

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.2 - phase: 0 /pseudo
(971 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
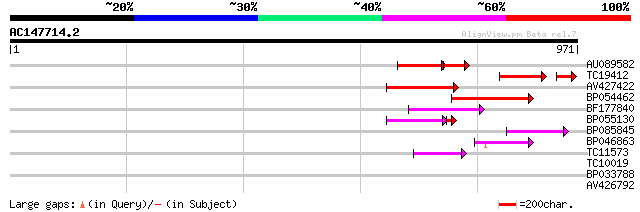
AU089582 104 1e-39
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 101 1e-27
AV427422 110 1e-24
BP054462 94 8e-20
BF177840 93 2e-19
BP055130 86 2e-18
BP085845 59 5e-09
BP046863 59 5e-09
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 55 4e-08
TC10019 weakly similar to UP|AAO17787 (AAO17787) Lecithin diacyl... 31 0.82
BP033788 30 2.4
AV426792 28 9.1
>AU089582
Length = 383
Score = 104 bits (260), Expect(2) = 1e-39
Identities = 50/83 (60%), Positives = 63/83 (75%)
Frame = +2
Query: 665 SQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSE 724
SQ +IY+ EIV GVP SI+SDR +FTS FW+S Q ALG++L++S+A+HPQTDGQSE
Sbjct: 14 SQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSE 193
Query: 725 RTIQSLEDLLRICVLEQGGTWDS 747
RTIQ LED+LR CV + G S
Sbjct: 194 RTIQILEDMLRACVXDLRGXLGS 262
Score = 77.0 bits (188), Expect(2) = 1e-39
Identities = 33/45 (73%), Positives = 38/45 (84%)
Frame = +3
Query: 743 GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESG 787
G+WD +L L+EF YNNSY SSI MAPFEALYGRRCR+P+ WFE G
Sbjct: 249 GSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPIGWFEVG 383
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 101 bits (252), Expect(2) = 1e-27
Identities = 48/79 (60%), Positives = 64/79 (80%)
Frame = +3
Query: 840 RVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYV 899
RV+PM GV R K KL+P+FIGP+++LERVG+V+YR+ LPP LS +H VFHVS LRKY+
Sbjct: 3 RVSPMKGVLRFGKKGKLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRKYL 182
Query: 900 PDPSHVIQSDDVQVRDNLT 918
DPSHVI+ +DVQ+ +L+
Sbjct: 183 YDPSHVIRHEDVQLDVHLS 239
Score = 39.7 bits (91), Expect(2) = 1e-27
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = +2
Query: 937 KEIPLVRVVWTGATSESLTWELESKMLESYPELF 970
K++ V+V+W G + E TWE E M E YP LF
Sbjct: 296 KDVGSVKVLWRGPSGEEATWEAEDIMREKYPHLF 397
>AV427422
Length = 417
Score = 110 bits (275), Expect = 1e-24
Identities = 52/124 (41%), Positives = 77/124 (61%)
Frame = +1
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
++ R K +HF+ + +I+++E+V+ GVP SIVSDRDP F S FWK L +
Sbjct: 46 VVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMSNFWKELFKM 225
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI 764
G+KL++S+AYHP++DGQ+E + LE LR + +Q +W +P E+ YN SYH S
Sbjct: 226 QGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYWYNTSYHVST 405
Query: 765 GMAP 768
G P
Sbjct: 406 GQTP 417
>BP054462
Length = 422
Score = 94.4 bits (233), Expect = 8e-20
Identities = 51/141 (36%), Positives = 85/141 (60%), Gaps = 1/141 (0%)
Frame = +1
Query: 757 NNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQS 816
N++Y+ S M+PF+ALYGR L ++ ++ E + ++ + SQ
Sbjct: 1 NSNYNRSAKMSPFQALYGREPPVLLQGTTIPSKIAAVNDLQVGRDELLSDLRANLLKSQD 180
Query: 817 RQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALK-SKKLTPKFIGPYQILERVGTVAY 875
++Y +K+R+D+++Q GD VFL++ P A K ++KL+P++ GPY I+ ++G VAY
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAY 360
Query: 876 RVGLPPHLSNLHNVFHVSQLR 896
R+ LP H S +H VFHVS L+
Sbjct: 361 RLELPAH-SRVHPVFHVSLLK 420
>BF177840
Length = 410
Score = 93.2 bits (230), Expect = 2e-19
Identities = 52/130 (40%), Positives = 74/130 (56%)
Frame = +2
Query: 683 SSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQG 742
+SIVSDRD +F S FW++L +G+KL S+ HPQTDGQ+E ++L LLR +
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 743 GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTE 802
W++ LP IEF YN HS+ +PFE +YG TPL +L + + E
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDLLPMPNTYLLKHKDGKAKAE 370
Query: 803 KVQMIQEKMK 812
V+ + EK+K
Sbjct: 371 FVKRLHEKIK 400
>BP055130
Length = 567
Score = 86.3 bits (212), Expect(2) = 2e-18
Identities = 46/105 (43%), Positives = 61/105 (57%)
Frame = +2
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
++ R K HF ++ SQ E ++ IVK G+P +IVSDRD FTS FWK +
Sbjct: 200 VVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHFFKL 379
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHL 749
G+ L +SS+YHPQTDGQ+E + LE LR V E W S+L
Sbjct: 380 HGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYL 514
Score = 24.3 bits (51), Expect(2) = 2e-18
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +3
Query: 749 LPLIEFTYNNSYHSSIG 765
LP E+ YN+S+ +SIG
Sbjct: 516 LPWAEYCYNSSFQTSIG 566
>BP085845
Length = 464
Score = 58.5 bits (140), Expect = 5e-09
Identities = 35/106 (33%), Positives = 57/106 (53%)
Frame = -2
Query: 852 KSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDV 911
K +K K IGP++ILE V +AY + +L H V HV+ L + + D S I + V
Sbjct: 421 KKRKTESKIIGPFKILEMVCLIAY*LTHSLYLLAAHIVLHVTLLWRNLYDQSQNICHEGV 242
Query: 912 QVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWE 957
+ ++ + P+ + D KV+ +R K I V+V+W G + +WE
Sbjct: 241 XLGEHWSHMEHPIVMVDMKVRCMRPKNIDDVKVIWRGLSG*EKSWE 104
>BP046863
Length = 580
Score = 58.5 bits (140), Expect = 5e-09
Identities = 38/113 (33%), Positives = 65/113 (56%), Gaps = 12/113 (10%)
Frame = +1
Query: 797 VQQTTEKVQMIQEKMKA-----------SQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMT 845
++ + E V M+ E+ A +Q + ++ +K R+ +++Q G+ VFL++ P
Sbjct: 244 IKLSVEAVNMLIEERDALLLELRGNLLKAQDQMRAQANKHRRYVDYQVGNWVFLKLQPYK 423
Query: 846 GVXRAL-KSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRK 897
A K++KL+P+F GP+++LERV VAY + L S +H VFH+S L K
Sbjct: 424 LQNLAQRKNQKLSPRFYGPFKVLERVVQVAY*LDLXSE-SRVHPVFHLSLLEK 579
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 55.5 bits (132), Expect = 4e-08
Identities = 29/90 (32%), Positives = 48/90 (53%)
Frame = +2
Query: 692 RFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPL 751
+FTS+ + + +G ++R SS HPQT+GQ+E + + ++ + E G W LP+
Sbjct: 20 QFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWIDELPI 199
Query: 752 IEFTYNNSYHSSIGMAPFEALYGRRCRTPL 781
+ ++YN SSI PF YG P+
Sbjct: 200 VLWSYNTMPQSSIKETPF*LTYGADTMLPV 289
>TC10019 weakly similar to UP|AAO17787 (AAO17787) Lecithin diacylglycerol
cholesterol acyltransferase, partial (10%)
Length = 570
Score = 31.2 bits (69), Expect = 0.82
Identities = 15/33 (45%), Positives = 20/33 (60%)
Frame = -2
Query: 450 DIFLHPIGSKSLSFAPRLNPSNIHS*NKIKCNL 482
DI +HPI S+ SF P NP NI + K+ N+
Sbjct: 116 DIRIHPITSQIFSFCPSCNPYNILNQCKVSHNI 18
>BP033788
Length = 518
Score = 29.6 bits (65), Expect = 2.4
Identities = 15/43 (34%), Positives = 25/43 (57%)
Frame = +3
Query: 699 KSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQ 741
KSLQ G++++L + P+ D ERT+Q D +I V ++
Sbjct: 75 KSLQTKSGARIQLIPQHLPEGDDSKERTVQVTGDKRQIEVAQE 203
>AV426792
Length = 432
Score = 27.7 bits (60), Expect = 9.1
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +1
Query: 798 QQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHV 837
Q+ K Q +EK KA ++ K+ HDK + EGD V
Sbjct: 280 QEARRKKQREEEKRKAEEAAAKATHDKTIEGGNQTEGDLV 399
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.350 0.155 0.524
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,275,120
Number of Sequences: 28460
Number of extensions: 195199
Number of successful extensions: 1587
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1574
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1583
length of query: 971
length of database: 4,897,600
effective HSP length: 99
effective length of query: 872
effective length of database: 2,080,060
effective search space: 1813812320
effective search space used: 1813812320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147714.2


