

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
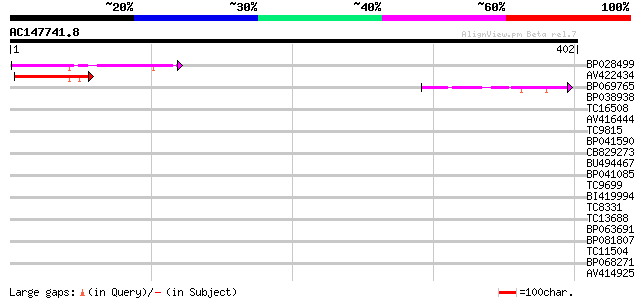
Query= AC147741.8 - phase: 0
(402 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP028499 87 4e-18
AV422434 62 1e-10
BP069765 60 8e-10
BP038938 32 0.19
TC16508 30 0.72
AV416444 30 0.72
TC9815 weakly similar to UP|Q9FEA1 (Q9FEA1) Anthocyanin 1, parti... 29 1.2
BP041590 29 1.6
CB829273 28 2.1
BU494467 28 2.1
BP041085 28 3.6
TC9699 similar to UP|APH1_ARATH (Q8L9G7) Gamma-secretase subunit... 28 3.6
BI419994 28 3.6
TC8331 similar to UP|O04616 (O04616) CODED for BY A. THALIANA CD... 27 4.6
TC13688 similar to UP|O22161 (O22161) At2g44900 protein, partial... 27 6.1
BP063691 27 6.1
BP081807 27 7.9
TC11504 similar to UP|O82273 (O82273) At2g31110 protein, partial... 27 7.9
BP068271 27 7.9
AV414925 27 7.9
>BP028499
Length = 491
Score = 87.4 bits (215), Expect = 4e-18
Identities = 55/129 (42%), Positives = 74/129 (56%), Gaps = 8/129 (6%)
Frame = +2
Query: 2 ATVDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSS----VPKHHHKFRFQFPLLK 57
A DWS+LP ELL+LISQR+ + L+ FRSVCSTWR SS +P HH F F FP L
Sbjct: 107 APPDWSQLPAELLHLISQRLNSPLYLLRFRSVCSTWRHSSASSFIPIHH--FPFNFPPL- 277
Query: 58 FPFNADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRI----CQNARGKSKLYHP 113
+D + + S +TK+ + LI PP T P L++I C R +++L+HP
Sbjct: 278 ----SDHDHHDASSFSLTKRTVVLITPPPPPHHQTQTPWLVKIGEDLCDGDRTRTRLWHP 445
Query: 114 LLQNGYKFP 122
L + G FP
Sbjct: 446 LFR-GPHFP 469
>AV422434
Length = 422
Score = 62.4 bits (150), Expect = 1e-10
Identities = 35/63 (55%), Positives = 41/63 (64%), Gaps = 7/63 (11%)
Frame = +3
Query: 4 VDWSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSS----VPKHHHK---FRFQFPLL 56
VDWS+LPTELL+LISQR+ + L+ FRSVCSTWR SS P HH R +F LL
Sbjct: 54 VDWSQLPTELLHLISQRLDSPLYLLRFRSVCSTWRRSSSSAVTPLHHFPNPLRRHRFHLL 233
Query: 57 KFP 59
P
Sbjct: 234 PLP 242
Score = 35.8 bits (81), Expect = 0.013
Identities = 26/87 (29%), Positives = 40/87 (45%), Gaps = 4/87 (4%)
Frame = +1
Query: 32 SVCSTWRSSSVPKHHHKFRFQFP----LLKFPFNADFINKNRSFCYITKQNIFLIKPPQQ 87
S+ + S+S P H P FP +D S +++++ +FLI PP
Sbjct: 106 SIARSTSSASAPSVPHGAALPHPPSRPFTTFPTLSD---DTASTFFLSQRTVFLIIPPPL 276
Query: 88 EQTLTLRPLLIRICQNARGKSKLYHPL 114
+ T L PLL +I Q+ +LY PL
Sbjct: 277 QPTEPLTPLLFQISQDNG*TRRLYDPL 357
>BP069765
Length = 417
Score = 59.7 bits (143), Expect = 8e-10
Identities = 44/117 (37%), Positives = 62/117 (52%), Gaps = 10/117 (8%)
Frame = -2
Query: 293 KKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFNNYIFESHRPLESECKEYV 352
+ WV + ++ DRVLF+G C+FS ASDL V +GN V+F + L+S +
Sbjct: 416 ESWVHVRNVRDRVLFIGDD--CAFSVSASDLGVDEGNYVVFRD------DDLKSGIGVFP 261
Query: 353 LDLDLGRHS----LLSDYPEYSNLFWPPPEW------IRPRYDVSYLASKGFYFPNV 399
LD D+G + L DYP +S+LF PPP+W R + Y+ S GF F V
Sbjct: 260 LD-DVGWYGDGILHLLDYPGFSSLFRPPPDWAMLH*QTREYVCMQYVTSNGF*FEAV 93
>BP038938
Length = 583
Score = 32.0 bits (71), Expect = 0.19
Identities = 24/104 (23%), Positives = 39/104 (37%), Gaps = 14/104 (13%)
Frame = +3
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSV--------------PKHHHKFRF 51
WS LP ELL L+ R+ + D + VC W S + PK+ + F
Sbjct: 135 WSDLPAELLELVMTRLALD-DNVRASVVCKRWHSVATAVRVVNQSPWLMYFPKYGDLYEF 311
Query: 52 QFPLLKFPFNADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRP 95
P+ + ++ + S TK L+ P+ + P
Sbjct: 312 YDPVQRKTYSLQMPELSGSRVCYTKDGWLLLYRPRTHRVFFFNP 443
>TC16508
Length = 572
Score = 30.0 bits (66), Expect = 0.72
Identities = 15/42 (35%), Positives = 20/42 (46%)
Frame = +3
Query: 83 KPPQQEQTLTLRPLLIRICQNARGKSKLYHPLLQNGYKFPHT 124
K P + L LR L R+C +RG HP + N + F T
Sbjct: 120 KQPSPLERLNLRQSLKRLCPRSRGILSSQHPCICNFFTFQET 245
>AV416444
Length = 414
Score = 30.0 bits (66), Expect = 0.72
Identities = 16/42 (38%), Positives = 24/42 (57%), Gaps = 2/42 (4%)
Frame = +1
Query: 32 SVCSTWRSSSVPKHHHKF--RFQFPLLKFPFNADFINKNRSF 71
++ S +R+ HHH++ F FPLLK P N F + + SF
Sbjct: 85 TLLSVYRTHDFFSHHHQYFLLFSFPLLKHP-NHSFSSSSHSF 207
>TC9815 weakly similar to UP|Q9FEA1 (Q9FEA1) Anthocyanin 1, partial (11%)
Length = 634
Score = 29.3 bits (64), Expect = 1.2
Identities = 21/65 (32%), Positives = 29/65 (44%), Gaps = 6/65 (9%)
Frame = -3
Query: 100 ICQNARGKSKLYHPLLQNGYKFPHTFMLDFNKLSILHLGSNFFMIDFDFTF------NDK 153
+CQ R S+L+HP F M+ FN L + FF + + FTF N
Sbjct: 299 VCQVVRQYSRLFHPCTFQTKTFLVCGMI*FNALFTSTM-LTFFPLTYSFTFALNSATNTP 123
Query: 154 LYNPD 158
L+N D
Sbjct: 122 LFNED 108
>BP041590
Length = 434
Score = 28.9 bits (63), Expect = 1.6
Identities = 16/53 (30%), Positives = 23/53 (43%), Gaps = 7/53 (13%)
Frame = +3
Query: 357 LGRHSLLSDYPEYSNLFWPPPEWIRPRYDVSYLAS-------KGFYFPNVTMI 402
LGR + DYP + P W++ R + AS +G+ P TMI
Sbjct: 111 LGRKCITEDYPSGIYELYQPKRWVKKRSEEMLWASDLQSH*YRGYEIPTSTMI 269
>CB829273
Length = 551
Score = 28.5 bits (62), Expect = 2.1
Identities = 19/53 (35%), Positives = 24/53 (44%), Gaps = 13/53 (24%)
Frame = -2
Query: 8 KLPTELLNLISQRIYEEVDLIHFRSVCSTW---------RSSSVP----KHHH 47
+LP+ LNL + LIH RS+C +W RSS P HHH
Sbjct: 370 RLPSHFLNLQHLHPWNHYRLIH*RSLCLSWMYHSQYFCCRSSLHPLQLLPHHH 212
>BU494467
Length = 334
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/34 (38%), Positives = 19/34 (55%), Gaps = 7/34 (20%)
Frame = -3
Query: 363 LSDYPEYSNL-------FWPPPEWIRPRYDVSYL 389
LS PE++ L FWPPP W+ ++V +L
Sbjct: 218 LSWSPEFAPLLIFALEKFWPPPPWLLNEFEVPWL 117
>BP041085
Length = 449
Score = 27.7 bits (60), Expect = 3.6
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = -1
Query: 373 FWPPPEWIRPRYDVSYL 389
FWPPP W+ ++V +L
Sbjct: 173 FWPPPPWLLNEFEVPWL 123
>TC9699 similar to UP|APH1_ARATH (Q8L9G7) Gamma-secretase subunit
APH1-like, partial (96%)
Length = 1070
Score = 27.7 bits (60), Expect = 3.6
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 6/59 (10%)
Frame = -1
Query: 269 FTDLCNP----DHNDCVRIDLFKLNE--KEKKWVKLTSLGDRVLFLGLGLVCSFSACAS 321
F LC P DH +CV D K N+ K++KW S+ + +CS++ C S
Sbjct: 593 FVALCIPIKRDDH*ECVNRDKCKTNKCRKKEKWNLRASVNE---------ICSWTKCRS 444
>BI419994
Length = 549
Score = 27.7 bits (60), Expect = 3.6
Identities = 20/66 (30%), Positives = 29/66 (43%)
Frame = +2
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNADFI 65
+S LP EL+ I R+ L+ F++VC +WR K F L P + F+
Sbjct: 38 FSHLPDELILQILLRLPVR-SLLRFKTVCKSWRYLISDPQFTKSHFN--LAASPTHRIFL 208
Query: 66 NKNRSF 71
N F
Sbjct: 209 NNTNGF 226
>TC8331 similar to UP|O04616 (O04616) CODED for BY A. THALIANA CDNA T46771
(AT4g01150/F2N1_18), partial (73%)
Length = 724
Score = 27.3 bits (59), Expect = 4.6
Identities = 16/45 (35%), Positives = 20/45 (43%), Gaps = 2/45 (4%)
Frame = +2
Query: 29 HFRSVCSTWRSSSVPKHHHKFRF--QFPLLKFPFNADFINKNRSF 71
H+ +C T SSS HHH F FP F F ++SF
Sbjct: 128 HYSLLCFTLSSSSCLHHHHHHHFVLTFP*TLFRVPKTFSASDKSF 262
>TC13688 similar to UP|O22161 (O22161) At2g44900 protein, partial (7%)
Length = 621
Score = 26.9 bits (58), Expect = 6.1
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 9/55 (16%)
Frame = -3
Query: 4 VDWSKLPTE-LLNLISQRIYEEVDLIHFRSVCSTWRS--------SSVPKHHHKF 49
V+W LP + ++ L+S Y D F S C TWRS +S+ HKF
Sbjct: 169 VEWRGLPDDTVIQLLSCLSYR--DRASFSSSCRTWRSLGSSPCLWTSLDLRSHKF 11
>BP063691
Length = 429
Score = 26.9 bits (58), Expect = 6.1
Identities = 10/38 (26%), Positives = 19/38 (49%)
Frame = -2
Query: 14 LNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRF 51
LN I + + E+ ++H+ + W S H H F++
Sbjct: 302 LNFIRELGFGELVILHYVDIFFVWNQGSC*DHXHSFQY 189
>BP081807
Length = 302
Score = 26.6 bits (57), Expect = 7.9
Identities = 13/49 (26%), Positives = 26/49 (52%)
Frame = +3
Query: 286 FKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFN 334
F + +K ++L + R++ G +CS S CAS + +++IF+
Sbjct: 108 FSSSHNQKLPLQLQQVVSRMVKSGFLFICSLSICASAF*FRRTHAIIFS 254
>TC11504 similar to UP|O82273 (O82273) At2g31110 protein, partial (40%)
Length = 579
Score = 26.6 bits (57), Expect = 7.9
Identities = 14/49 (28%), Positives = 25/49 (50%)
Frame = -2
Query: 313 VCSFSACASDLCVAKGNSVIFNNYIFESHRPLESECKEYVLDLDLGRHS 361
+ SFS C C KGN + + YI+ + S+C + L ++ ++S
Sbjct: 569 ITSFSCCILI*CFRKGNFIRY--YIYNEPHQVLSKCLIFTLQNNIWKNS 429
>BP068271
Length = 437
Score = 26.6 bits (57), Expect = 7.9
Identities = 13/29 (44%), Positives = 14/29 (47%)
Frame = -2
Query: 361 SLLSDYPEYSNLFWPPPEWIRPRYDVSYL 389
SLL + S WPPP W R V YL
Sbjct: 241 SLLDEMRRKSAHRWPPPPWRRNSNGVCYL 155
>AV414925
Length = 414
Score = 26.6 bits (57), Expect = 7.9
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +1
Query: 8 KLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHK 48
++P E++ I R+ L+ FRS+ +WRS KH K
Sbjct: 94 EIPVEVIADILSRL-PVTSLLRFRSISKSWRSLIDSKHFMK 213
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.142 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,355,399
Number of Sequences: 28460
Number of extensions: 162017
Number of successful extensions: 782
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 777
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 777
length of query: 402
length of database: 4,897,600
effective HSP length: 92
effective length of query: 310
effective length of database: 2,279,280
effective search space: 706576800
effective search space used: 706576800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147741.8


