

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
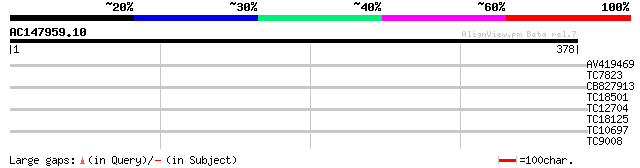
Score E
Sequences producing significant alignments: (bits) Value
AV419469 31 0.30
TC7823 similar to UP|O24470 (O24470) Lipoxygenase , partial (96%) 29 1.5
CB827913 28 3.3
TC18501 27 4.3
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 27 5.6
TC18125 similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X carboxypep... 26 9.5
TC10697 26 9.5
TC9008 homologue to PIR|T06773|T06773 ferredoxin-NADP reductase ... 26 9.5
>AV419469
Length = 305
Score = 31.2 bits (69), Expect = 0.30
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +2
Query: 1 DGRRVQDFFLSMEVAIRNHPLWA 23
D VQ+F +ME A + HPLWA
Sbjct: 236 DSAAVQEFLANMEAAFKAHPLWA 304
>TC7823 similar to UP|O24470 (O24470) Lipoxygenase , partial (96%)
Length = 2909
Score = 28.9 bits (63), Expect = 1.5
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 3/66 (4%)
Frame = -3
Query: 85 LHNEASWLLAEKELQKINAFKAPQEKLSTIMNC--CRVI-NNLLLNAAMSEYVPAGADDF 141
LH WL+A+ + ++ F P LS +NC C VI ++LLN + + +
Sbjct: 2127 LHLRPPWLVAQVSMTSLHNFFPPSL*LSVFLNCIICWVI*RDILLNPLRNSIPNL*SINS 1948
Query: 142 IPVLIY 147
I V++Y
Sbjct: 1947 IRVVLY 1930
>CB827913
Length = 567
Score = 27.7 bits (60), Expect = 3.3
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 16/90 (17%)
Frame = -3
Query: 148 VTIKARLALLVQTSNSLSCIEGKQNSSQKLSIISQILCQLRH--------------L*LN 193
+TI+ L LL+ +S C + L + Q L QL+H +
Sbjct: 340 LTIQLHL*LLIIQLDSCCCHNPQPQEESSLHFLFQ-LSQLQH*SPFQKFQLFEPFPFHSH 164
Query: 194 *ILNPSQLMKSNLRSAC--KQPNWPRKSQV 221
+L QL+ NL +QP+WPR SQ+
Sbjct: 163 PLLQHQQLLSGNLSHQLHPRQPHWPRPSQM 74
>TC18501
Length = 591
Score = 27.3 bits (59), Expect = 4.3
Identities = 11/24 (45%), Positives = 19/24 (78%)
Frame = -3
Query: 169 GKQNSSQKLSIISQILCQLRHL*L 192
G+ +SSQ L+++ ILC ++H+*L
Sbjct: 421 GQSSSSQSLTLLDIILCLVQHI*L 350
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 26.9 bits (58), Expect = 5.6
Identities = 25/70 (35%), Positives = 35/70 (49%), Gaps = 6/70 (8%)
Frame = +1
Query: 67 ISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQK--INAFKAPQEKL----STIMNCCRV 120
I+LL +PE+ P + + L EKEL K + AF + L ST NC RV
Sbjct: 2185 IALLCVQQRPEYR---PDMLSVVLMLNGEKELPKPSLPAFYTGNDDLLWPESTSKNCERV 2355
Query: 121 INNLLLNAAM 130
I +++LN M
Sbjct: 2356 IKHVILNGFM 2385
>TC18125 similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X carboxypeptidase,
partial (22%)
Length = 585
Score = 26.2 bits (56), Expect = 9.5
Identities = 20/81 (24%), Positives = 32/81 (38%)
Frame = +3
Query: 16 IRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKIDHEISEKISLLQTFLK 75
+ P W T TE + L+K+ +FS + ISE I L T
Sbjct: 87 VEPRPKWIT-TEFGGHNILAPLKKFGSNIIFSNGLLDPWTGGSVLQNISESIVSLVTKEG 263
Query: 76 PEHLDIPPVLHNEASWLLAEK 96
H+D+ N+ WL+ ++
Sbjct: 264 AHHIDLRASTGNDPDWLVEQR 326
>TC10697
Length = 487
Score = 26.2 bits (56), Expect = 9.5
Identities = 19/45 (42%), Positives = 27/45 (59%), Gaps = 1/45 (2%)
Frame = +2
Query: 171 QNSSQKLSIISQILCQLRHL*LN*ILNPSQLMKSNLRSAC-KQPN 214
QNSS + ++ +LC+L +L L+* L + L K NL C K PN
Sbjct: 287 QNSSDVVFVLHILLCELLYL-LS*ALLYATLEKLNLTYDCMKVPN 418
>TC9008 homologue to PIR|T06773|T06773 ferredoxin-NADP reductase - garden
pea (fragment) {Pisum sativum;} ,
partial (18%)
Length = 737
Score = 26.2 bits (56), Expect = 9.5
Identities = 17/62 (27%), Positives = 27/62 (43%)
Frame = +2
Query: 290 VLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMSEKEPLLHQLE 349
V+ HG N PY E +S D+LL I+C + +S +E ++ L
Sbjct: 203 VIDHGGNVPYWEGQSYGFIPPVSDLLL---------ILILCFRSSLYLISGRESIMIDLR 355
Query: 350 MQ 351
+Q
Sbjct: 356 LQ 361
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,276,069
Number of Sequences: 28460
Number of extensions: 79176
Number of successful extensions: 482
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 481
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 482
length of query: 378
length of database: 4,897,600
effective HSP length: 92
effective length of query: 286
effective length of database: 2,279,280
effective search space: 651874080
effective search space used: 651874080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147959.10


