1
Query: 211 KDQNRNYVPFHTRSIRWSYNWNICWSSSRTNS 242
+D RN+ H I W+Y W CW+ R N+
Sbjct: 19 QDHTRNFNHLHRCCIWWTYFWI*CWNCRRCNN 114
>TC18890 similar to UP|Q7YS88 (Q7YS88) Receptor activity modifying protein
3, partial (9%)
Length = 525
Score = 30.4 bits (67), Expect = 0.87
Identities = 26/75 (34%), Positives = 36/75 (47%), Gaps = 14/75 (18%)
Frame = -1
Query: 449 KLLCKGCRLRINQAD*CWNFISSHC*YGWHI--------WLHATRICI------W*CFFK 494
+++C G RL + + C +ISSH GW + WL A R+C * F
Sbjct: 228 RVVCGGGRLPMGASRACLAWISSHFTVGWDLG*QEKEGEWLSA-RVCESEKSK*V*VFST 52
Query: 495 NRCICFWSSSL*TNF 509
NRC WS+S+ NF
Sbjct: 51 NRCKT-WSTSII*NF 10
>TC10812 similar to UP|O65440 (O65440) CLV1 receptor kinase like protein,
partial (14%)
Length = 671
Score = 30.4 bits (67), Expect = 0.87
Identities = 15/48 (31%), Positives = 27/48 (56%), Gaps = 4/48 (8%)
Frame = +2
Query: 466 WNFISSHC*YGWHIWLHATRICIW*CFFKNR----CICFWSSSL*TNF 509
W+F H + W +WLH++R+CI ++R C+ WS + T++
Sbjct: 65 WDF-RVHVFHCWFLWLHSSRVCI---HIESR*EK*CL*LWSGAARTHY 196
>TC8153 weakly similar to UP|Q8VAJ3 (Q8VAJ3) Wsv416 (WSSV475), partial
(18%)
Length = 476
Score = 30.0 bits (66), Expect = 1.1
Identities = 21/64 (32%), Positives = 32/64 (49%), Gaps = 4/64 (6%)
Frame = +1
Query: 32 IALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQS----FTRVNVPF 87
+A S Q TN+TYVS++ ++ +++ ITN F S F++V VP
Sbjct: 58 VAFLSSSAQYLTNITYVSSLCSTH--------AHHSSPIITNIFFFFSPPPYFSQVAVPT 213
Query: 88 PCDC 91
PC C
Sbjct: 214 PCSC 225
>TC19855 similar to UP|Q84Y10 (Q84Y10) Carbonic anhydrase 2 , partial (27%)
Length = 642
Score = 30.0 bits (66), Expect = 1.1
Identities = 17/53 (32%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = +1
Query: 126 EWLQNFNSYPSNDIPDTGTLNVTVNC-SCGNSDVSKDYGLFITYPLRPEDSLE 177
+W+Q N S DTG+L+ + C +C V+ G +TYP E L+
Sbjct: 34 QWVQICNPAKSKVKTDTGSLSFSEQCTNCEKEAVNVSLGNLLTYPFVKERVLD 192
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +2
Query: 416 CSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCR 456
CS S RS I + R + + HK+++Y+ +C GCR
Sbjct: 2474 CSWCSSRSGISAHISRTFCNTQGHKIKQYTSGS*HVC*GCR 2596
>BI418064
Length = 667
Score = 28.9 bits (63), Expect = 2.5
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Frame = +3
Query: 3 PIKFRLSFLFMLLASKSFIAESKCSKTC--NIALASYYLQDDTNL 45
P + L+ L M LAS + CSKTC N+ L S Y + T +
Sbjct: 291 PATYHLTLLPMKLASHTQFHAPWCSKTCHLNLPLTSLYAKKSTRV 425
>TC8515
Length = 627
Score = 28.5 bits (62), Expect = 3.3
Identities = 20/58 (34%), Positives = 28/58 (47%)
Frame = -3
Query: 91 CIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVT 148
CIHD+F+G+ + Y+ ++NNY T EW N NS+ N T T T
Sbjct: 616 CIHDDFIGN---QPLNV*KKYVQSSNNNYI-YTR*EW--NINSFEDN*HAATTTTTTT 461
>TC11204 similar to GB|AAO64909.1|29029060|BT005974 At1g77410 {Arabidopsis
thaliana;}, partial (13%)
Length = 568
Score = 28.1 bits (61), Expect = 4.3
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +2
Query: 200 SQGSGLVYIPGKDQNRNYVPFHTRSIRWSYNW 231
S G G+ ++ G+ R +V FHT S NW
Sbjct: 242 SMGKGITWVNGQGIGRYWVSFHTPDGTSSQNW 337
>TC11074
Length = 759
Score = 28.1 bits (61), Expect = 4.3
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = -3
Query: 125 SEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSD 157
S WLQNF Y S DT + + + CSC + +
Sbjct: 232 STWLQNFQ*YSSRTKSDTVYVPLGL*CSCSSDN 134
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,538,892
Number of Sequences: 28460
Number of extensions: 234648
Number of successful extensions: 2323
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 2276
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2313
length of query: 617
length of database: 4,897,600
effective HSP length: 96
effective length of query: 521
effective length of database: 2,165,440
effective search space: 1128194240
effective search space used: 1128194240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147960.5


